3.1: Amino Acids and Peptides
- Page ID
- 14927
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)-
Understand the Role and Diversity of Proteins:
- Describe the various functions of proteins (structural, regulatory, enzymatic, transport, etc.) and explain why proteins are considered the most functionally diverse macromolecules.
- Recognize that proteins are linear polymers of alpha-amino acids linked by peptide bonds.
-
Comprehend Alpha Amino Acid Structure:
- Identify the common structural features of an alpha amino acid (α-amino group, α-carboxylic acid, hydrogen, and a variable R-group).
- Memorize the 20 naturally occurring amino acids along with their three-letter and one-letter abbreviations.
- Explain how variations in the R-group confer different chemical properties (nonpolar, polar uncharged, acidic, and basic) to amino acids.
-
Master Peptide Bond Formation and Protein Primary Structure:
- Describe the mechanism of peptide bond formation via nucleophilic attack and the release of water.
- Illustrate the concept of primary protein structure as the linear sequence of amino acids and appreciate the enormous diversity of possible sequences.
-
Analyze Amino Acid Classification and Side Chain Characteristics:
- Categorize amino acids based on their side chain properties (e.g., aliphatic vs. aromatic nonpolar, polar uncharged, acidic, and basic).
- Discuss how side chain properties influence protein folding, stability, and function.
- Interpret hydrophobicity (or hydropathy) scales (e.g., Kyte-Doolittle, Hopp-Woods) and relate these to protein topology (buried versus surface residues).
-
Grasp Stereochemistry and Chirality in Amino Acids:
- Explain the concept of chirality and why all naturally occurring proteinogenic amino acids (except glycine) are L isomers.
- Differentiate between the D/L nomenclature and the R/S system, and understand why the D/L system is preferred in biochemistry.
-
Apply Acid-Base Chemistry to Proteins:
- Utilize the Henderson–Hasselbalch equation to predict the ionization state of amino acid side chains and terminal groups.
- Interpret titration curves for amino acids and proteins and calculate the isoelectric point (pI) of proteins based on the ionizable groups present.
-
Examine Chemical Reactivity of Amino Acid Side Chains:
- Identify which side chains serve as hydrogen bond donors and acceptors, and explain their roles in catalysis and substrate binding.
- Compare the nucleophilicity and basicity of key amino acid side chains (e.g., lysine, cysteine, histidine) and discuss factors (like electronegativity and steric effects) that influence reactivity.
- Describe specific chemical reactions involving amino acid side chains, including acylation, Schiff base formation, and nucleophilic substitution reactions used in protein modification.
-
Explore Post-Translational Modifications (PTMs) and Their Biological Significance:
- List common PTMs (e.g., phosphorylation, acetylation, glycosylation, oxidation) and explain how these modifications alter protein structure and function.
- Understand how aberrant or deleterious PTMs (e.g., glycation, carbonylation) can lead to altered protein activity and contribute to disease processes.
These learning goals are intended to help students integrate structural, chemical, and functional perspectives on proteins, preparing them for more advanced topics in enzymology, protein engineering, and molecular biology.
Introduction
Proteins are one of the most abundant organic molecules in living systems and have the most diverse range of functions of all macromolecules. Proteins may be structural, regulatory, contractile, or protective. They may serve in transport, storage, or membranes. They may be toxins or enzymes. Each cell in a living system may contain thousands of proteins, each with a unique function. Their structures, like their functions, vary greatly. They are all, however, polymers of alpha amino acids arranged in a linear sequence and connected by covalent bonds.
Alpha Amino Acid Structure
The major building blocks of proteins are called alpha (α) amino acids. As their name implies, they contain a carboxylic acid and an amine functional group. The alpha designation indicates that a single carbon atom separates these two functional groups. In addition to the amine and carboxylic acid, the alpha carbon is also attached to a hydrogen atom and to one additional group that can vary in size and length. In the diagram below, this group is designated as an R-group. Within living organisms, 20 common amino acids are used as protein building blocks. They differ only in the R-group position. The fully protonated structure of an amino acid (at low pH) is shown in Figure \(\PageIndex{1}\).
The twenty common naturally occurring amino acids contain an alpha-carbon, an amino, a carboxylic acid, and an R group (or side chain). The R group side chains may be either nonpolar, polar uncharged, or charged, depending on the functional group, the pH, and the pKa of any ionizable group in the side chain.
Two other amino acids occasionally appear in proteins. One is selenocysteine, found in Archaea, eubacteria, and animals. Another is pyrrolysine, found in Archaea. Bacteria have been modified to incorporate two new amino acids, O-methyl-tyrosine and p-aminophenylalanine. The yeast strain Saccharomyces cerevisiae has been engineered to incorporate five new unnatural amino acids (using the TAG nonsense codon and new, modified tRNA and tRNA synthetases) with keto groups that allow chemical modifications to the protein. We will concentrate only on the 20 abundant, naturally occurring amino acids.
Proteins are polymers of monomeric amino acids with an amide link (also called a peptide bond) between the α-carboxylic group of one amino acid and the α-amine of the next one. Figure \(\PageIndex{2}\) shows the twenty naturally occurring α-amino acids as they would appear internally within a protein sequence. The squiggles show the connecting amide/peptide bond between adjacent amino acids. Students often assume that the α-amino and α-carboxylic acid groups within a protein sequence are free and not part of the peptide bond. This figure should help in resolving that misconception. The three-letter and one-letter abbreviations of each amino acid are shown, as well as the typical pKa values of side chains (R groups). It is important to memorize the three-letter and one-letter codes for the amino acids.
Amino acids form polymers through a nucleophilic attack by the amino group of an amino acid at the electrophilic carbonyl carbon of the carboxyl group of another amino acid. The carboxyl group of the amino acid must first be activated to provide a better leaving group than OH-. The resulting link between the amino acids is an amide link, which biochemists call a peptide bond. In this reaction, water is released. In a reverse reaction, the peptide bond can be cleaved by water (hydrolysis). This is illustrated in Figure \(\PageIndex{3}\).
Proteins are polymers of twenty naturally occurring amino acids. In contrast, nucleic acids are polymers of just four different monomeric nucleotides. The sequence of a protein and its total length differentiate one protein from another. Just for an octapeptide, there are over 25 billion different possible arrangements of amino acids. Compare this to just 65536 oligonucleotides (4 different monomeric deoxynucleotides) of 8 monomeric units, an 8mer. Hence, the diversity of possible proteins is enormous.
The resulting structure is called a dipeptide when an amide bond links two amino acids. Likewise, we can have tripeptides, tetrapeptides, and other polypeptides. At some point, when the structure is long enough, it is called a protein. The average molecular weight of yeast proteins is about 50,000, with about 450 amino acids. The largest human protein might be titin, with a molecular weight of about 3 million (about 30,0000 amino acids). A new class of very small proteins (30 or fewer amino acids, and perhaps better named polypeptides) called smORFs (small open reading frames) has recently been discovered to have significant biological activity. These are encoded directly in the genome and are produced by the same processes that produce regular proteins (DNA transcription and RNA translation). They are not the result of the selective cleavage of a larger protein into smaller peptide fragments.
Figure \(\PageIndex{4}\) shows several ways to represent the structure of a polypeptide or protein, each showing differing amounts of information. Note that the atoms in the side chains are denoted alpha, beta, gamma, delta, epsilon ...
Characteristics of Amino Acids
The different R-groups exhibit distinct characteristics depending on the nature of the atoms incorporated into the functional groups. Some R-groups predominantly contain carbon and hydrogen and are very nonpolar or hydrophobic. Others contain polar uncharged functional groups such as alcohols, amides, and thiols. A few amino acids are basic (containing amine functional groups) or acidic (containing carboxylic acid functional groups). These amino acid side chains can carry full charges and engage in ionic interactions. Each amino acid can be abbreviated using a three-letter and a one-letter code. Figure \(\PageIndex{5}\) shows groupings of the amino acids based on their side chain properties.
The above classification for a few amino acids is somewhat arbitrary, as described below. For example, cysteine is often buried in protein in a more hydrophobic environment.
Nonpolar (Hydrophobic) Amino Acids
The nonpolar amino acids can be subdivided into two more specific classes: aliphatic and aromatic. The aliphatic amino acids (glycine, alanine, valine, leucine, isoleucine, and proline) typically contain straight or branched hydrocarbon chains, with two exceptions. Glycine, the simplest amino acid, has a single hydrogen atoms as the "side" chain. Proline is also classified as an aliphatic amino acid, but the hydrocarbon side chain has cyclized with the terminal amine, creating a unique 5-membered ring, non-aromatic ring structure. As we will see in the next section on primary structure, proline can significantly alter the 3-dimensional structure of the protein due to the rigidity of its ring when incorporated into the polypeptide chain, and it is commonly found in regions of the protein where folds or turns occur.
As their names imply, the aromatic amino acids (phenylalanine, tyrosine, and tryptophan) contain an aromatic functional group, making them largely nonpolar and hydrophobic due to their high carbon-to-hydrogen ratio. However, it should be noted that hydrophobicity and hydrophilicity represent a sliding scale, and each amino acid can have different physical and chemical properties depending on its structure. For example, the hydroxyl group in tyrosine increases its reactivity and solubility compared to phenylalanine.
Methionine, one of the sulfur-containing amino acids, is usually classified as a nonpolar, hydrophobic amino acid. The terminal methyl group is a thioether, which generally cannot form a permanent dipole within the molecule and retains low solubility.
Polar (Hydrophilic) Amino Acids
The polar, hydrophilic amino acids can be subdivided into three major classes: the polar uncharged, acidic, and basic functional groups. Within the polar uncharged class, the side chains contain heteroatoms (O, S, or N) capable of forming permanent dipoles within the R-group. These include the hydroxyl- and sulfhydryl-containing amino acids, serine, threonine, and cysteine, and the amide-containing amino acids, glutamine and asparagine. Two amino acids, glutamic acid (glutamate), and aspartic acid (aspartate) constitute the acidic amino acids and contain side chains with carboxylic acid functional groups capable of fully ionizing in solution. The basic amino acids, lysine, arginine, and histidine, contain amine functional groups that can be protonated to carry a full charge.
Many amino acids with hydrophilic R groups can participate in the active sites of enzymes. An active site is the part of an enzyme that directly binds to a substrate and carries out a reaction. Protein-derived enzymes contain catalytic groups consisting of amino acid R-groups that promote the formation and degradation of bonds. The amino acids that play a significant role in the binding specificity of the active site are usually not adjacent to each other in the primary structure but form the active site as a result of folding in creating the tertiary structure, as you will see later in the chapter.
Tryptophan contains an amine functional group. Why isn't tryptophan basic?
Answer
Tryptophan contains an indole ring structure with the amine functional group. However, due to the proximity of and electron-withdrawing nature of the aromatic ring structure, the lone pair of electrons on the nitrogen is unavailable to accept a proton. Instead, they form pi-bonds within several of the different resonance structures possible for the indole ring. Figure 2.3A shows four of the possible resonance structures for indole. Conversely, within the imidazole ring structure found in histidine, there are two nitrogen atoms, one of which is involved in the formation of resonance structures (Nitrogen #1 in Figure 2.3B) and cannot accept a proton, and the other (Nitrogen #3) that has a lone pair of electrons that is available to accept a proton.

Exercise \(\PageIndex{1}\)
Given the example above, describe, using a chemical diagram, why the amide nitrogen atoms found in asparagine and glutamine are not basic.
- Answer
-
The lone pair is delocalized into the peptide bond (via resonance), so it is unavailable for sharing.
Recent Updates: 5/5/24
Quantitative measures of amino acid polarity and hydrophobicity
There are quantitative methods to determine the relative polarity of an amino acid side chain, so it's not just a matter of visual inspection or guesswork. Let's consider hydrophobicity or hydropathy scales. Most are based on the standard free energy of transfer of a side chain from water to a nonpolar solvent. Each amino acid side chain is assigned a number ranging from negative to positive. The Kyte-Doolittle and the Hopp-Woods hydropathy scales are the two commonly used scales, as shown in Table \(\PageIndex{1}\) below.
|
Amino Acid |
Kyte-Doolittle |
Hopp-Woods |
|---|---|---|
|
Alanine |
1.8 |
-0.5 |
|
Arginine |
-4.5 |
3.0 |
|
Asparagine |
-3.5 |
0.2 |
|
Aspartic acid |
-3.5 |
3.0 |
|
Cysteine |
2.5 |
-1.0 |
|
Glutamine |
-3.5 |
0.2 |
|
Glutamic acid |
-3.5 |
3.0 |
|
Glycine |
-0.4 |
0.0 |
|
Histidine |
-3.2 |
-0.5 |
|
Isoleucine |
4.5 |
-1.8 |
|
Leucine |
3.8 |
-1.8 |
|
Lysine |
-3.9 |
3.0 |
|
Methionine |
1.9 |
-1.3 |
|
Phenylalanine |
2.8 |
-2.5 |
|
Proline |
-1.6 |
0.0 |
|
Serine |
-0.8 |
0.3 |
|
Threonine |
-0.7 |
-0.4 |
|
Tryptophan |
-0.9 |
-3.4 |
|
Tyrosine |
-1.3 |
-2.3 |
|
Valine |
4.2 |
-1.5 |
Note that the Hopp-Woods scale is more like a hydrophilicity scale, since more polar residues have higher positive values. It was developed to identify likely antibody or other protein interaction sites on protein surfaces that display more hydrophilic side chains.
Some discrepancies exist in which amino acid side chains are nonpolar between the Kyte-Doolittle values and Figure \(\PageIndex{5}\). The Kyte-Doolittle scale shows that glycine, the two large aromatic tyrosine and tryptophan, and proline are more polar than nonpolar, and that cysteine is quite nonpolar.
As we will see in subsequent sections, a continuous stretch of amino acids found to have a high average hydrophobicity (low hydrophilicity) is probably buried in the interior of a protein away from the aqueous environment. Conversely, a continuous stretch with low hydrophobicity (high hydrophilicity) is likely buried in a protein or a membrane bilayer. Consider the example of the water-soluble bovine alpha-chymotrypsinogen, a 245 amino acid protein, whose sequence is shown below in single-letter code.
1 CGVPAIQPVLSGLSRIVNGEEAVPGSWPWQVSLQDKTGFHFCGGSLINENWVVTAAHCGV
61 TTSDVVVAGEFDQGSSSEKIQKLKIAKVFKNSKYNSLTINNDITLLKLSTAASFSQTVSA
121 VCLPSASDDFAAGTTCVTTGWGLTRYTNANTPDRLQQASLPLLSNTNCKKYWGTKIKDAM
181 ICAGASGVSSCMGDSGGPLVCKKNGAWTLVGIVSWGSSTCSTSTPGVYARVTALVNWVQQ
241 TLAAN
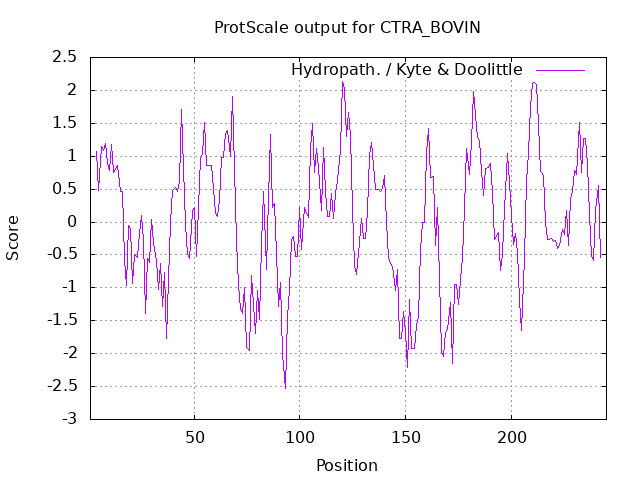
Let's use the ExPasy Prot Scale server to produce hydrophobicity plots of the protein bovine α-chymotrypsinogen. Input the Uniprot number P00766 for the protein or the sequence into the appropriate boxes. Select a length of continuous amino acids (called a window) of 7, and the program will calculate an average hydrophobicity for the "window." The window slides down the linear sequence, and a new value is calculated to determine a series of values for the entire sequence. Hydropathy plots (average score for the midpoint amino acid in the window) for chymotrypsinogen (window of seven consecutive residues) are shown in Figure \(\PageIndex{6}\) below. The Kyte-Doolittle scale (+ is hydrophobic) shows many stretches with high average values. The amino acids at those positions are likely buried in the protein's interior. The Hoop-Woods value (+ represents hydrophilic) indicates stretches with high average values. These amino acids are likely exposed to water. The two plots are complementary.
| Kyte-Doolittle (7 aa window) | Hopp-Woods (7 aa window) |
 |
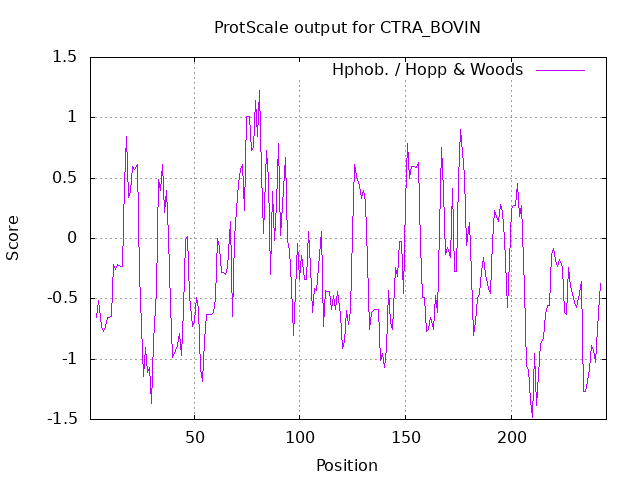 |
Figure \(\PageIndex{6}\): Kyte-Doolittle and Hopp-Woods plots for bovine α-chymotrypsinogen
Amino Acid Stereochemistry
The amino acids are all chiral except glycine, whose side chain is H. A chiral molecule is not superimposable with its mirror image. Like left and right hands that have a thumb and fingers in the same order but are mirror images and not the same, chiral molecules have the same things attached in the same order but are mirror images and not the same. The mirror image versions of chiral molecules have nearly identical physical properties, making it very difficult to tell them apart or separate them. Because of this, they are given a special stereoisomer name called enantiomers, and the compounds themselves are given the same name! These molecules differ in how they rotate plane-polarized light and in their interactions with biological molecules. Molecules that rotate light in the right-handed direction are called dextrorotary and are given a small "d" letter designation. Molecules that rotate light to the left are called levorotary and are designated with a small "l" to distinguish one enantiomer from the other. Biochemists also use the older nomenclature of the large "L" and "D" forms to characterize the 3D stereochemistry of amino acids. All naturally occurring proteins from all living organisms consist of L-amino acids based on their structural similarities to L-glyceraldehyde.
Again, the d- and l-designations are specific terms for how a molecule rotates plane-polarized light. It does not denote the absolute stereo configuration of a molecule. An absolute configuration refers to the spatial arrangement of the atoms of a chiral molecular entity (or group) and its modern stereochemical description, e.g., R or S, referring to Rectus or Sinister, respectively. Absolute configurations for a chiral molecule (in pure form) are often determined by X-ray crystallography. Alternative techniques are optical rotatory dispersion, vibrational circular dichroism, chiral shift reagents in proton NMR, and Coulomb explosion imaging. When the absolute configuration is known, the assignment of R or S is based on the Cahn–Ingold–Prelog priority rules. The absolute stereochemistry is related to L-glyceraldehyde, as shown below in Figure \(\PageIndex{6}\).
All naturally occurring amino acids in proteins are L, which corresponds to the S isomer, except cysteine. As shown in the bottom left of Figure \(\PageIndex{7}\) below, the absolute configuration of the amino acids can be shown with the H pointed to the rear, the COOH groups pointing out to the left, the R group to the right, and the NH3 group upwards. You can remember this with the mnemonic "CORN".
Figure \(\PageIndex{7}\): Stereochemistry of amino acids
Why does Biochemistry still use D and L for sugars and amino acids? This explanation (taken from a website that may no longer be available, so no reference is available) seems reasonable.
"In addition, however, chemists often need to define a configuration unambiguously in the absence of any reference compound, and for this purpose, the alternative (R,S) system is ideal, as it uses priority rules to specify configurations. These rules sometimes produce absurd results when applied to biochemical molecules. For example, as we have seen, all common amino acids are L because they all have the same structure, including the position of the R group (if we just write the R group as R). However, they do not all have the same configuration in the (R, S) system: L-cysteine is also (R)-cysteine. Still, all the other L-amino acids are (S), but this reflects the human decision to give a sulfur atom a higher priority than a carbon atom and does not reflect a real difference in configuration. Worse problems can sometimes arise in substitution reactions: sometimes, inversion of configuration can result in no change in the (R) or (S) prefix, and sometimes, retention of configuration can result in a change of prefix.
It follows that it is not just conservatism or a failure to understand the (R, S) system that causes biochemists to continue using D and L; it is simply that the DL system meets their needs much better. As mentioned, chemists also use D and L when appropriate. The explanation
given above of why the (R, S) system is little used in biochemistry is thus almost the exact opposite of reality. This system is the only practical way to unambiguously represent the stereochemistry of complex molecules with several asymmetric centers. Still, it is inconvenient with a regular series of molecules like amino acids and simple sugars."
If you are told to draw the correct stereochemistry of a molecule with one chiral C (S isomer, for example) and are given the substituents, you could do so easily following the R, S priority rules. However, how would you draw the correct isomer for the L isomer of the amino acid alanine? You couldn't do it without prior knowledge of the absolute configuration of the related molecule, L glyceraldehyde, or unless you remembered the anagram CORN. However, this disadvantage is more than made up for, as different L-amino acids with the same absolute stereochemistry might be labeled R or S, making this nomenclature unappealing to biochemists.
Amino Acid Charges
Monomeric amino acids have an alpha-amino group and a carboxyl group, both of which may be protonated or deprotonated, and an R group, some of which may be protonated or deprotonated. When protonated, the amino group carries a +1 charge, and the carboxyl group carries a zero charge. When deprotonated, the amino group is neutral, while the carboxyl group carries a -1 charge. The R groups that can be protonated/deprotonated include Lys, Arg, and His, which have a + 1 charge when protonated, and Glu and Asp (carboxylic acids), Tyr and Ser (alcohols), and Cys (thiol), which have zero charges when protonated. Of course, when the amino acids are linked by peptide bonds (amide link), the alpha N and the carboxyl C are in an amide link and are not charged.
However, the amino group of the N-terminal amino acid and the carboxyl group of the C-terminal amino acid of a protein may be charged. The Henderson-Hasselbalch equation allows us to determine the charge state of any ionizable group given its pKa. Write each functional group capable of being deprotonated as an acid, HA, and the deprotonated form as A. The charge of HA and A can be determined for the functional groups using the Henderson-Hasselbalch equation from Chapter 2.2.
\begin{equation}
\mathrm{pH}=\mathrm{p} K_{\mathrm{a}}+\log \frac{\left[\mathrm{A}^{-}\right]}{[\mathrm{HA}]}
\end{equation}
The titration curve for a single ionizable acid with different pKa values is shown below.
At the curve's inflection point, pH = pKa, the system is most resistant to changes in pH when either acid or base is added. At this pH, [HA]=[A-].
The properties of a protein will be determined partly by whether the side chain functional groups, the N-terminal, and the C-terminal are charged or not. The HH equation tells us that this will depend on the pH and the pKa of the functional group.
- If the pH is 2 units below the pKa, the HH equation becomes -2 = log A/HA, or .01 = A/HA. This means the functional group will be about 99% protonated (with either 0 or +1 charge, depending on the functional group).
- If the pH is 2 units above the pKa, the HH equation becomes 2 = log A/HA, or 100 = A/HA. Therefore, the functional group will be 99% deprotonated.
- If the pH = pKa, the HH equation becomes 0 = log A/HA or 1 = A/HA. Therefore, the functional group will be 50% deprotonated.
From these simple examples, we have derived the +2 rule. This rule is used to quickly determine protonation, and hence charge state, and is extremely important to know (and easy to derive). Titration curves for Gly (no ionizable side chain), Glu (carboxylic acid side chain), and Lys (amine side chain) are shown in Figure \(\PageIndex{8}\). You should be able to associate various sections of these curves with the titration of specific ionizable groups in the amino acids.

New 5/16/23: Download this Excel spreadsheet for Titration Curves for a Triprotic Acid. It has adjustable scroll bars to change pKa values.
Buffer Review
The Henderson-Hasselbalch equation is also useful in calculating the composition of buffer solutions. Remember that buffer solutions are composed of a weak acid and its conjugate base. Consider the equilibrium for a weak acid, like acetic acid, and its conjugate base, acetate:
\[\ce{CH3CO2H + H2O <=> H3O^{+} + CH3CO2^{-}} \nonumber \]
If the buffer solution contains equal concentrations of acetic acid and acetate, the pH of the solution is:
or pH = pKa + log [A]/[HA] = 4.7 + log 1 = 4.7
A look at the titration curve for the carboxyl group of Gly (see above) shows that when the pH = pKa, the slope of the curve (i.e. the change in pH on addition of base or acid) is at a minimum. As a general rule of thumb, buffer solutions can be made for a weak acid or base within ±1 pH units of its pKa. At pH = pKa, the buffer solution best resists the addition of either an acid or a base and has its greatest buffering capacity. The weak acid can react with the added strong base to form the weak conjugate base, and the conjugate base can react with the added strong acid to form the weak acid (as shown below), so pH changes with the addition of strong acid and base are minimized.
- addition of a strong base produces a weak conjugate base: CH3CO2H + OH- ↔ CH3CO2- + H2O
- addition of strong acid produces weak acid: H3O+ + CH3CO2 → CH3CO2H + H2O
There are two simple ways to make a buffered solution. Consider an acetic acid/acetate buffer solution.
- Make equal molar solutions of acetic acid and sodium acetate and mix them, monitoring pH with a pH meter, until the desired pH is reached (+/- 1 unit from the pKa).
- Add acetic acid solution to NaOH in stoichiometric amounts until the desired pH is reached (+/- 1 unit from the pKa). In this method, you are forming the conjugate base, acetate, with the addition of NaOH:
CH3CO2H + OH- → CH3CO2- + H2O
- Buffers for pH control: Recipes based on pKas for acids, temperature, and ionic strength
Isoelectric Point
What happens if you have many ionizable groups in a single molecule, as is the case with a polypeptide or a protein? Consider a protein. At a pH of 2, all ionizable groups would be protonated, and the overall charge of the protein would be positive. (Remember, when carboxylic acid side chains are protonated, their net charge is 0.) As pH increases, the most acidic groups will begin to deprotonate, and the net charge will become less positive. At high pH, all the ionizable groups will become deprotonated in the strong base, and the overall charge of the protein will be negative. At some pH, then, the net charge will be 0. This pH is called the isoelectric point (pI). The pI can be determined by averaging the pKa values of the two groups closest to and on either side of the pI. One of the online problems will address this in more detail.
Remember that pKa is a measure of the equilibrium constant for the reaction. And, of course, you remember that ΔGo = -RT ln Keq. Therefore, pKa is independent of concentration and depends only on the intrinsic stability of the reactants relative to the products. This is true only AT A GIVEN SET OF CONDITIONS, SUCH AS T, P, AND SOLVENT CONDITIONS.
Consider, for example, acetic acid, which in aqueous solution has a pKa of about 4.7. It is a weak acid, which dissociates only slightly to form H+ (in water, the hydronium ion, H3O+, is formed) and acetate (Ac-). These ions are moderately stable in water but reassociate readily to form the starting product. The pKa of acetic acid in 80% ethanol is 6.87. This can be accounted for by the decrease in stability of the charged products, which are less shielded from each other by the less polar ethanol. Ethanol has a lower dielectric constant than water. The pKa increases to 10.32 in 100% ethanol and a whopping 130 in air!
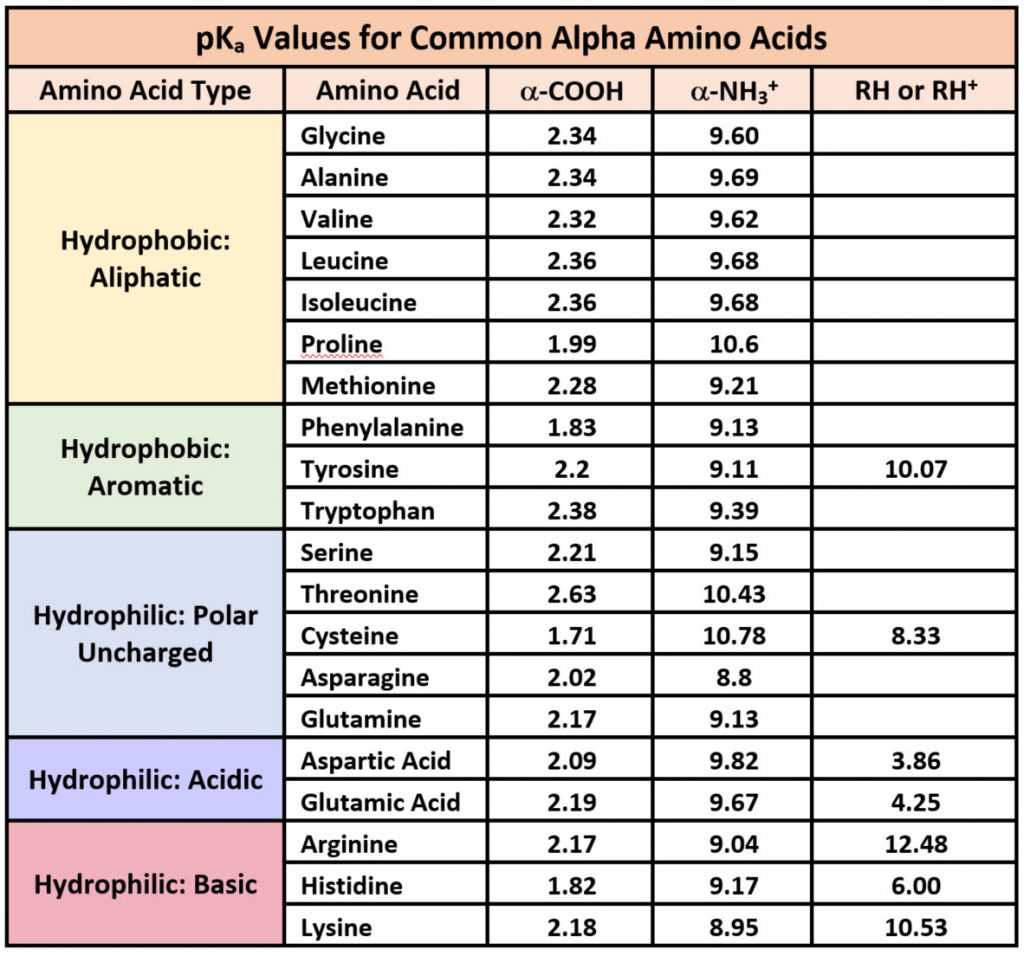
Because amino acids are zwitterions and several also contain ionizable R-groups, their charge state in vivo —and thus their reactivity —can vary with pH, temperature, and the local microenvironment's solvation status. Table \(\PageIndex{2}\) shows the standard pKa values for the amino acids and can be used to predict their ionization/charge status and their resulting peptides/proteins.
Table \(\PageIndex{2}\): Summary of pKas of amino acids
However, it should be noted that the solvation status in the microenvironment of an amino acid can alter the relative pKa values of these functional groups and provide unique reactive properties within the active sites of enzymes. A more in-depth discussion of the effects of desolvation will be given in Chapter 6, discussing enzyme reaction mechanisms.
- Printable Version of pKa Values
- Expasy pI and molecular weight calculator for any protein sequence
Introduction to Amino Acid Reactivity
You should be able to identify which side chains contain H-bond donors and acceptors. Likewise, some are acids and bases. You should know the approximate pKas of the side chains and the N and C-terminal groups. Three of the amino acid side chains (Trp, Tyr, and Phe) contribute significantly to the UV absorption of a protein at 280 nm. This section will primarily address the chemical reactivity of the side chains, which is important for understanding protein properties. Many of the side chains are nucleophiles. Nucleophilicity measures how rapidly molecules with lone pairs of electrons can react in nucleophilic substitution reactions. It correlates with basicity, which measures the extent to which a molecule with lone pairs can react with an acid (Bronsted or Lewis). The properties of the atom that holds the lone pair are important in determining both nucleophilicity and basicity. In both cases, the atom must be willing to share its unbonded electron pair. If the atoms holding the nonbonded pair are more electronegative, they will be less likely to share electrons, and that molecule will be a poorer nucleophile (nu:) and weaker base. Using these ideas, it should be clear that RNH2 is a better nucleophile than ROH, OH- is better than H2O, and RSH is better than H2O. In the latter case, S is larger, and its electron cloud is more polarizable—hence, it is more reactive. The important side chain nucleophiles (in order from most to least nucleophilic) are Cys (RSH, pKa 8.5-9.5), His (pKa 6-7), Lys (pKa 10.5), and Ser (ROH, pKa 13). The side chain of serine is generally no more reactive than ethanol. It is a potent nucleophile in a certain class of proteins (such as proteases) when deprotonated. The amino group of lysine is a potent nucleophile only when deprotonated.
An understanding of the chemical reactivity of the various R group side chains of the amino acids in a protein is important since chemical reagents that react specifically with a given amino acid side chain can be used to:
- Identify the presence of the amino acids in unknown proteins or
- Determine if a given amino acid is critical for the structure or function of the protein. For example, suppose a reagent that covalently interacts with only Lys is found to inhibit the protein's function. Lysine might be considered important for the protein's catalytic activity in that case.
Figure \(\PageIndex{9}\) summarizes nucleophilic addition and substitution at carbonyl carbons.
The rest of the section will summarize the chemistry of the side chains of reactive amino acids. Historically, the function of a given amino acid in a protein has been studied by reacting it with side chain-specific chemical modifying agents. In addition, some side chains are covalently modified after synthesis in vivo (post-translational modifications—see below).
Reactions of Lysine
Figure \(\PageIndex{10}\) shows the reaction of lysine with anhydrides and ethylacetimidate.
- reacts with anhydrides in a nucleophilic substitution reaction (acylation).
- reacts reversibly with methylmaleic anhydride (also called citraconic anhydride) in a nucleophilic substitution reaction.
- reacts with high specificity with ethylacetimidate in a nucleophilic substitution reaction (ethylacetimidate is like ethylacetate, only with an imido group replacing the carbonyl oxygen). Ethanol leaves as the amidino group forms. (has two N -i.e., din - attached to the C)
Figure \(\PageIndex{11}\) shows a second set of common reactions of lysine, including those used to attach a chromophore or a fluorescent label to the side chain.
- reacts with O-methylisourea in a nucleophilic substitution reaction with the expulsion of methanol to form a guanidino group (has 3 N attached to C, nidi)
- reacts with fluorodinitrobenzene (FDNB or Sanger's reagent) or trinitrobenzenesulfonate (TNBS, as we saw with the reaction with phosphatidylethanolamine) in a nucleophilic aromatic substitution reaction to form 2,4-DNP-lysine or TNB-lysine.
- reacts with dimethylaminonapthelenesulfonylchloride (Dansyl Chloride) in a nucleophilic substitution reaction.
Figure \(\PageIndex{12}\) shows a final common reaction we will encounter: the formation of an imine or Schiff base on the reaction of lysine with an aldehyde or ketone.
- reacts with high specificity toward aldehydes to form imines (Schiff bases), which can be reduced with sodium borohydride or cyanoborohydride to form a secondary amine.
Reactions of Cysteine
Cysteine is a potent nucleophile that often forms a covalent disulfide bond with another Cys.
Figure \(\PageIndex{13}\) shows common reagents used in the lab to label free Cys side chains. These reagents are used to alter Cys side chains to determine if they have functional significance in a protein (such as an active nucleophile in an enzyme-catalyzed reaction).
- reacts with iodoacetic acid in an SN2 reaction, adding a carboxymethyl group to the S.
- reacts with iodoacetamide in an SN2 reaction, adding a carboxyamidomethyl group to S.
- reacts with N-ethylmaleimide in an addition reaction to the double bond
Sulfur is directly below oxygen in the periodic table, and, in analogy to water, sulfur-containing amino acids are found in different redox states, as illustrated in Figure \(\PageIndex{14}\).
Cystine Chemistry
Two cysteine side chains can covalently interact in a protein to form a disulfide bond (RS-SR), called cystine. Just as HOOH (hydrogen peroxide) is more oxidized than HOH (O in H2O2 has an oxidation number of 1- while the O in H2O has an oxidation number of -2), RSSR is the oxidized form (S oxidation number -1), and RSH is the reduced form (S oxidation number -2) of thiols. Their oxidation numbers are analogous since O and S are both in Group 6 of the periodic table and are more electronegative than C.
Cystine can react with a free sulfhydryl (RSH) in a thermodynamically non-challenging disulfide exchange reaction, which, when conducted with excess free sulfhydryls, results in the reduction of cystine in the protein, as shown in Figure \(\PageIndex{15}\).
Figure \(\PageIndex{15}\): Disulfide interchange and reduction of protein disulfides
This reaction is often used in the lab to quantify the amount of free cysteine side chains in a protein using Ellman's reagent, as shown in Figure \(\PageIndex{16}\).
The 2-nitro-5-thiobennzoate anion leaving group absorbs at 412 nm, making quantitation easy. However, unless the protein is unfolded to expose all the cysteines, only surface-exposed, not buried, free cysteines will be labeled.
When a protein folds, two Cys side chains might approach each other and form an intra-chain disulfide bond. Likewise, two Cys side chains on separate proteins might approach each other and form an inter-chain disulfide. For protein structure analysis, disulfides are typically cleaved, and the chains are separated. The disulfides can be cleaved by reducing agents such as beta-mercaptoethanol, dithiothreitol, or tris(2-carboxyethyl)phosphine (TCEP), or by oxidizing agents such as performic acid, which further oxidizes the disulfide to separate cysteic acids. Three common reagents used in disulfide cleavage reactions in the lab are shown in Figure \(\PageIndex{17}\).
The reaction for beta-mercaptoethanol (BME) and performic acid is shown in Figure \(\PageIndex{18}\) below.
Figure \(\PageIndex{19}\) shows the reaction for dithiothreitol (DTT). Note that it forms a stable cyclohexane-like ring, which thermodynamically favors this reaction. It does not require as much excess DTT as the reaction with BME.
The reaction with tris (2-carboxyethyl) phosphine (TCEP) is not a disulfide interchange reaction, as is shown in Figure \(\PageIndex{20}\).
Cells maintain a reducing environment using many "reducing" agents, such as the tripeptide gamma-Glu-Cys-Gly (glutathione). Hence, intracellular proteins usually do not contain disulfides, which are abundant in extracellular proteins (such as those found in blood) or in certain organelles, such as the endoplasmic reticulum and mitochondrial intermembrane space, where disulfides can be introduced.
Sulfur redox chemistry is very important biologically. As described above, the sulfur in cysteine is redox-active. Hence, it can exist in various states, depending on the local redox environment and the presence of oxidizing and reducing agents. A potent oxidizing agent produced in cells is hydrogen peroxide, which can lead to more drastic, irreversible chemical modifications of Cys side chains. Suppose a reactive Cys is important to protein function. In that case, the function of the protein can be modulated (sometimes reversibly, sometimes irreversibly) with various oxidizing agents, as shown in Figure \(\PageIndex{21}\).
Reactions of Histidine
Histidine is one of the most important bases at physiological pH. Remember from introductory chemistry that for any acid/conjugate base pair, the pKa and pKb of the acid and base are related by this expression:
pKa + pKb = 14
Table \(\PageIndex{3}\) below shows the pKa and pKb of three amino acids.
| amino acid | pKa | pKb |
|---|---|---|
| Histidine | 6.5 | 7.5 |
| Lysine | 10.5 | 3.5 |
| Arginine | 12.5 | 1.5 |
The deprotonated forms of Lys and Arg with lower pKbs are much stronger bases than the deprotonated form of His, so at physiological pH, they would always be protonated (unless their local environment lowers their pKa and pKb values). In contrast, His exists in both protonated and deprotonated states at physiological pH, so it can readily gain a proton and act as a general base in reactions.
Histidine can exist as two tautomers, as shown in Figure \(\PageIndex{22}\). NMR studies show that in model peptides, the proton is predominantly on the ε2, N3, or tele N in the imidazole ring, as it has a pKa 0.6 units higher than δ1, N1, or pro N.
The nitrogen atom in a secondary amine might be expected to be a stronger nucleophile than a primary amine through electron release to that N in a secondary amine. Opposing this effect is the steric hindrance by the two attached Cs of the N on attachment to an electrophile. However, in His, this steric effect is minimized since the ring restrains the 2Cs. With a pKa of about 6.5, this amino acid is one of the strongest available bases at physiological pH (7.0). Hence, it can often cross-react with many reagents used to modify Lys side chains. Histidine reacts with reasonably high selectivity with diethylpyrocarbonate as shown in Figure \(\PageIndex{23}\).
In vivo Post-Translational Modification of Amino Acids
Amino acids in naturally occurring proteins are also subjected to chemical modifications within cells. These modifications alter the properties of the amino acid, thereby affecting the protein's structure and function. Most chemical modifications of proteins within cells occur after they are synthesized through a process called translation. The resulting chemical changes are termed post-translational modifications. Several are shown in Figure \(\PageIndex{24}\). Simple acid/base reactions are included, but these are not considered examples of post-translational modifications.
There are 100s of PTMs, and many are part of an elaborate system within a cell to respond to both external (hormones, neurotransmitters, nutrients, metabolites) and internal chemical signals. PTMs (such as phosphorylation and acetylation) and their removal by enzymes are part of an elaborate cell signaling system that we will explore in great detail in Chapter 28. However, not all PTMs are benign. Examples include glycation, oxidation, citrullination, and carbonylation of protein side chains. These are often increased during periods of inflammatory stress (both acute and chronic). These modified proteins are degraded within the cell to short peptides that retain the chemical modification. Unfortunately, these can be recognized by the immune system as foreign, triggering an immune response against the body's own tissues and leading to autoimmune disease. One potentially deleterious PTM is the carboxyethylation of cysteine, catalyzed by the enzyme cystathionine β-synthase as shown in Figure \(\PageIndex{25}\) below.
Figure \(\PageIndex{25}\): Carboxyethylation of cysteine
The product is very similar to the carboxymethylation of cysteine shown in Figure 13 above. The modifying reagent, 3-hydroxypropionic acid, is a metabolite produced by gut microbes. This modification has been shown to produce an autoimmune response in the disease ankylosing spondylitis.
Summary
This chapter provides an extensive overview of proteins, their building blocks, and the chemical principles that govern their structure and function. It begins by emphasizing that proteins are not only among the most abundant macromolecules in living systems, but also the most functionally diverse. Proteins serve many roles—including structural support, catalysis, regulation, transport, and defense—and their diversity arises from the linear polymerization of alpha-amino acids linked via peptide bonds.
Key topics include:
-
Alpha-Amino Acid Structure and Protein Primary Structure:
Every protein is composed of 20 naturally occurring alpha-amino acids, each sharing a common backbone (an α-amino group, an α-carboxyl group, and a hydrogen atom) and a unique R-group that defines its chemical properties. The chapter clarifies the process of peptide bond formation (and hydrolysis) through nucleophilic attack, and it dispels common misconceptions about the free nature of the terminal groups in a protein chain. -
Amino Acid Side Chain Properties:
The diverse chemical nature of proteins is largely due to the variability of the amino acid side chains, which can be classified as nonpolar (hydrophobic), polar uncharged, acidic, or basic. These classifications are critical for understanding protein folding, stability, and function. The chapter introduces hydropathy scales, such as the Kyte-Doolittle and Hopp-Woods scales, to quantitatively assess the relative hydrophobicity or hydrophilicity of side chains, providing insight into protein topology. -
Stereochemistry and Chirality:
Except for glycine, all amino acids are chiral and naturally occur as L isomers in proteins. The chapter contrasts the D/L nomenclature with the R/S system and explains why biochemists favor the former, particularly when describing the stereochemistry of amino acids in the context of protein structure. -
Acid-Base Chemistry and Ionization:
The ionizable groups on amino acids, including the α-amino, α-carboxyl, and various side chains, determine the overall charge and reactivity of proteins. The Henderson–Hasselbalch equation is introduced as a tool to predict the protonation state of these groups under varying pH conditions, leading to the concept of the isoelectric point (pI) — the pH at which a protein has no net charge. -
Chemical Reactivity of Side Chains:
Beyond structural roles, the chemical reactivity of amino acid side chains is central to protein function. The chapter details how key side chains—such as those of lysine, cysteine, and histidine—participate in nucleophilic reactions, form covalent modifications (e.g., Schiff bases), and are targeted by specific chemical reagents. This reactivity is foundational for both enzyme catalysis and experimental methods used to probe protein structure and function. -
Post-Translational Modifications (PTMs):
The chapter concludes by discussing PTMs, chemical modifications that occur after protein synthesis. These modifications, including phosphorylation, acetylation, glycosylation, and oxidation, can dramatically alter a protein's activity, stability, and interactions. While many PTMs are critical for normal cellular signaling, aberrant modifications may contribute to disease.
Overall, the chapter integrates structural biology with chemical reactivity, providing a comprehensive framework that links the primary structure of proteins to their three-dimensional architecture and dynamic functions in the cell. This foundation is essential for understanding advanced topics in enzymology, protein engineering, and molecular regulation.