1.5: Gene Expression- Transcription
- Page ID
- 73670
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the roles that the promoter, coding region and untranslated regions of a gene play in gene expression.
- Describe mRNA processing steps.
- Draw the process of transcription and include the following in your drawing. DNA template and non-template strands, RNA polymerase, new RNA strand, and direction of RNA synthesis.
- Draw the process of mRNA processing and include the following in your diagram, Gene (DNA), promoter, coding region, introns, exons, pre-mRNA, mature mRNA, poly A tail, cap.
Introduction
Genes are DNA sequences that control traits in an organism by coding for proteins (Figure 1). Organisms such as plants and animals have tens of thousands of genes. The impact that a single gene’s information can have on an organism, however, is tremendous. Furthermore, organisms have all their genes in each of their cells, but they only need to use the information from a subset of these genes, depending on the type of cell and the cell’s stage of development. Therefore, the key to gene function is controlling its expression.
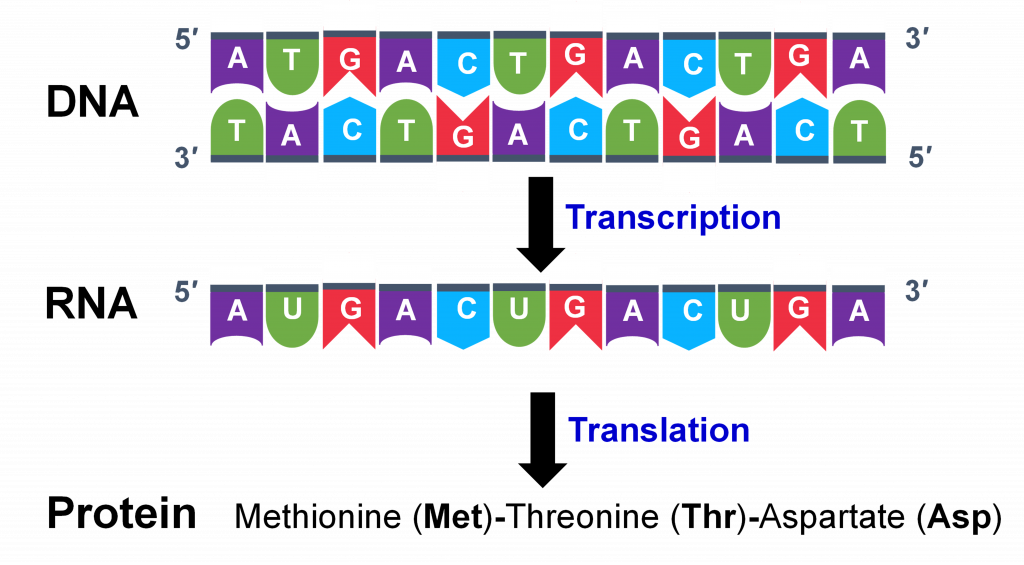
Gene structure and transcription
The DNA sequence contains the information to control all biological functions, including the manifestation of traits important to agriculture (yield, drought tolerance, disease resistance, etc., etc.) How is the information contained in DNA sequences converted into the cellular activities necessary for plants and other organisms to function? DNA sequences are used to direct the synthesis of other molecules that actually perform these cellular functions.
Most typical genes encode proteins. The production of a protein from a gene involves several different processes (Figure 1). Transcription involves the copying of the DNA nucleotide sequence into an intermediate nucleotide molecule called RNA (ribonucleic acid). The primary RNA molecule is processed into a mature messenger RNA (mRNA) which then provides the information for the synthesis of a protein through the process of translation. Proteins are composed of amino acids connected by peptide bonds. The sequence of amino acids is determined by the sequence of nucleotide bases in the mRNA. The 20 amino acids have different chemical and physical properties and the sequence of amino acids determine the structure and function of the protein.
Gene structure
Only particular regions of chromosomal DNA are transcribed. A gene can be considered as the region of transcribed DNA, along with associated regions of DNA important for the regulation of transcription (Figure 1). A gene has several parts that are each important to the function of the gene.
The regulatory region (also known as the promoter) contains DNA sequence involved in the control of where and when the genes will be turned on to produce mRNA. The coding region is the part of the gene that is used as template to produce RNA molecules in a process called transcription. Some RNA molecules perform cellular functions directly while many others (messenger RNAs) are used to direct the synthesis of proteins in a process called translation.
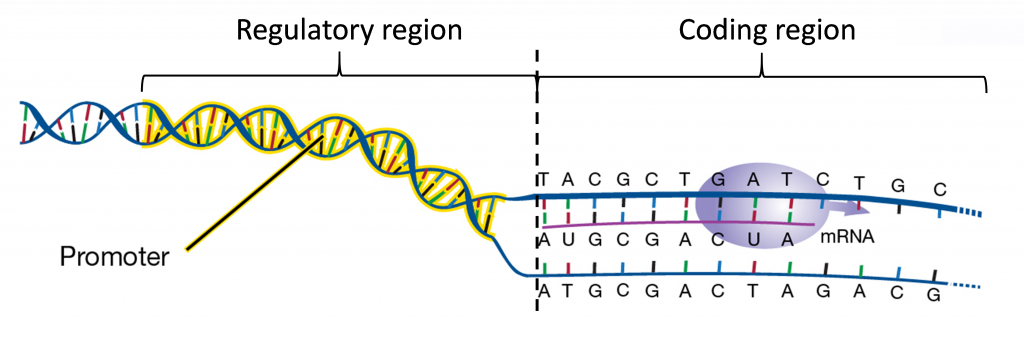
a. Gene Promoter
The signals for starting and stopping transcription are located within DNA sequences. Specific nucleotide segments called promoters are recognized by RNA polymerase to start RNA synthesis. After the transcription of full-length RNA strand is completed, a second segment of DNA called terminator invokes termination of RNA synthesis and the detachment of RNA polymerases from the DNA template.
b. Protein-coding region
The protein-coding region of a gene is composed of the sequence of nucleotides that codes for amino acids. As described further in the section on translation, the coding region begins with an ATG start codon (AUG in RNA) and then ends with one of three stop codons. These sequences include only exons, but not all exonic sequences are protein coding as they may include untranslated regions.

c. Untranslated regions (UTRs)
Mature transcripts contain some sequences that do not code for amino acid sequences in proteins. These are referred to as untranslated regions or UTRs. Most mRNA transcripts contain a 5′ and a 3′ UTR. The 5′ UTR contains sequences toward the 5′ end of the mRNA sequence, before the start codon. These sequences can often be important for translational regulation, and sometimes other functions. The sequences following the stop codon are the 3′ UTR. The 3′ UTR may also have important functions regulating transcript stability or directing transcript localization within cells, or sometimes even transport (trafficking) between cells.
UTRs and introns are often useful in genetic studies. Protein coding regions are under strong selective pressure to produce functional proteins and so sequence variation is relatively rare. UTRs and introns on the other hand are under less stringent selection and are therefore sources of sequence variants that can be used to develop genetic markers.
Transcription
The genetic information of DNA is transferred to an intermediate molecule called RNA that is often translated to amino acid sequences used to build proteins. RNA is a nucleic acid, like DNA, but with some important differences. RNA contains a ribose sugar group instead of the deoxyribose found in DNA. RNA molecules are single stranded, instead of being double stranded. RNA contains a uridine (U) base and does not contain a thymidine base. The other bases (A, C, G) are contained in both RNA and DNA. U has the property of base-pairing with A.
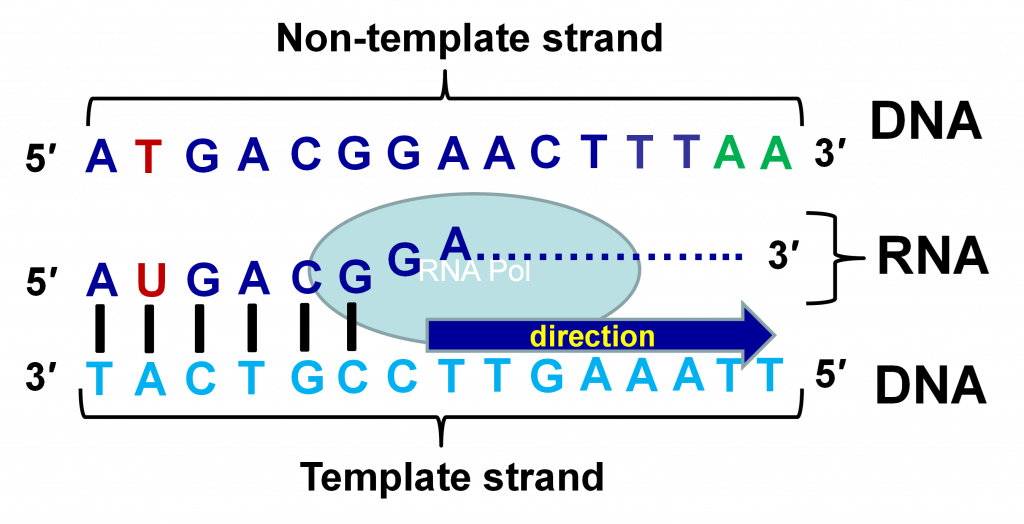
RNA synthesis is directed by a DNA template in a process called transcription. A protein complex containing the enzyme RNA polymerase synthesizes an RNA molecule by adding nucleosides to the 3′ end of a growing chain. The principle of base pairing is used again and each nucleotide base added is complementary to the corresponding base on the DNA template. Thus, the RNA is complementary in sequence to the template strand of DNA, which is also referred to as “antisense” or “negative” strand (Figure 4). The RNA is identical in sequence (except U replaces T) to the other strand, which is called the “sense” or “positive” strand. Because RNA molecules are produced by the process of transcription, they are often referred to as transcripts.
RNA processing
Coding (transcribed) region
This is the region that is transcribed by RNA polymerase, also known as the RNA coding region. As described below, it may include introns, sequences that are removed from the mature RNA molecule during RNA processing. The transcribed region is demarcated by promoter and terminator sequences.
Introns and exons
As mentioned, and described in detail below, introns are sequences that are removed from transcripts during RNA processing. Sequences that are retained in mature transcripts are called exons. The corresponding stretches of DNA are typically referred to with the same terms. Introns are commonly found in genes of eukaryotes but are rare in prokaryotic organisms.
Intron splicing
The process of transcription produces pre-mRNA that contains both introns and exons. The process of splicing involves removal of introns from pre-mRNA and joining together the exons. A complex group of proteins that form a spliceosome perform the splicing reaction.
Introns sometimes serve as boundaries for sequences encoding functional protein domains, leading to possibility for new and variant proteins by exon shuffling. Also, introns can provide possibility for productions of variant RNA forms through alternate splicing allowing more than one gene product from a single gene. Some introns result from the insertion of transposable elements and may be spliced perfectly of imperfectly, offering more possibility for new genetic diversity.
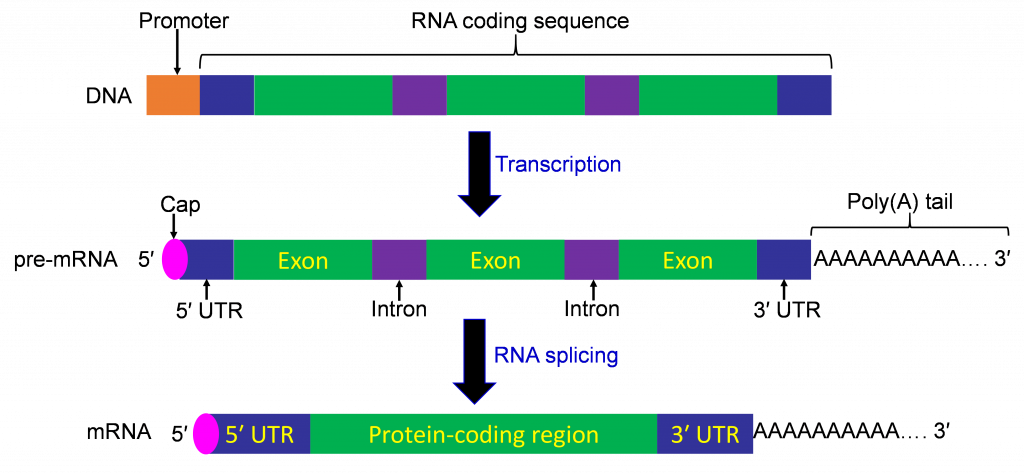
5′ Capping
The 5’ capping is the addition of a 7-methyl guanidine to the first nucleotide of mRNA molecule, usually and adenine or guanidine. The phosphodiester linkage between 7-methyl guanidine and the target nucleotide is 5′-5′ instead of 5′-3′, and 3 phosphates rather than 1 are retained in the linkage. The cap stabilizes the 5′ end of the mRNA and plays a role in translation initiation.
Poly adenylation
The transcription of a gene may proceed beyond what ends up as 3′ end of mature mRNA. Thus the 3′ end of mRNA is formed after transcription. The enzyme poly(A) polymerase adds numerous adenosines to the 3′ end to result in what is called the poly(A) tail. The poly(A) tail is necessary for proper processing and transport of mRNA to the cytoplasm. The poly(A) tail is also important for the stability of mRNA, and initiation of translation in eukaryotic organisms.
The genetic information of DNA is transferred through transcription to an intermediate molecule called RNA. The signals for starting and stopping transcription are located within the DNA sequence and referred to as promoter and terminator sequences. The coding region of a gene is composed of a sequence of nucleotides that are transcribed into RNA. These sequences include exons and introns. Exons are the sequences that code for proteins. The coding region of a gene contains exons and introns. Also, pre-mRNA contains both introns and exons. The introns in pre-mRNA are removed through a process called intron splicing. The mRNA is processed by 5′ capping and addition of a poly(A) tail.
Learning Activities
Given the following sequence of double-stranded DNA, predict the sequence of the RNA strand.
Watch This Video: Transcription Detail
A video element has been excluded from this version of the text. You can watch it online here: https://iastate.pressbooks.pub/genagbiotech/?p=24


