11.7: Restriction Fragment Length Polymorphisms
- Page ID
- 4928
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Restriction enzymes cut DNA at precise points producing a collection of DNA fragments of precisely defined length. These can be separated by electrophoresis, with the smaller fragments migrating farther than the larger fragments. One or more of the fragments can be visualized with a "probe" — a molecule of single-stranded DNA that is complementary to a run of nucleotides in one or more of the restriction fragments and is radioactive (or fluorescent). If probes encounter a complementary sequence of nucleotides in a test sample of DNA, they bind to it by Watson-Crick base pairing and thus identify it. Polymorphisms are inherited differences found among the individuals in a population.
Restriction Fragment Length Polymorphisms (RFLPs) have provided valuable information in many areas of biology, including screening human DNA for the presence of potentially deleterious genes ("Case 1") and providing evidence to establish the innocence of, or a probability of the guilt of, a crime suspect by DNA "fingerprinting" ("Case 3").
Case 1: Screening for the sickle-cell gene
Sickle-cell disease is a genetic disorder in which both genes in the patient encode the amino acid valine (Val) in the sixth position of the beta chain (betaS) of the hemoglobin molecule. "Normal" beta chains (betaA) have glutamic acid at this position. The only difference between the two genes is the substitution of a T for an A in the middle position of codon 6. This converts a GAG codon (for Glu) to a GTG codon for Val and abolishes a sequence (CTGAGG, which spans codons 5, 6, and 7) recognized and cut by one of the restriction enzymes.
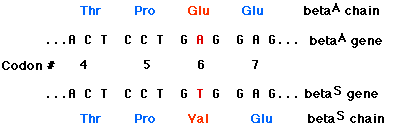
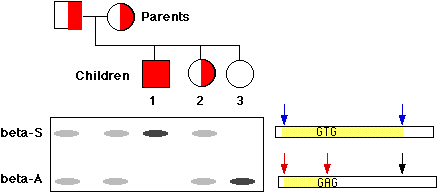
When the normal gene (betaA) is digested with the enzyme and the fragments separated by electrophoresis, the probe binds to a short fragment (between the red arrows). However, the enzyme cannot cut the sickle-cell gene at this site, so the probe attaches to a much larger fragment (between the blue arrows).
Figure \(\PageIndex{1}\) shows the pedigree of a family whose only son has sickle-cell disease. Both his father and mother were heterozygous (semifilled box and circle respectively) as they had to be to produce an afflicted child (solid box). The electrophoresis patterns for each member of the family are placed directly beneath them. Note that the two homozygous children (1 and 3) have only a single band, but these are more intense because there is twice as much DNA in them. In this example, a change of a single nucleotide produced the RFLP. This is a very common cause of RFLPs and now such polymorphisms are often referred to as single nucleotide polymorphisms or SNPs. (However, not all RFLPs arise from SNPs.
By testing the DNA of prospective parents, their genotype can be determined and their odds of producing an afflicted child can be determined. In the case of sickle-cell disease, if both parents are heterozygous for the genes, there is a 1 in 4 chance that they will produce a child with the disease. Amniocentesis and chorionic villus sampling make it possible to apply the same techniques to the DNA of a fetus early in pregnancy. The parents can learn whether the unborn child will be free of the disease or not. They may choose to have an abortion rather than bring an afflicted child into the world.
Three problems:
- The mutations that cause most human genetic diseases are more varied than the single mutation associated with sickle-cell disease. Over a thousand different mutations in the cystic fibrosis gene can cause the disease. A probe for one will probably fail to identify a second. A mixture of probes, one for each of the more common mutations, can be used. But there remains the problem of "false negatives": people who are falsely told they do not carry a mutant gene.
- There are many diseases which result from several mutant genes working together to produce the disease phenotype.
- There are still genetic diseases for which no gene has yet been discovered. Until the gene can be located, cloned, and sequenced, no probe can be made to detect it directly. However, it is sometimes possible to find a genetic "marker" that can serve as a surrogate for the gene itself. Let's see how.
Case 2: Screening for a RFLP "marker"
If a particular RFLP is usually associated with a particular genetic disease, then the presence or absence of that RFLP can be used to counsel people about their risk of developing or transmitting the disease. The assumption is that the gene they are really interested in is located so close to the RFLP that the presence of the RFLP can serve as a surrogate for the disease gene itself. But people wanting to be tested cannot simply walk in off the street. Because of crossing over, a particular RFLP might be associated with the mutant gene in some people, with its healthy allele in others. Thus it is essential to examine not only the patient but as many members of the patient's family as possible.
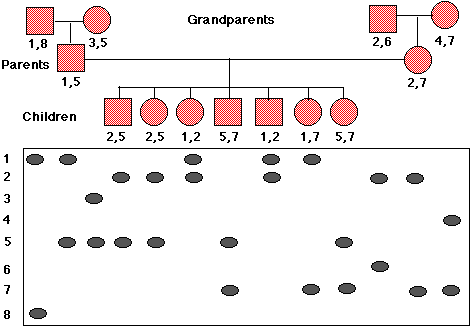
The most useful probes for such analysis bind to a unique sequence of DNA; that is, a sequence occurring at only one place in the genome. Often this DNA is of unknown, if any, function. This can actually be helpful as this DNA has been freer to mutate without harm to the owner. The probe will hybridize (bind to) different lengths of digested DNA in different people depending on where the enzyme cutting sites are that each person has inherited. Thus a large variety of alleles (polymorphisms) may be present in the population. Some people will be homozygous and reveal a single band; others (e.g., all the family members shown below) will be heterozygous with each allele producing its band.
The pedigree in Figure \(\PageIndex{2}\) shows the inheritance of a RFLP marker through three generations in a single family. A total of 8 alleles (numbered to the left of the blots) are present in the family. The RFLPs of each member of the family are placed directly below his (squares) or her (circles) symbol and RFLP numbers.
If, for example, everyone who inherited RFLP 2 also has a certain inherited disorder, and no one lacking RFLP 2 has the disorder, we deduce that the gene for the disease is closely linked to this RFLP. If the parents decide to have another child, prenatal testing could reveal whether that child was apt to come down with the disease. But note, that crossing over during gamete formation could have moved the RFLP to the healthy allele. So the greater the distance between the RFLP and the gene locus, the lower the probability of an accurate diagnosis.
Case 3: DNA "typing"
Each human cell contains 6 x 109 base pairs of DNA. Some of this represents protein-encoding genes (e.g., for the beta chain of hemoglobin) that are identical in a large proportion of people. But long stretches of DNA do not encode for anything and are free to mutate extensively. It seems certain that if we could read the entire sequence of DNA in each human, we would never find two that were identical (unless the samples were from identical siblings; i.e., derived from a single zygote).
So each person's DNA is as unique as a fingerprint. This truth has not escaped the law enforcement and legal professions. Analysis of DNA, called DNA typing, is widely used to identify rapists and other criminals, determine paternity; that is, who the father of the child really is, determine whether a hopeful immigrant is, as he or she claims, really a close relative of already established residents.
Figure \(\PageIndex{3}\) shows the test results in a rape case. Two probes were used: one revealing the bands at the top, the other those at the bottom.
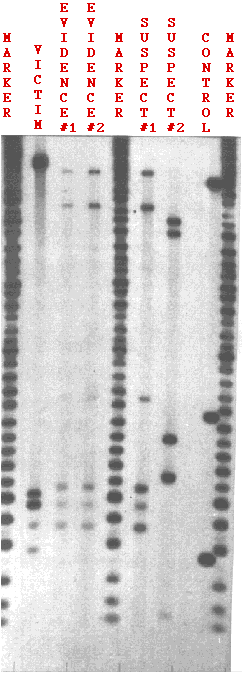 3
3DNA was tested from
- semen removed from the vagina of the rape victim (EVIDENCE #2);
- a semen stain left on the victim's clothing (EVIDENCE #1);
- the DNA of the victim herself (VICTIM) to be sure that the DNA didn't come from her cells;
- DNA from two suspects (SUSPECT #1, SUSPECT #2);
- a set of DNA fragments of known and decreasing length (MARKER). They provide a built-in ruler for measuring the exact distance that each fragment travels.
- the cells of a previously-tested person to be sure the probes are performing properly (CONTROL).
One the basis of this test, suspect #2 can clearly be ruled out. None of his bands matches the bands found in the semen.
Is suspect #1 guilty?
We can never be certain. The best we can do is to estimate the probability that another person, picked at random, could provide the same DNA fingerprint. As a conservative estimate, a given allele (band) might be found in 25% of the people tested. The probability of a random match of two alleles is (0.25)2 or 1 in 16. The probability that 6 alleles match, as in this case, is (0.25)6 or 1 in 4096. However, the suspect was not picked at random, so you may feel that the evidence of guilt is strong.
The more probes you use, the more confident you can be that you have gotten the right man. If, for example, a set of probes revealed 14 bands in a suspect's DNA identical to those in the semen sample, the probability that you have the wrong man drops to less than 1 in 268 million (0.25)14 = 1/268,435,456, which is almost as great as the entire population, males and females, in the United States.
Starting in 1999, law enforcement agencies in both Great Britain and the United States began switching to a new version of RFLP analysis using shorter sequences called STRs ("Short Tandem Repeats"). STRs are repeated sequences of a few (usually four) nucleotides, e.g., TCATTCATTCATTCAT. They often occur in the untranslated parts of known genes (whose sequence can be used for the PCR primers). The exact number of repeats (6, 7, 8, 9, etc.) varies in different people (and, often, in the gene on each chromosome; that is, people are often heterozygous for the marker).
When 13 STR loci — scattered over different chromosomes — are examined, the chance that two people picked at random have the same pattern is less than 1 in 1 trillion. The U.S. Federal Bureau of Investigation (FBI) wants to increase the number of loci examined to 20 further eliminating the possibility of false positives.


