22A: Identification of Staphylococcus Species
- Page ID
- 3629
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Objectives
- Become familiar with the speciation of the genus Staphylococcus.
- Grow and identify different staphylococci species using selective and differential agar
- Identify the 3 hemolytic types on blood agar.
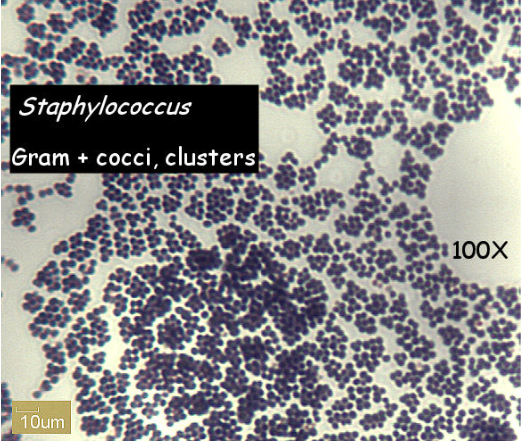
Staphylococcus is a genus of Gram +, nonspore-forming cocci belonging to the family Micrococcaceae that are often found as normal human microbiota of the skin and nasal cavity. There are five organisms to consider as potential human pathogens in this genus: S. aureus, S. epidermidis, S. saprophiticus, S. haemolyticus, and S. hominis but the first three are the most common isolates. S.aureus is often considered to be the most problematic of the three pathogens and is distinguished from the other two by being the only one able to coagulate plasma. S. aureusis able to cause many superficial pyogenic (pus-forming) infections of the dermis and underlying tissues as well as serious systemic infections. It can produce a range of toxins including enterotoxins (food poisoning), cytotoxins (general systemic toxins), and toxic shock superantigens. The other coagulase-negative staphylococci (S. epidermidis and S. saprophiticus) are much less frequently found as pathogens but are occasionally associated with endocarditis, prosthetic joint infections,and wound infections, just to name a few.
This exercise gives you the opportunity to use selective media, in this case based on high sodium chloride (MSA and SM1 10 are both selective media for the isolation of Staphylococci- 7.5% NaCl). A selective medium has an inhibitory agent which favors the growth of certain bacteria by inhibiting others. MSA contains an additional indicator for monitoring mannitol fermentation, which makes it a differential media also. Of the bacteria which can grow in the presence of high NaCl, some are halophilic (requiring a certain concentration of salt to grow) while other are haloduric (do not use the salt, but can tolerate it). Staphylococcusis not halophilic, but rather haloduric, in that it can live in or endure high NaCl concentrations. The high salt content in SM1 10 and MSA inhibits other common skin microorganisms. The other media being used in this exercise are for differentiating pathogenic Staphylococcus from nonpathogenic, and for identification of the species.
Not only salt resistant, Staphylococcus is always facultatively anaerobic. When stained, it will be seen in small clusters (staphylo = cluster). Staphylococcus is usually either beta hemolytic or not hemolytic at all (called gamma hemolysis). Pathogenic Staphylococci can produce a variety of virulence factors, including toxins,coagulase, leucocidins, and hydrolytic enzymes that can damage host tissues.
Blood agar (BAP) is made with 5% sheep blood. It is a common medium used to culture bacteria because:
- It is a great enrichment medium for fastidious bacteria.
- Hemolysis of blood cells can be very useful as an identification test.
CNA agar is a type of blood agar. The only difference is that CNA has an antibiotic, naladixic acid, that inhibits gram - bacteria.
Hemolysis is the breakdown of red blood cells. Hemolysins are enzymes produced by some bacteria and are released into the medium around the bacterial colony. It can be a complete breakdown of the cells, with the release of hemoglobin and a clearing of the red from the surrounding medium around the colony. Or the hemolysis can be a partial breakdown, resulting in a greenish or green-yellow zone around the colony.

MATERIALS NEEDED: per table
 Sterile swabs
Sterile swabs- Sterile saline solution
- 1 Mannitol salt agar (MSA) plates
- Containers of alcohol + forceps
- 1 Columbia naladixic acid blood agar plate (CNA)
- 1 Staphylococcus medium 110 (SM110) agar plate
- 1 DNase agar plates
- Novobiocin (5 microgram) antibiotic discs
- Metric rulers
- Rabbit plasma (frozen) for coagulase test
- Other media for identification:
- arginine and ornithine decarboxylase broths
- urea broth
- ONPG
- sucrose, trehalose, maltose
- VP broth
- nitrate broth
- thioglycollate broth (for anaerobic growth)
SCHEMATIC OF IDENTIFICATION PROCEDURE
1st session ⇒ 2nd session ⇒ 3rd session ⇒ 4th session
CNA DNAse Coagulase Run other needed tests Identify species
MSA SM110 ONPG
Catalase Oxidase
Gram Phenol red lactose
Stain Broth
THE PROCEDURES
Note
Be sure to keep a list of all test results for your isolates.
1st Session
- Each table will be given a Staphylococcal species growing on aNA or TSA plate.
- Using an isolation streak technique, inoculate the Columbia naladixic acid (CNA) and place the novobiocin disc in section 1.
- Novobiocin sensitivity is a key differentiating features among some of the Staphylococcus species. Place forceps into the alcohol and then sterilize the forceps by running them through the flame quickly. The alcohol will catch on fire and when evaporated from the forceps, they will then be sterile. You may now pick a novobiocin disc from the holder to place on the CNA plate—in the FIRST streak section (where there will be confluent growth of the bacterium).
- For the other agar plates--SM 110 plate,mannitol salt agar (MSA) plate, DNAse agar plate—an inoculation line down the center of the plate is adequate for growth results.
- Incubate media at 37°C.
- Run oxidase and catalase tests on plate culture.
- Gram stain the isolate to get shape and arrangement as well as gram reaction.
- We will be doing the ONPG test the next session. That test requires the turning on of a set of genes, the lac operon. To induce this activity, the bacterium has to be exposed to lactose. This is why you will also inoculate a tube of phenol red lactose broth.
2nd Session
- Go to individual lab exercises to interpret the mannitol salt and DNAse test (requires HCl).
- On the CNA plate, you are checking for 2 reactions---sensitivity to novobiocin antibiotic AND hemolysis of blood.
- Novobiocin sensitivity - A zone of 17 mm indicates sensitivity
- Hemolytic reactions :
- alpha (α) hemolysis–green zone around colony, caused by leaking hemoglobin converted to biliverdin, called a partial hemolysis
- beta (β) hemolysis–complete clearing around colony caused by breakdown of RBCs by streptolysin enzymes
- gamma (γ) hemolysis - no hemolysins, no zone
- Staphylococcus species are either beta hemolytic or gamma (not hemolytic). Staph aureus produces alpha toxin which typically causes wide zones of beta (complete) hemolysis.
- Check the SM 110 for growth and for pigment. Nutrients and vitamins in this medium enhance the pigmentation of the pathogenic Staphylococcus, those colonies becoming ayellow-orange colony.
- Run the coagulase test: there is a linked exercise for this test. There are only a few species of Staphylococcus that are positive for the coagulase test (see table below), and S. aureus is the most common. Since there are 2 kinds of coagulase enzyme—bound and free---there are 2 different tests that can be used to identify these enzymes. The TUBE method is the definitive test of the 2, the most reliable. The coagulase test exercise in the lab manual clearly describes these 2 versions.
- The ONPG test will be run. Look in your lab exercises for this test for the directions on how to run the test. Since there is very little liquid in the tube, you will want to cover the top of the tube will parafilm and then place the tube top over that. This reduces dehydration.
3rd Session
- The major test reaction to use in Staphylococcus identification is the coagulase test reaction, which divides the genus Staphylococcus into 2 groups—coagulase negative species and coagulase positive species. The test media that you will run for identification depends on which category your organism falls in. You may want to run some of the following tests.
- Available media:
- arginine and ornithine
- decarboxylase broths
- urea broth
- ONPG
- sucrose, trehalose, maltose sugars
- VP broth
- nitrate broth
- If you have a coagulase positive Staph, there are only a few species available as your choice: there is a table with those species below.
- If your organisms is coagulase negative, check the flow chart below, as well as the Bergey’s Manual or other identification books.
- Run necessary tests for identification. IT IS UP TO YOU TO DETERMINE WHAT EXTRA TESTS TO RUN FOR A CLEAR IDENTIFICATION.
4th Session
- Read any tests from last session (see lab exercises for information on test interpretations.
- Write up a report for your Staphylococcus species identification, if your instructor directs you to do so.
| Tube coagulase | Acid from trehalose | VP test | ONPG test | |
| S. aureus | + | + | + | - |
| S. intermedius | + | + | - | + |
| S. hyicus | variable | + | - | - |
| S. schleiferi | + | - | + | ? |

Common coagulase negative Staphylococcal species
| S. aureus | S. epidermidis | S. saprophiticus | |
| Alpha toxin (ß-hemolysis) | + (most strains) | - | - |
| Growth on 7.5% salt (mannitol salt) | |||
| Acid from mannitol | + | - | + (most strains) |
| Coagulase reaction | + | - | - |
| Pigment production (SM110) | Usually golden | Usually white | Usually white |
| DNase production | + (usually) | - | - |
| Sensitivity to novobiocin | sensitive | sensitive | resistant |
* Novobiocin sensitivity = >17mm zone size
QUESTIONS
- Which type of hemolysis is often associated with pathogenicity?
- What are the distinguishing features of the genus Staphylococcus?
- Can you give the test reaction of S. aureus for each of the major tests run---MSA, SM110, coagulase, catalase, oxidase, and DNAse?
- What happens to RBCs in beta and alpha hemolysis?
Contributors and Attributions
Jackie Reynolds, Professor of Biology (Richland College)


