18.5: Pathways of Amino Acid Degradation
- Page ID
- 37268
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)-
Differentiate Amino Acid Fates:
• Categorize amino acids as purely glucogenic, purely ketogenic, or both, and explain the structural reasons behind these classifications.
• Identify which amino acids are converted to pyruvate, acetyl-CoA (or acetoacetate), TCA cycle intermediates (e.g., α-ketoglutarate, succinyl-CoA), or have multiple fates. -
Trace Metabolic Pathways:
• Map the key pathways for converting amino acid carbon skeletons into central metabolites, such as the conversion of alanine to pyruvate or threonine to glycine, propionyl-CoA, and ultimately succinyl-CoA.
• Explain how different pathways (e.g., the VOMIT pathway for valine, isoleucine, threonine, and methionine) replenish the TCA cycle via anaplerotic reactions. -
Understand Enzymatic Reactions and Cofactor Roles:
• Describe the role of PLP-dependent enzymes in transamination and deamination reactions that facilitate the conversion of amino acids to pyruvate and other intermediates.
• Identify the key cofactors (such as PLP, NAD⁺/NADH, tetrahydrobiopterin, and biotin) involved in these pathways and explain their specific functions. -
Analyze Pathway Integration:
• Explain how the degradation of amino acids integrates with gluconeogenesis, ketogenesis, and the TCA cycle, emphasizing the importance of metabolic flexibility.
• Discuss the interplay between amino acid degradation and energy metabolism, including the implications for fasting states and overall metabolic homeostasis. -
Explore Specific Pathway Examples:
• Detail the conversion of tryptophan to alanine and acetoacetate, highlighting the branching of pathways that contribute to both glucogenic and ketogenic outcomes.
• Compare and contrast the metabolism of amino acids like lysine (via saccharopine and acetoacetyl-CoA formation) with the pathways leading from branched-chain amino acids (Val, Ile, and Leu) to succinyl-CoA or acetyl-CoA. -
Relate Metabolic Pathways to Physiology:
• Interpret how these catabolic processes contribute to overall cellular metabolism and the maintenance of energy balance, especially during protein breakdown or fasting.
• Evaluate the significance of amino acid catabolism in the context of systemic nitrogen and carbon balance as well as its clinical implications, as highlighted by integrated pathway diagrams like the Reactome summary.
These learning goals are designed to help students build a comprehensive understanding of how amino acid carbon skeletons are processed during degradation, reinforcing both the biochemical mechanisms and the broader physiological context of these pathways.
Introduction
In previous sections, we saw how nitrogen is removed from amino acids to produce urea or NH4+, that some amino acids are glucogenic, ketogenic, or both, and the role of tetrahydrofolate derivatives and S-adenosylmethionine in 1C transfer reactions. Now, we can focus on how the carbon skeletons of amino acids are processed during degradation.
Here are some key features of amino acid catabolism that were discussed in the previous section.
- some are converted to pyruvate, the end product of glycolysis and the start reactant of gluconeogenesis. Hence, these amino acids are glucogenic;
- some are converted to acetoacetate-CoA and or acetyl-CoA. Both can be converted into ketone bodies (acetoacetate and β-hydroxybutyrate), which are considered ketogenic. Since the two carbons of the acetyl group of acetyl-CoA are lost as CO2 in the TCA cycle, and there is no reverse for the pyruvate dehydrogenase reaction (pyr → acetyl-CoA), acetyl-CoA formed by amino acid degradation can not be used to create glucose in a net fashion;
- some are metabolized to form TCA intermediates. Since they are added in a net fashion to the TCA cycle and don't remove the existing pool of TCA intermediates, they can produce, in a net fashion, either directly or indirectly, molecules that can be used to produce glucose. These entry reactions to the TCA, which replenish or add to TCA intermediates, are called anaplerotic (replenishing) reactions. Hence, these amino acids are also glucogenic.
- some have multiple ways to be degraded and can produce acetyl-CoA and pyruvate, so they are both glucogenic and ketogenic.
Let's get more explicit:
- purely ketogenic: only Leu and Lys (the only amino acids whose name starts with L, and you have to Love them since there are only two amino acids in this category)
- both: 5 are, including the aromatics - Trp, Tyr, Phe - along with Ile/Thr
- purely glucogenic: the rest
Figure \(\PageIndex{1}\), also shown in the previous sections, summarizes the fates of the 20 amino acids in their catabolic reactions
Figure \(\PageIndex{1}\): Fates of the 20 amino acids in their catabolic reactions
Given the myriad of enzymes and pathways involved, we won't delve into the mechanisms for the reactions or the structures of the enzymes, except one for lysine metabolism.
Conversion to Pyruvate: Ala, Trp, Cys, Ser, Gly, Thr
We concluded Section 18.3 with a discussion of the Ser Gly One Carbon Cycle (SGOC), so some of this will be a review.
Figure \(\PageIndex{2}\) shows an overview of the conversion of amino acids to pyruvate. More details are provided for each step below.
Figure \(\PageIndex{2}\): Overview of conversion of amino acids to pyruvate
The metabolic steps for the chemical transformations shown in A-F are described in more detail below.
Tryptophan to alanine and on to acetoacetate
This is a multistep process, as shown in Figure \(\PageIndex{3}\).
Figure \(\PageIndex{3}\): Conversion of Tryptophan to Alanine and to acetoacetate
The starting material, tryptophan, is highlighted in a red box while the end product of specific interest, Ala, is highlighted in a green box. No reaction occurs in isolation in a cell, but rather as part of a more complex pathway. In the figure above, Ala is presented almost as a side product as the modified aromatic ring found in either anthranilate or 3-hydroxyanthranilate continues to form either acetoacetate, a ketone body which can break down to acetyl-CoA (making tryptophan ketogenic as well as glucogenic) or NAD+.
Alanine to Pyruvate
As described in 18.2 and shown in Figure \(\PageIndex{4}\), the PLP-dependent enzyme ALanine Amino Transferase (ALT), also known as Glutamate Pyruvate Transaminase (GPT), catalyzes this simple transamination reaction:
alanine +α−ketoglutarate ↔ pyruvate + glutamate
Figure \(\PageIndex{4}\): Alanine to pyruvate
The glutamate produced in this reaction can be oxidatively deaminated to give NH4+ and α-ketoglutarate again, as shown below:
glutamate + H2O + NAD+ → α-ketoglutarate + NADH + H+
The sum (net) of the two reactions is:
Alanine + H2O + NAD+ → pyruvate + NADH + H+
Threonine to Glycine
There are several pathways for this conversion.
One involves the conversion of Thr to 2-amino-3-ketobutyrate by threonine-3-dehydrogenase.
Rx: Thr + NAD+ ↔ 2-amino-3-ketobutyrate + NADH
This is followed by the conversion of 2-amino-3-ketobutyrate to glycine by the enzyme 2-amino-3-ketobutyrate coenzyme A ligase.
Rx: 2-amino-3-ketobutyrate + CoASH ↔ Gly + acetyl-CoA
The net of these reactions is
Rx: Thr + NAD+ + CoASH ↔ Gly + acetyl-CoA + NADH
These reactions are illustrated in Figure \(\PageIndex{5}\).
Figure \(\PageIndex{5}\): Threonine to glycine
A second and predominant reaction involves the conversion of Thr to NH4 + and α-ketobutyrate by the PLP-dependent enzyme Ser/Thr dehydratase (also called threonine ammonia-lyase). We saw this enzyme in the previous section. Note this reaction does NOT produce glycine but is an intermediate, α-ketobutyrate.
Rx: Thr ↔ NH4 + + α-ketobutyrate
α-ketobutyrate can then be converted to propionyl-CoA.
Rx: α-ketobutyrate + NAD+ + CoASH ↔ propionyl-CoA + NADH + CO2 + H+
This reaction, catalyzed by the inner mitochondrial membrane branched-chain α-ketoacid dehydrogenase complex (BCKDC or BCKDH complex), is an oxidative decarboxylation reaction. BCKDC is a member of two other enzymes, pyruvate dehydrogenase and alpha-ketoglutarate dehydrogenase, both of which act on short alpha-keto acids to produce key Kreb cycle metabolites.
Propionyl-CoA is then converted, in several mitochondrial steps, to succinyl-CoA for entry into the TCA cycle. Three enzymes are required for this conversion: propionyl-CoA carboxylase, methylmalonyl-CoA epimerase, and methylmalonyl-CoA mutase. Propionyl carboxylase, like another alpha-keto acid carboxylase (pyruvate carboxylase), requires ATP, Biotin, and CO2 (as a substrate) for the carboxylation reaction and hence is often referred to as an ABC enzyme.
The three-step conversion pathway of propionyl CoA to succinyl CoA is also used for the degradation of Valine, Odd-chain fatty acids (which form multiple 2-carbon acetyl CoA units and 1 3-C propionyl CoA unit), Methionine, and Isoleucine, along with Threonine. This three-step pathway is sometimes referred to as the VOMIT pathway.
The third pathway, which we just saw in the previous section, is catalyzed by serine hydroxymethyltransferase (SHMT) (but also called glycine hydroxymethyltransferase or threonine aldolase) and requires the use of both PLP and tetrahydrofolate as cofactors. A 1C methylene is added to tetrahydrofolate (FH4). PLP forms bonds to the alpha-carbon of amino acids that are labile to cleavage. In this case, the amino acid threonine dehydrates through an alpha-elimination reaction. However, threonine has an extra CH3 group, which is released as acetaldehyde. Here is the overall reaction.
Rx: Thr+ FH4 + ↔ Glycine + N5,N10-FH4 + acetaldehyde + H2O
The literature is a bit unclear about the enzymes involved in this reaction. SHMT appears to act on Thr at a lower rate, but a second enzyme, threonine aldolase, which seems to be afunctional in mammals, acts in other organisms.
Glycine to Serine
As mentioned above, this reversible reaction is catalyzed by serine hydroxymethyltransferase (SHMT) (see the mechanism in section 18.4) and uses tetrahydrofolate and PLP as cofactors. Here is the overall reaction, the reverse of the Gly ↔ Ser we saw in 18.4.
Rx: Glycine + N5,N10-CH2-FH4 + H2O ↔ Serine + FH4
Figure \(\PageIndex{6}\) shows the serine dehydratase reaction presented in Chapter 18.4. Figure A below shows the dehydration reaction and formation of glycine. using PLP as a cofactor. Figure B shows how the released formaldehyde reacts with FH4 to form N5,N10-methylene FH4, using FH4 as a cofactor.
Figure \(\PageIndex{6}\): Reversible reaction of Serine to Glycine
Serine to Pyruvate
This reaction is analogous to the Ala → Pyr reaction in Rx B above and is catalyzed by the PLP-dependent enzyme serine/threonine dehydratase/threonine deaminase.
Rx: Serine ↔ Pyr + NH4+
The enzyme is found in the cytoplasm and is mainly involved in gluconeogenesis.
Cysteine to Pyruvate
The overall reactions for this conversion are shown in the figure below. The aspartate aminotransferase used in the production of 3-sulfinylpyruvate is cytosolic and not the same as the more abundant version in the mitochondria.
The reaction pathway is shown in Figure \(\PageIndex{7}\).
Figure \(\PageIndex{7}\): Cysteine to pyruvate
Other important metabolites are made from cysteine catabolic pathways. One is taurine, the most abundant free amino acid in the body. It is especially abundant in development and early milk. It is synthesized predominantly in the liver. It is unclear if hypotaurine is converted to taurine in a non-enzymatic fashion or by an oxidase/dehydrogenase.
The sulfate produced in these pathways is utilized to synthesize an interesting derivative of ATP, 3′-phosphoadenosine-5′-phosphosulfate (PAPS), which is subsequently used to produce sulfated sugars essential for glycolipid and proteoglycan synthesis. This is illustrated in Figure \(\PageIndex{8}\).
Figure \(\PageIndex{8}\): Sulfate conversion to PAPS
Conversion to Acetyl-CoA: Trp, Lys, Phe, Tyr, Leu, Ile, Thr
An overview of the many reactions in ketogenic amino acid degradation is shown in Figure \(\PageIndex{9}\). The red-boxed amino acids are those that form either acetoacetate (a ketone body) or acetyl-CoA directly (green boxes). Some of the carbons are color-coded red or green to indicate where they end up.
Figure \(\PageIndex{9}\): Ketogenic amino acid pathway
Trp to acetyl-CoA
Fortunately, we have explored the conversion of the non-ring part of tryptophan to alanine and a precursor of acetoacetyl CoA (2-amino-3-carboxymuconate 6-semialdehyde - ACMS) and to NAD+ (quinolinate). ACMS, through the action of ACMS decarboxylase, leads to acetoacetyl CoA and then to acetyl-CoA as shown in Figure \(\PageIndex{10}\). As Trp is a ketogenic amino acid, it seems appropriate to show the steps that lead to acetyl-CoA even at the risk of providing too much detail.
Figure \(\PageIndex{10}\): Part 2 - Tryptophan to acetyl-CoA
Lys metabolism
In the liver, the main pathway (of several) starts with the formation of saccharopine from the transamination reaction of lysine and α-ketoglutarate, allowing the ε-amino group of lysine to enter the nitrogen metabolic pool. This transamination does not use pyridoxal phosphate (PLP). The first two steps of the reaction are catalyzed by an enzyme, α-aminoadipic semialdehyde synthase, with two activities (condensation/reduction and hydrolysis/oxidation). Lysine is an essential amino acid since the transamination is not reversible. Figure \(\PageIndex{11}\) shows pathways for the conversion of lysine to acetoacetyl-CoA and acetyl-CoA.
Figure \(\PageIndex{11}\): Pathways for conversion of lysine to acetoacetyl-CoA and acetyl-CoA.
The lysine-oxoglutarate reductase (LOR) and saccharopine dehydrogenase (SDH) are found in a single bifunctional enzyme, often referred to as aminoadipic semialdehyde synthase.
Figure \(\PageIndex{12}\) shows an interactive iCn3D model of the AlphaFold predicted structure of aminoadipic semialdehyde synthase (Q9UDR5)
Figure \(\PageIndex{12}\): AlphaFold predicted structure of aminoadipic semialdehyde synthase (Q9UDR5). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...Pi1c7Y5A9DmLa8
The cyan domain represents the lysine-oxoglutarate reductase (LOR) domain, which is connected by an alpha helix to the magenta saccharopine dehydrogenase (SDH) domain.
Phenylalanine is converted to Tyrosine and continues to acetoacetate
We'll follow the conversion of phenylalanine to tyrosine, which proceeds to acetoacetate, resulting in both phenylalanine and tyrosine (ketogenic amino acids), and subsequently produces fumarate. They can enter the TCA cycle, leading to the net production of oxaloacetate, which can be pulled off into gluconeogenesis, making both Phe and Try glucogenic as well.
A new cofactor is required that facilitates electron flow in the conversion of Phe to Try in the first step, catalyzed by the enzyme tyrosine hydrolase. That cofactor is tetrahydrobiopterin (BH4). The reaction involves the hydroxylation of BH4, followed by its transfer to phenylalanine. Figure \(\PageIndex{13}\) shows a possible mechanism for the conversion of phenylalanine to tyrosine with tetrahydrobiopterin (BH4).
Figure \(\PageIndex{13}\): Conversion of phenylalanine to tyrosine with tetrahydrobiopterin (BH4)
As in the case of converting dihydrofolate back to tetrahydrofolate (FH4) by dihydrofolate reductase, the 4a-OH-BH4 is converted to dihydrobiopterin and then to tetrahydrobiopterin by dihydrobiopterin reductase.
Figure \(\PageIndex{14}\) shows the full pathway for the conversion of Phe and Tyr to acetoacetate and fumarate.
Figure \(\PageIndex{14}\): Conversion of phenylalanine and tyrosine to acetoacetate and fumarate
Leu to Acetoacetate
The conversion of leucine to acetoacetate is shown in Figure \(\PageIndex{15}\).
Figure \(\PageIndex{15}\): Conversion of leucine to acetoacetate
The first reaction is a transamination using the PLP-dependent branched-chain aminotransferase (BCAT) with α-ketoglutarate.
Isoleucine to Acetyl-CoA
Figure \(\PageIndex{16}\) shows the pathway for the conversion of isoleucine to acetyl-CoA.
Figure \(\PageIndex{16}\): Pathway for converting isoleucine to acetyl-CoA.
Conversion to α-ketoglutarate: Pro, Glu, Gln, Arg,His
Proline and Arginine
The conversion of proline (bottom left) to glutamate (top left) is shown in Figure \(\PageIndex{17}\).
Figure \(\PageIndex{17}\): Conversion of proline (bottom left) to glutamate (top left)
Glutamate can then form α-ketoglutarate, so the reaction is glucogenic.
The conversions of arginine (and proline) to α-ketoglutarate are shown in Figure \(\PageIndex{18}\).
Figure \(\PageIndex{18}\): Conversion of arginine and proline to α-ketoglutarate
Histidine
The conversion of histidine to α-ketoglutarate is shown in Figure \(\PageIndex{19}\).
Figure \(\PageIndex{19}\): Conversion of histidine to α-ketoglutarate
As described in the reactions above, histidine can be converted to α-ketoglutarate through transamination reactions. Additionally, we described in a previous section how glutamine can be deaminated through the action of glutaminase to form glutamate, which can subsequently form α-ketoglutarate, a gluconeogenic intermediate.
Conversion to succinyl-CoA: Met and the branched-chain amino acids Ile, Thr, Val
We have just seen that two branched-chain amino acids, Leu and Ile, are converted to acetyl-CoA and, therefore, are ketogenic (E and F above). The branched-chain hydrophobic amino acids, Val and Leu, can be converted to succinyl-CoA, which can be converted to α-ketoglutarate in the Krebs' cycle in a net fashion and hence are glucogenic amino acids. We saw in the introduction to amino acids that produce acetyl-CoA that threonine and isoleucine, two branched-chain amino acids, also form propionyl-CoA, which goes on to succinyl-CoA. So, let's consider Val, another branched-chain amino acid, before we consider Met, both of which have three C atoms in their side chains.
Valine
The conversion of valine to succinyl-CoA is shown in Figure \(\PageIndex{20}\).
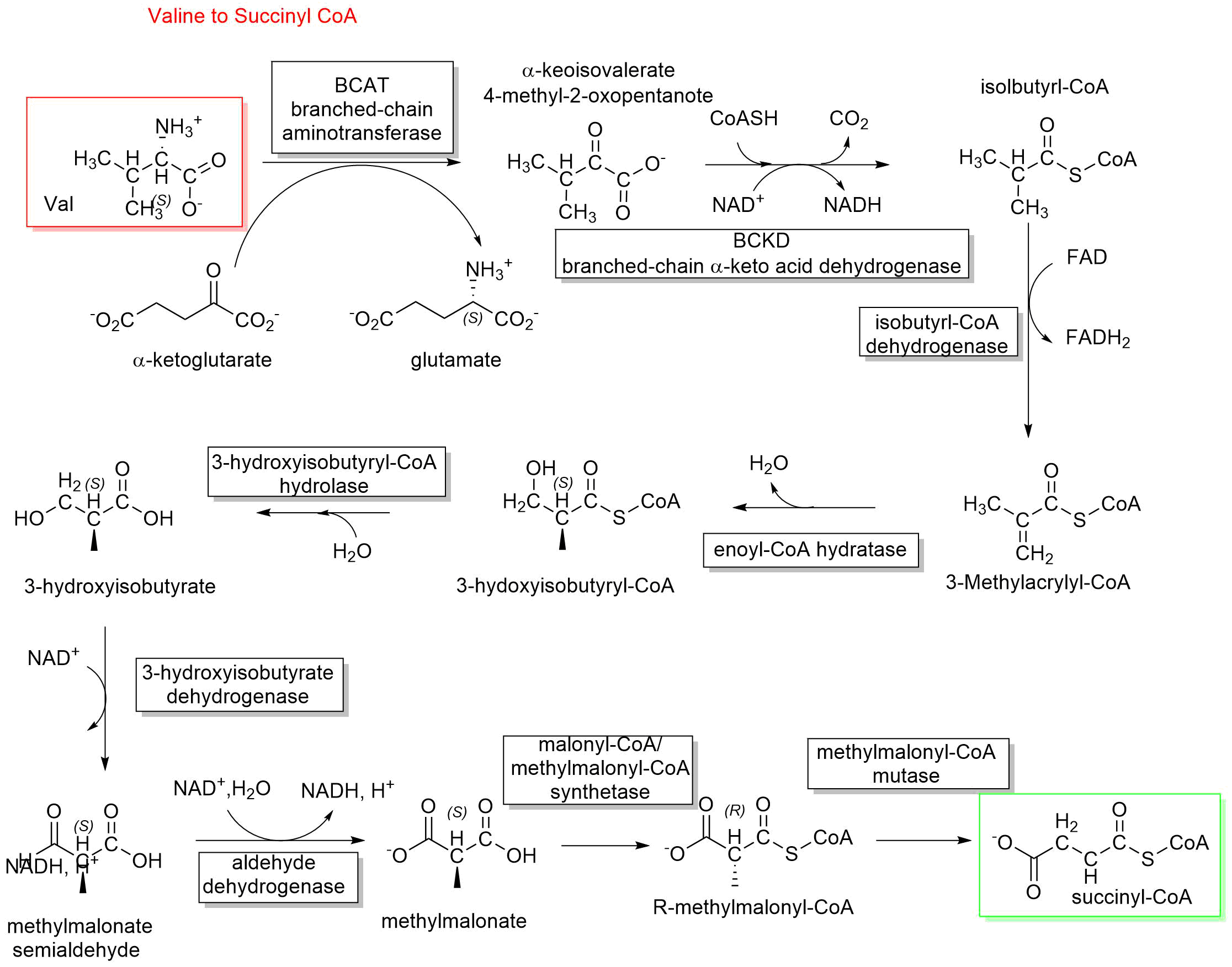
Figure \(\PageIndex{20}\): Conversion of valine to succinyl-CoA
The other two amino acids with branched-chain carbon chains (isoleucine and leucine) use the same enzymes as valine to enter the degradation pathway. They start with branched-chain transaminases (BCATc or BCATm) followed by oxidative decarboxylation reactions catalyzed by branched-chain ketoacid dehydrogenase (BCKD). Three different enzymes are required for the following dehydrogenase reaction. These are short/branched-chain acyl-CoA dehydrogenase (SBCAD) for isoleucine, isovaleryl-CoA dehydrogenase (IVD) for leucine, and isobutyryl-CoA dehydrogenase (IBD) for valine.
Methionine
Methionine can be metabolized to S-adenosylhomocysteine (SAM) and on to cysteine and α-ketobutyrate, which can also be produced by a transsulfuration reaction that produces cysteine. That product is metabolized using branched-chain dehydrogenases to produce succinyl-CoA, a key intermediate in the TCA cycle. Three enzymes are needed to convert the α-ketobutyrate to succinyl-CoA. Propionyl-CoA carboxylase uses ATP, biotin, and CO2, while the methylmalonyl-CoA mutase requires vitamin B12. An additional enzyme is involved in an epimerase reaction, in which D-methylmalonyl-CoA is converted into L-methylmalonyl-CoA. The conversion of propionyl-CoA to succinyl-CoA also occurs for branched-chain amino acids (Val, Ile, Thr) as well as Met, and in addition, Odd number fatty acids. This odd assortment of substrates for conversion to succinyl-CoA leads to the name VOMIT pathways. These reactions are illustrated in Figure \(\PageIndex{21}\).
Figure \(\PageIndex{21}\): Methionine conversion to succinyl-CoA
Finally, Aspartate and Asparagine
Using the enzyme L-asparaginase, asparagine is converted to NH3 and aspartate. Aspartate is then used in a transamination reaction to form oxaloacetate, a reactant in gluconeogenesis. Aspartate participates in the urea cycle, which helps eliminate nitrogen. Glutamate also acquires NH3 through the reaction catalyzed by glutamine synthase.
Both amino acids serve as substrates for transamination reactions, which produce TCA intermediates. Glutamate dehydrogenase can lead to alpha-ketoglutarate.
Glutamate and aspartate play important roles in collecting and eliminating amino nitrogen through glutamine synthetase and the urea cycle, respectively. The catabolic path of the carbon skeletons involves simple 1-step aminotransferase reactions that directly produce net quantities of a TCA cycle intermediate. The glutamate dehydrogenase reaction operating in the direction of 2-oxoglutarate (α-ketoglutarate) production provides a second avenue leading from glutamate to gluconeogenesis.
This SUMMARY GRAPHIC From Reactome shows "Cellular metabolism of amino acids and related molecules includes the pathways for the catabolism of amino acids, the biosynthesis of the nonessential amino acids (alanine, arginine, aspartate, asparagine, cysteine, glutamate, glutamine, glycine, proline, and serine) and selenocysteine, the synthesis of urea, and the metabolism of carnitine, creatine, choline, polyamides, melanin, and amine-derived hormones. The metabolism of amino acids provides a balanced supply of amino acids for protein synthesis. In the fasting state, the catabolism of amino acids derived from the breakdown of skeletal muscle protein and other sources is coupled to the processes of gluconeogenesis and ketogenesis to meet the body’s energy needs in the absence of dietary energy sources."
Provided by Reactome. Citation Accessed on Wed, May 15, 2024. Fabregat A, Sidiropoulos K, Viteri G, Marin-Garcia P, Ping P, Stein L, D'Eustachio P, Hermjakob H. Reactome diagram viewer: data structures and strategies to boost performance. Bioinformatics (Oxford, England). 2018 Apr;34(7) 1208-1214. doi: 10.1093/bioinformatics/btx752. PubMed PMID: 29186351. PubMed Central PMCID: PMC6030826. Image: https://reactome.org/PathwayBrowser/#/R-HSA-71291&PATH=R-HSA-1430728
Summary
This chapter builds on earlier discussions of nitrogen removal and one‐carbon metabolism by focusing on the fate of amino acid carbon skeletons after deamination. It explores how these carbon fragments are channeled into central metabolic pathways, illustrating their diverse roles in energy production and biosynthesis.
Key Concepts:
-
Classification of Amino Acids by Catabolic Fate:
Amino acids are categorized based on their degradation products:
• Glucogenic Amino Acids produce pyruvate or TCA cycle intermediates (e.g., α-ketoglutarate, oxaloacetate), which can be used for gluconeogenesis.
• Ketogenic Amino Acids yield acetyl-CoA or acetoacetyl-CoA, precursors for ketone body formation.
• Dual-Function Amino Acids provide both glucogenic and ketogenic intermediates. -
Major Pathways for Carbon Skeleton Conversion:
• Conversion to Pyruvate:
Amino acids such as alanine, tryptophan, cysteine, serine, glycine, and threonine are converted to pyruvate via transamination and deamination reactions. This process is crucial for fueling gluconeogenesis and energy production.
• Conversion to Acetyl-CoA and Acetoacetyl-CoA:
Several amino acids, including tryptophan, lysine, phenylalanine, tyrosine, leucine, isoleucine, and threonine, are degraded into acetyl-CoA or acetoacetyl-CoA. These intermediates serve as building blocks for ketone bodies, although the irreversible loss of acetyl carbons as CO₂ prevents their net use in glucose synthesis.
• Anaplerotic Reactions and TCA Cycle Replenishment:
Amino acids such as proline, arginine, histidine, glutamate, aspartate, and asparagine contribute carbon to the TCA cycle by forming intermediates like α-ketoglutarate, succinyl-CoA, or oxaloacetate. These reactions help replenish the cycle’s intermediates, supporting both energy generation and biosynthetic pathways. -
Detailed Metabolic Pathways:
• Pyruvate Formation:- Alanine to Pyruvate: Catalyzed by alanine aminotransferase (ALT/GPT), producing pyruvate and glutamate.
- Threonine Pathways: Threonine can be converted to glycine (via threonine dehydrogenase or serine hydroxymethyltransferase) or deaminated to form α-ketobutyrate, which feeds into further metabolic steps. • Conversion to Acetyl-CoA:
- Tryptophan: Undergoes a complex multi-step pathway yielding both alanine and a precursor for acetoacetyl-CoA.
- Lysine: Metabolized via saccharopine formation and further processing to yield acetoacetyl-CoA and acetyl-CoA.
- Aromatic Amino Acids (Phe/Tyr): Convert to acetoacetate and eventually to acetyl-CoA, with tetrahydrobiopterin (BH₄) facilitating key hydroxylation reactions. • Entry into the TCA Cycle:
- Branched-Chain Amino Acids (Val, Ile, Leu) and Methionine: Degraded via pathways (sometimes grouped under the VOMIT pathway) that generate propionyl-CoA and ultimately succinyl-CoA, replenishing the TCA cycle.
- Glutamate, Aspartate, and Related Amino Acids: Directly produce TCA intermediates through transamination reactions and subsequent dehydrogenase activity.
-
Metabolic Integration and Physiological Implications:
The degradation of amino acid carbon skeletons is tightly integrated with gluconeogenesis, ketogenesis, and the TCA cycle, ensuring that cells maximize energy extraction and maintain metabolic balance during fasting or high-protein conditions. These interconnected pathways exemplify metabolic flexibility and the efficient recycling of carbon, which is critical for overall cellular homeostasis. -
Visual Integration:
A comprehensive diagram (e.g., from Reactome) illustrates the interconnected nature of these pathways, emphasizing how amino acid degradation contributes to the broader network of cellular metabolism.
In summary, this chapter provides a detailed overview of how amino acid carbon skeletons are transformed and integrated into central metabolism. It underscores the dual role of these pathways in energy production and biosynthesis, highlighting the sophisticated regulatory mechanisms that maintain metabolic balance in response to varying physiological demands.




.png?revision=1&size=bestfit&width=484&height=285)