3.3: Introduction of recombinant DNA into cell and replication: Vectors
- Page ID
- 10523
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Vectors used to move DNA between species, or from the lab bench into a living cell, must meet three requirements (Figure \(\PageIndex{1}\)):
- They must be autonomously replicating DNA molecules in the host cell. The most common vectors are designed for replicating in bacteria or yeast, but there are vectors for plants, animals and other species.
- They must contain a selectable marker so cells containing the recombinant DNA can be distinguished from those that do not. An example is drug resistance in bacteria.
- They must have an insertion site to accommodate foreign DNA. Usually a unique restriction cleavage site in a nonessential region of the vector DNA. Later generation vectors have a set of about 15 or more unique restriction cleavage sites.
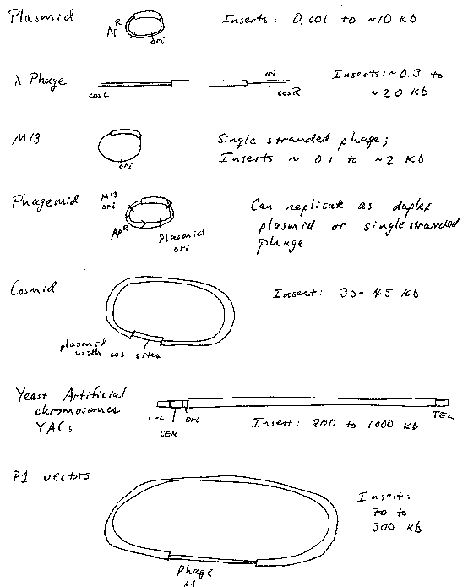
Plasmid Vectors
Plasmids are autonomously replicating circular DNA molecules found in bacteria. They have their own origin of replication, and they replicate independently of the origins on the "host" chromosome. Replication is usually dependent on host functions, such as DNA polymerases, but regulation of plasmid replication is distinct from that of the host chromosome. Plamsids, such as the sex-factor F, can be very large (94 kb), but others can be small (2‑4 kb). Plasmids do not encode an essential function to the bacterium, which distinguishes them from chromosomes. Plasmids can be present in a single copy, such as F, or in multiple copies, like those used as most cloning vectors, such as pBR322, pUC, and pBluescript.
In nature, plasmids provide carry some useful function, such as transfer (F), or antibiotic resistance. This is what keeps the plasmids in a population. In the absence of selection, plasmids are lost from bacteria. The antibiotic resistance genes on plasmids are often carried within, or are derived from, transposons, a types of transposable element. These are DNA segments that are capable of "jumping" or moving to new locations (Chapter 9).
A plasmid that was widely used in many recombinant DNA projects is pBR322 (Figure \(\PageIndex{2}\)). It replicates from an origin derived from a colicin-resistance plasmid (ColE1). This origin allows a fairly high copy number, about 100 copies of the plasmid per cell. Plasmid pBR322 carries two antibiotic resistance genes, each derived from different transposons. These transposons were initially found in R-factors, which are larger plasmids that confer antibiotic resistance.
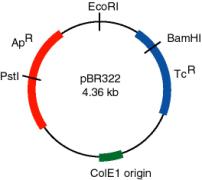
Use of the TcR and ApR genes allows for easy screening for recombinants carrying inserts of foreign DNA. For instance, insertion of a restriction fragment in the BamHI site of the TcR gene inactivates that gene. One can still select for ApR colonies, and then screen to see which ones have lost TcR .
Exercise \(\PageIndex{1}\)
What effects on drug resistance are seen when you use the EcoRI or PstI sites in pBR322 for inserting foreign DNA?
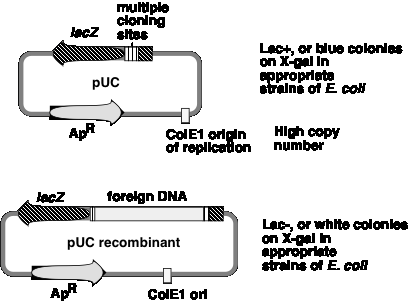
A generation of vectors developed after pBR322 are designed for even more efficient screening for recombinant plasmids, i.e. those that have foreign DNA inserted. The pUC plasmids (named for plasmid universal cloning) and plasmids derived from them use a rapid screen for inactivation of the b-galactosidase gene to identify recombinants (Figure \(\PageIndex{3}\)).
One can screen for production of functional b‑galactosidase in a cell by using the chromogenic substrate X‑gal (a halogenated indoyl b‑galactoside). When cleaved by b‑galactosidase, the halogenated indoyl compound is liberated and forms a blue precipitate. The pUC vector has the b‑galactosidase gene {actually only part of it, but enough to form a functional enzyme with the rest of the gene that is encoded either on the E. coli chromosome or an F' factor}. When introduced into E. coli, the colonies are blue on plates containing X‑gal.
The multiple cloning sites (unique restriction sites) are in the b‑galactosidase gene (lacZ). When a restriction fragment is introduced into one or more of these sites, the b‑galactosidase activity is lost by this insertional mutation. Thus cells containing recombinant plasmids form white(not blue) colonies on plates containing X‑gal.
The replication origin is a modified ColE1 origin of replication that has been mutated to eliminate a negative control region. Hence the copy number is very high(several hundred or more plasmid molecules per cell), and one obtains an very high yield of plasmid DNA from cultures of transformed bacteria. The plasmid has ApR as a selectable marker.