Lab 9: Standard Plate Count
- Page ID
- 24024
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)How do we know how many bacteria are in a liquid?
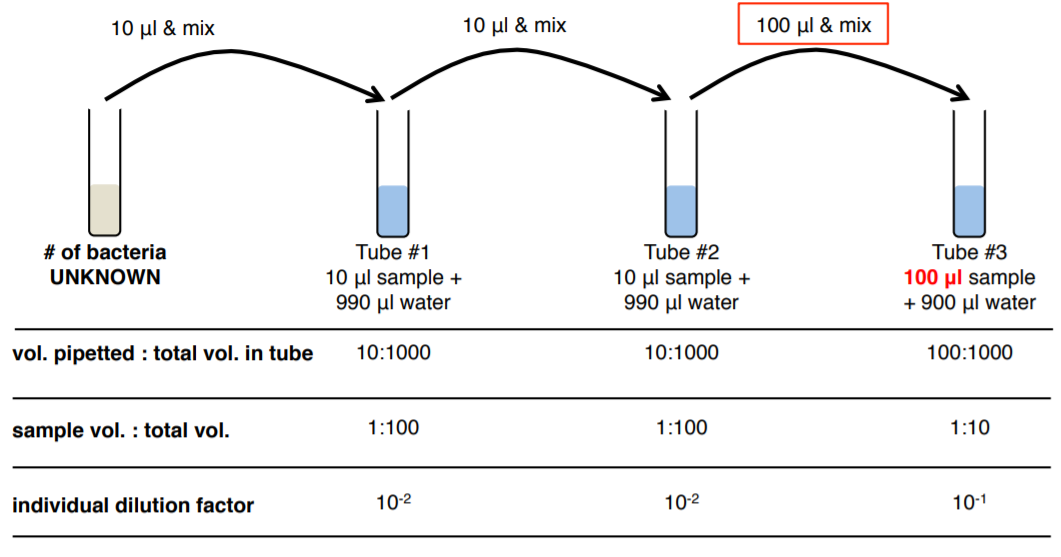
Microbiologists use a technique called the ‘standard plate count’ to estimate the population density of bacteria in a broth by plating a small and dilute portion of the sample and counting the number of bacteria colonies. We use serial dilutions to create decreasing concentrations of the original sample that are then plated so that a plate will be created with a low enough number of bacteria that we can count individual colonies. From that number, we can calculate the original cell density in the broth.
what do we dilute?
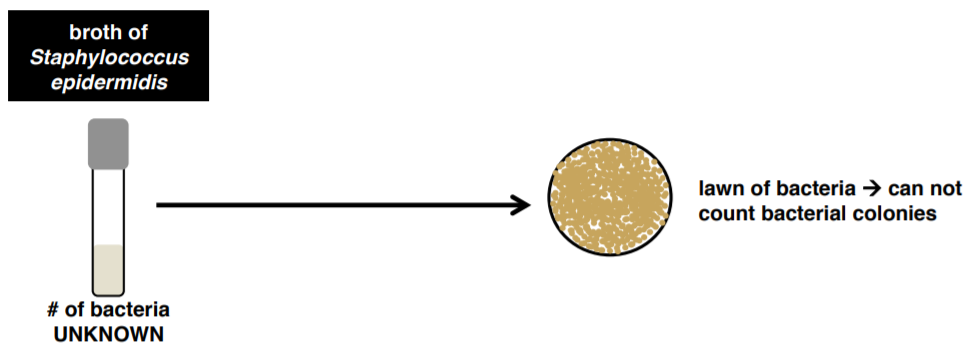
A turbid broth tube contains millions of bacteria. If we transferred directly from that tube, we would get a plate that had a lawn of bacteria. This plate would be uncountable and we could not use it to estimate bacterial cell numbers. Therefore, we need to dilute our original sample before plating.
Why do we do a dilution series?
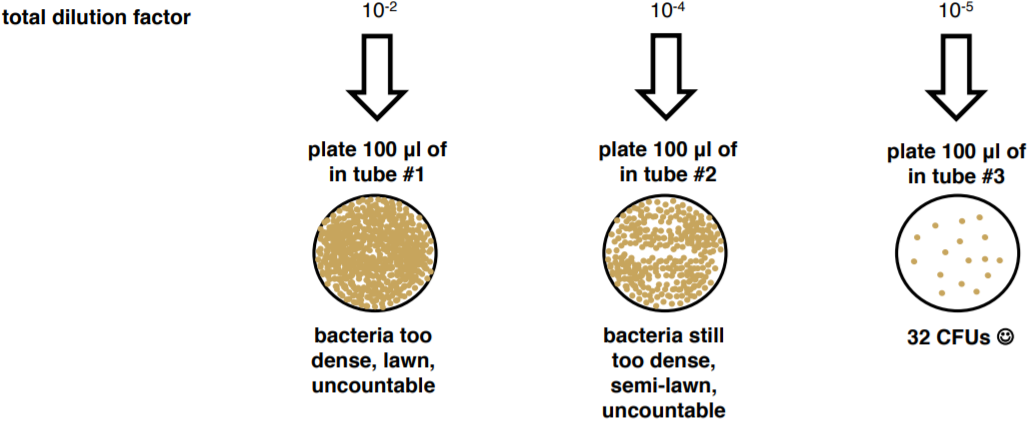
Since we don’t know how much bacteria is in our broth sample we don’t know how much we need to dilute it in order to get a ‘countable’ plate (between 30-300 bacteria). Therefore, we do a dilution series to create a number of plates, with a range of dilutions, in the hope of producing a countable plate. The number of colonies we count on a plate gives us the CFU or colony forming units, when we divide the CFU, by the volume we plated we get CFU/volume à CFU/mL. For example, if we counted 32 bacteria on a plate that had received 200 μl of a 10-6 dilution, we would do the following math:
\[\frac{32\;CFUs}{0.2\;mL}= 160\;\frac{CFU}{mL}\]
We then divide this by how much we diluted the original sample to get the estimated concentration of bacteria in the original tube:
\[\frac{160\;\frac{CFU}{mL}}{10^{-6}}= 160*10^6\;\frac{CFU}{mL}\]
Putting this in correct scientific notation this would be:
\[1.6*10^8 \frac{CFU}{mL}\]
NOTE
Calculate the CFU/mL for this experiment, show your work, and have the instructor sign off before proceeding.
Exercise
Fill in the chart below before proceeding (you do not need to turn this in).
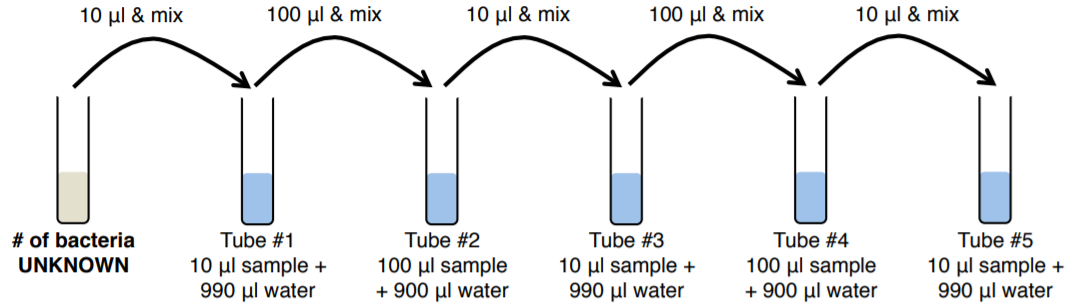
|
Volume Pipetted: Total Volume (Ratio) |
|||||
|
Sample Volume: Total Volume (Ratio) |
|||||
|
Individual Dilution Factor |
|||||
|
Total Dilution Factor |
Complete the Following Lab Individually:
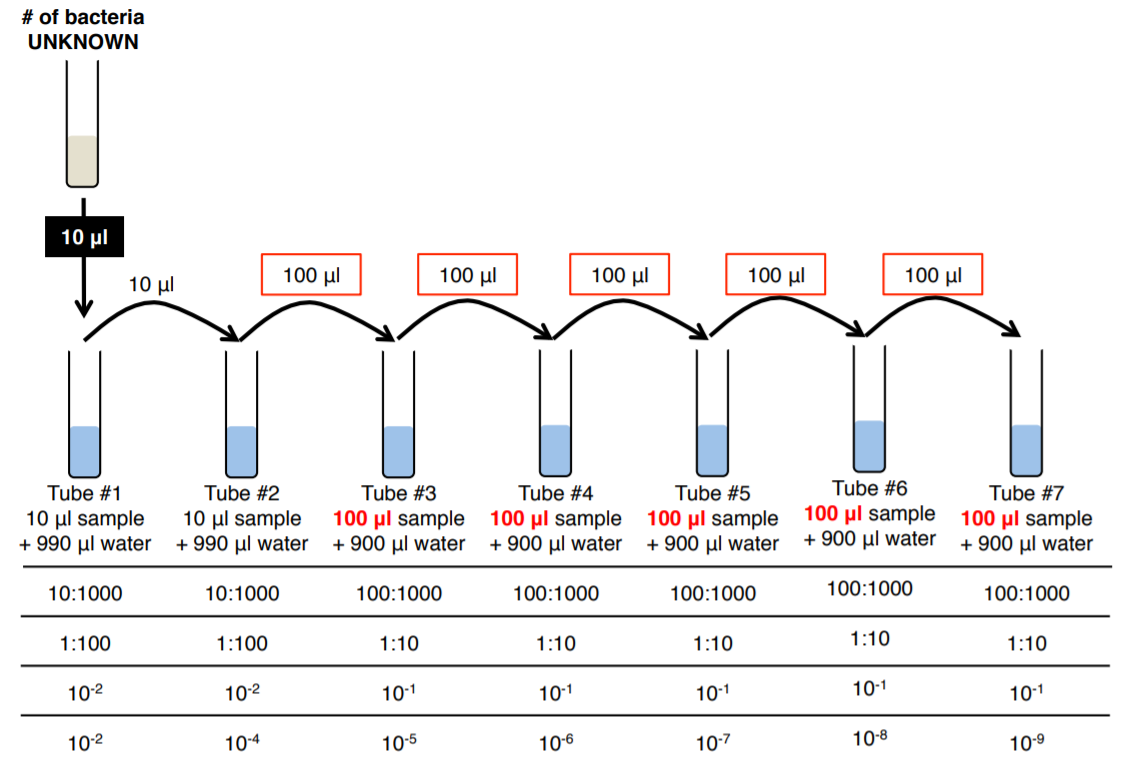
1. Obtain 7 microcentrifuge tubes and label them #1-7.
2. Aseptically add 990 μL of sterile water to tubes #1 and #2.
3. Aseptically add 900 μL of sterile water to tubes #3-7.
4. Mix the broth culture, then aseptically add 10 μL to tube #1 and vortex.
5. Aseptically transfer 10 μL from tube #1 to tube #2, close the lid and vortex.
6. Aseptically transfer 100 μL from tube #2 to tube #3, close the lid and vortex.
7. Aseptically transfer 100 μL from tube #3 to tube #4, close the lid and vortex.
8. Aseptically transfer 100 μL from tube #4 to tube #5, close the lid and vortex.
9. Aseptically transfer 100 μL from tube #5 to tube #6, close the lid and vortex.
10. Aseptically transfer 100 μL from tube #6 to tube #7, close the lid and vortex.
CONGRATULATIONS YOU HAVE NOW COMPLETED YOUR DILUTION SERIES!
11. Obtain 4 TSA plates and label them.
12. Using the magic of fire to bend a glass Pasteur pipette into a spreader. Place it in your beaker of alcohol.
13. Flame your glass spreader, wait for ALL the alcohol to evaporate and repeat this 3 times.
14. Aseptically pipette 100 μL of tube #4 onto the center of a TSA plate while making sure the tip does not touch the agar.
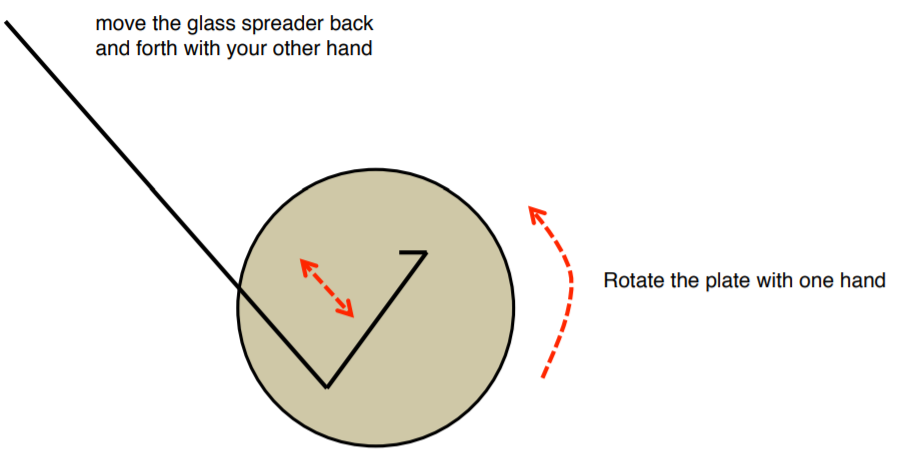
15. Place the sterilized spreader onto the center of your plate (DO NOT PRESS DOWN).
16. Turn the plate at least 5 times as you move the glass spreader to distribute the bacteria over the plate. At this point, the liquid has been absorbed by the plate.
17. Place the glass spreader back into your beaker of alcohol.
18. Repeat this procedure for tubes #5-7, incubate at 37°C.