19.1.12: Caenorhabditis Elegans
- Page ID
- 5960
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Caenorhabditis elegans is a microscopic (~1 mm) nematode (roundworm) that normally lives in soil. It has become one of the "model" organisms in biology. It is a true animal with at least rudiments of the physiological systems - feeding, nervous, muscle, reproductive - found in "higher" animals like mice and humans. However, it is so small that large numbers can be raised in petri dishes (where it is fed E. coli - another model organism). It reproduces rapidly.
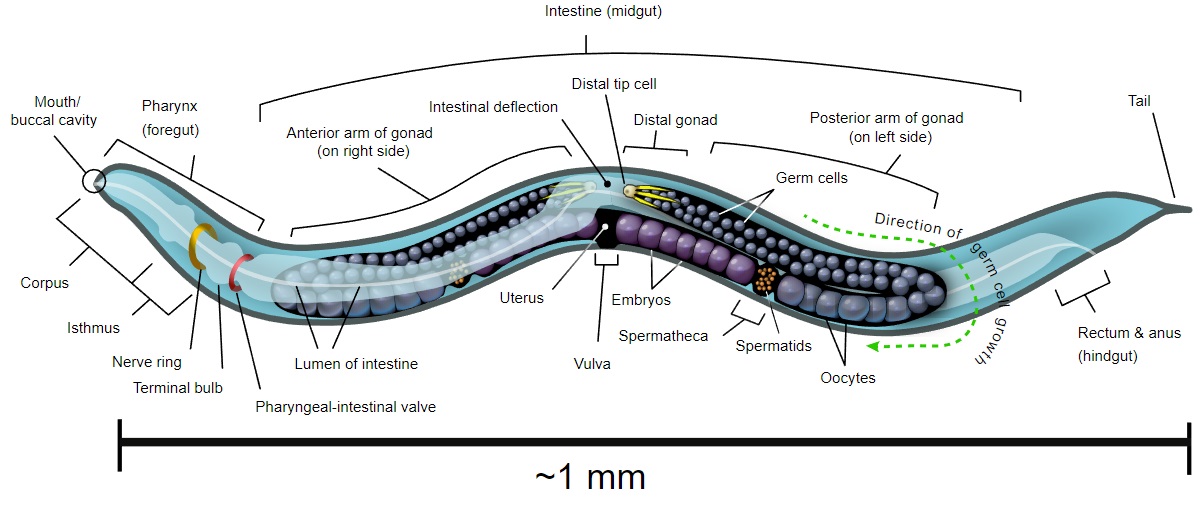
It is transparent so that every cell in the living animal can be seen under the microscope from the fertilized egg to the 556 cells of the newly-hatched worm and, later, the 959 somatic cells, and a variable number of germ cells, of the adult worm. It can be easily transformed with transgenes - DNA injected into the animal. It can also be treated with antisense RNA. Before it dies (after 2–3 weeks), it shows signs of aging and thus may provide general clues as to the aging process.
Its cells contain 5 pairs of autosomes and 2 X chromosomes. These animals are hermaphrodites, producing both sperm and eggs. Most of the time they fertilize themselves, so that any recessive alleles quickly become homozygous and affect the phenotype. On rare occasions, nondisjunction occurs during meiosis with the loss of one X chromosome. Animals with a single X are males and are able to fertilize the eggs of the hermaphrodites (with more success than they have themselves).
C. elegans was the first multicellular eukaryote to have its entire genome sequenced. It contains some 19–20,000 protein-encoding genes incorporated in 100,258,171 base pairs of DNA. In contrast to other eukaryotes, some 13–15% of its genes are grouped in operons containing 2–8 genes each.
C. elegans Fertilization
Like all animals, C. elegans starts life as a fertilized egg (zygote) which then undergoes the mitotic divisions needed to produce the adult. Because the worm is transparent and the pattern of differentiation is so rigid it has been possible to trace the lineage of every single somatic cell in the animal. Just after hatching, the it contains 556 cells and is approximately 0.3 mm long. After reaching maturity, it will contain 959 somatic cells and a variable number of germ cells in its gonad.
The diagrams below show the pathway by which each of the 556 cells in the larva of C. elegans has developed from the zygote. The relative length of the vertical lines indicates the length of the interval before the next mitosis. Some pathways end in the programmed death of the cell (apoptosis) even before the larva is complete.

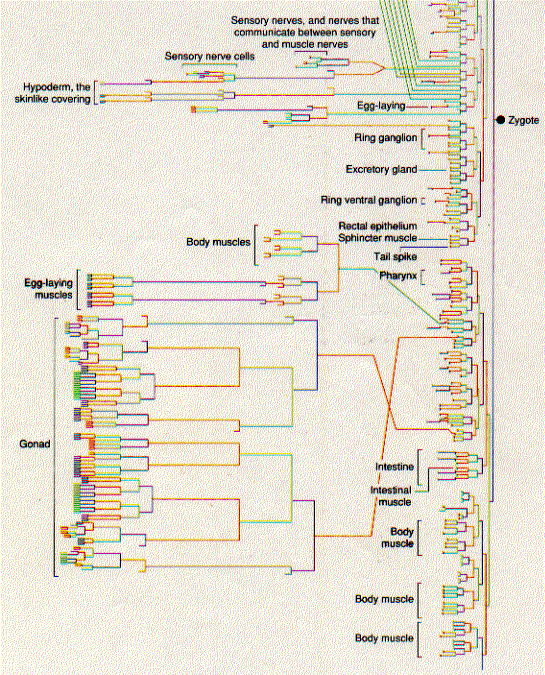
Several remarkable features have been found from these studies. The pattern of development is invariable from worm to worm. Every one of the 556 cells that make up the newly-hatched larva develops from a rigid pattern of mitotic division leading back to the zygote. 131 cells in the developing embryo die by apoptosis. This cell death is not random; which cells will die and at what stage is completely predictable. Any failures in this programmed cell death can lead to serious abnormalities.
Each organ - skin, nerve, muscle, etc. - is made of cells derived from several different lineages. One might have expected that the earliest cell divisions would produce daughter cells destined to go on to form a single structure in the embryo. But that is not generally the case. Instead, most of the earliest cells will produce descendants that team up with other groups of cells to form the various organs of the animal.
There is less flexibility of cell fate than occurs in mammals (or even amphibians). With a microscopic laser beam, a single cell can be killed in the developing embryo. Often the result is that all the cells that would have descended from that cell fail to form. Neighboring cells may not compensate for the loss as they do so freely in mammals. So it appears that the developmental controls of C. elegans rely more on cell-intrinsic signals rather than inductive signals. But this distinction is relative, not absolute. Cells originally destined to a different fate will in some cases switch their path of differentiation to replace the cells killed by the laser. In these cases, the switch is clearly mediated by inductive signals liberated by nearby cells.


