11.1: Recombinant DNA and Gene Cloning
- Page ID
- 4918
Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single recombinant molecule.
Making Recombinant DNA (rDNA) - An Overview
- Treat DNA from both sources with the same restriction endonuclease (BamHI in this case).
- BamHI cuts the same site on both molecules
5' GGATCC 3'
3' CCTAGG 5'
- The ends of the cut have an overhanging piece of single-stranded DNA.
- These are called "sticky ends" because they are able to base pair with any DNA molecule containing the complementary sticky end.
- In this case, both DNA preparations have complementary sticky ends and thus can pair with each other when mixed.
- A DNA ligase covalently links the two into a molecule of recombinant DNA.
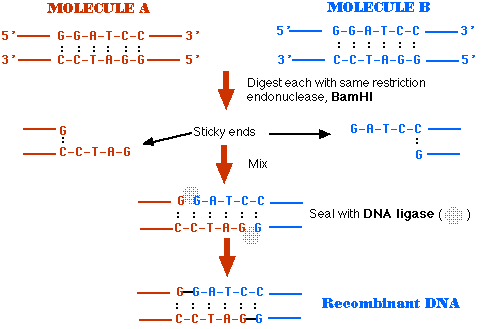
To be useful, the recombinant molecule must be replicated many times to provide material for analysis, sequencing, etc. Producing many identical copies of the same recombinant molecule is called cloning. Cloning can be done in vitro, by a process called the polymerase chain reaction (PCR). Here, however, we shall examine how cloning is done in vivo.
Cloning in vivo can be done in
- unicellular microbes like E. coli
- unicellular eukaryotes like yeast and
- in mammalian cells grown in tissue culture.
In every case, the recombinant DNA must be taken up by the cell in a form in which it can be replicated and expressed. This is achieved by incorporating the DNA in a vector. A number of viruses (both bacterial and of mammalian cells) can serve as vectors. But here let us examine an example of cloning using E. coli as the host and a plasmid as the vector.
Plasmids
Plasmids are small (a few thousand base pairs), usually carry only one or a few genes, are circular and have a single origin of replication. Plasmids are replicated by the same machinery that replicates the bacterial chromosome. Some plasmids are copied at about the same rate as the chromosome, so a single cell is apt to have only a single copy of the plasmid. Other plasmids are copied at a high rate and a single cell may have 50 or more of them.
Genes on plasmids with high numbers of copies are usually expressed at high levels. In nature, these genes often encode proteins (e.g., enzymes) that protect the bacterium from one or more antibiotics. Plasmids enter the bacterial cell with relative ease. This occurs in nature and may account for the rapid spread of antibiotic resistance in hospitals and elsewhere. Plasmids can be deliberately introduced into bacteria in the laboratory transforming the cell with the incoming genes.
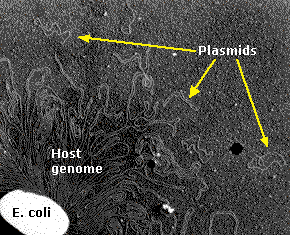
In the above figure, the tangle is a portion of a single DNA molecule containing over 4.6 million base pairs encoding approximately 4,300 genes. The small circlets are plasmids.
Examples of Plasmids
pAMP
- 4539 base pairs
- a single replication origin
- a gene (ampr)conferring resistance to the antibiotic ampicillin (a relative of penicillin)
- a single occurrence of the sequence
5' GGATCC 3'
3' CCTAGG 5'that, as we saw above, is cut by the restriction enzyme BamHI
- a single occurrence of the sequence
5' AAGCTT 3'
3' TTCGAA 5'that is cut by the restriction enzyme HindIII
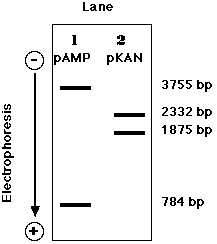
Treatment of pAMP with a mixture of BamHI and HindIII produces:
- a fragment of 3755 base pairs carrying both the ampr gene and the replication origin
- a fragment of 784 base pairs
- both fragments have sticky ends
pKAN
- 4207 base pairs
- a single replication origin
- a gene (kanr) conferring resistance to the antibiotic kanamycin.
- a single site cut by BamHI
- a single site cut by HindIII
Treatment of pKAN with a mixture of BamHI and HindIII produces:
- a fragment of 2332 base pairs
- a fragment of 1875 base pairs with the kanr gene (but no origin of replication)
- both fragments have sticky ends
These fragments can be visualized by subjecting the digestion mixtures to electrophoresis in an agarose gel. Because of its negatively-charged phosphate groups, DNA migrates toward the positive electrode (anode) when a direct current is applied. The smaller the fragment, the farther it migrates in the gel.
Ligation Possibilities
If you remove the two restriction enzymes and provide the conditions for DNA ligase to do its work, the pieces of these plasmids can rejoin (thanks to the complementarity of their sticky ends).
Mixing the pKAN and pAMP fragments provides several (at least 10) possibilities of rejoined molecules. Some of these will not produce functional plasmids (molecules with two or with no replication origin cannot function).

Fig.11.1.4 Recombinant Plasmid
One interesting possibility is the joining of
- the 3755-bp pAMP fragment (with ampr and a replication origin) with the
- 1875-bp pKAN fragment (with kanr)
Sealed with DNA ligase, these molecules are functioning plasmids that are capable of conferring resistance to both ampicillin and kanamycin. They are molecules of recombinant DNA.
Because the replication origin, which enables the molecule to function as a plasmid, was contributed by pAMP, pAMP is called the vector.
Transforming E. coli
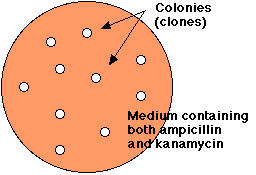
Treatment of E. coli with the mixture of religated molecules will produce some colonies that are able to grow in the presence of both ampicillin and kanamycin.
- A suspension of E. coli is treated with the mixture of religated DNA molecules.
- The suspension is spread on the surface of agar containing both ampicillin and kanamycin.
- The next day, a few cells — resistant to both antibiotics — will have grown into visible colonies containing billions of transformed cells.
- Each colony represents a clone of transformed cells.
However, E. coli can be simultaneously transformed by more than one plasmid, so we must demonstrate that the transformed cells have acquired the recombinant plasmid.
Electrophoresis of the DNA from doubly-resistant colonies (clones) tells the story.
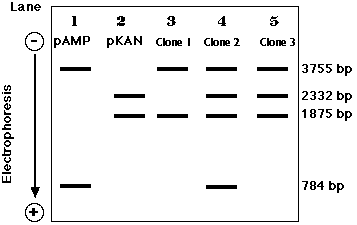
- Plasmid DNA from cells that acquired their resistance from a recombinant plasmid only show only the 3755-bp and 1875-bp bands (Clone 1, lane 3).
- Clone 2 (Lane 4) was simultaneous transformed by religated pAMP and pKAN. (We cannot tell if it took up the recombinant molecule as well.)
- Clone 3 (Lane 5) was transformed by the recombinant molecule as well as by an intact pKAN.
Cloning other Genes
The recombinant vector described above could itself be a useful tool for cloning other genes. Let us assume that within its kanamycin resistance gene (kanr) there is a single occurrence of the sequence
5' GAATTC 3'
3' CTTAAG 5'
This is cut by the restriction enzyme EcoRI, producing sticky ends.
If we treat any other sample of DNA, e.g., from human cells, with EcoRI, fragments with the same sticky ends will be formed. Mixed with EcoRI-treated plasmid and DNA ligase, a small number of the human molecules will become incorporated into the plasmid which can then be used to transform E. coli.
But how to detect those clones of E. coli that have been transformed by a plasmid carrying a piece of human DNA?
The key is that the EcoRI site is within the kanr gene, so when a piece of human DNA is inserted there, the gene's function is destroyed.
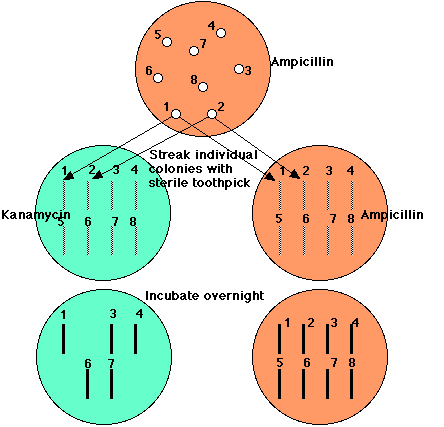
All E. coli cells transformed by the vector, whether it carries human DNA or not, can grow in the presence of ampicillin. But E. coli cells transformed by a plasmid carrying human DNA will be unable to grow in the presence of kanamycin. So,
- Spread a suspension of treated E. coli on agar containing ampicillin only
- grow overnight
- with a sterile toothpick transfer a small amount of each colony to an identified spot on agar containing kanamycin
- (do the same with another ampicillin plate)
- Incubate overnight
All those clones that continue to grow on ampicillin but fail to grow on kanamycin (here, clones 2, 5, and 8) have been transformed with a piece of human DNA.
Some recombinant DNA products being used in human therapy
Using procedures like this, many human genes have been cloned in E. coli or in yeast. This has made it possible — for the first time — to produce unlimited amounts of human proteins in vitro. Cultured cells (E. coli, yeast, mammalian cells) transformed with a human gene are being used to manufacture more than 100 products for human therapy. Some examples:
- insulin for diabetics
- factor VIII for males suffering from hemophilia A
- factor IX for hemophilia B
- human growth hormone (HGH)
- erythropoietin (EPO) for treating anemia
- several types of interferons
- several interleukins
- granulocyte-macrophage colony-stimulating factor (GM-CSF) for stimulating the bone marrow after a bone marrow transplant
- granulocyte colony-stimulating factor (G-CSF) for stimulating neutrophil production (e.g., after chemotherapy) and for mobilizing hematopoietic stem cells from the bone marrow into the blood.
- tissue plasminogen activator (TPA) for dissolving blood clots
- adenosine deaminase (ADA) for treating some forms of severe combined immunodeficiency (SCID)
- parathyroid hormone
- many monoclonal antibodies
- hepatitis B surface antigen (HBsAg) to vaccinate against the hepatitis B virus
- C1 inhibitor (C1INH) used to treat hereditary angioedema


