15: Positive and negative control of gene expression
- Page ID
- 386
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Operons
An operon is a cluster of coordinately regulated genes. It includes structural genes (generally encoding enzymes), regulatory genes (encoding, e.g. activators or repressors) and regulatory sites (such as promoters and operators). The type of control is defined by the response of the operon when no regulatory protein is present. In the case of negative control, the genes in the operon are expressed unless they are switched off by a repressor protein. Thus the operon will be turned on constitutively (the genes will be expressed) when the repressor in inactivated. In the case of positive control, the genes are expressed only when an active regulator protein, e.g. an activator, is present. Thus the operon will be turned off when the positive regulatory protein is absent or inactivated.
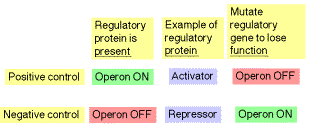
Table 4.1.1. Positive vs. negative control
Catabolic versus Biosynthetic Operons
Catabolic pathways catalyze the breakdown of nutrients (the substrate for the pathway) to generate energy, or more precisely ATP, the energy currency of the cell. In the absence of the substrate,there is no reason for the catabolic enzymes to be present, and the operon encoding them is repressed. In the presence of the substrate, when the enzymes are needed, the operon is induced or de-repressed.
| Operon encodes | Absence of | Effect | Presence of | Effect |
|---|---|---|---|---|
| catabolic enzymes | substrate | repressed | substrate | derepressed (induced) |
| biosynthetic enzymes | product | induced | product | repressed |
For example, the lac operon encodes the enzymes needed for the uptake (lactose permease) and initial breakdown of lactose (the disaccharide b-D-galactosyl-1->4-D-glucose) into galactose and glucose (catalyzed by b-galactosidase). These monosaccharides are broken down to lactate (principally via glycolysis, producing ATP), and from lactate to CO2 (via the citric acid cycle), producing NADH, which feeds into the electron-transport chain to produce more ATP (oxidative phosphorylation). This can provide the energy for the bacterial cell to live. However, the initial enzymes (lactose permease and b-galactosidase) are only needed, and only expressed, in the presence of lactose and in the absence of glucose. In the presence of the substrate lactose, the operon in turned on, and in its absence, the operon is turned off.
Anabolic, or biosynthetic, pathways use energy in the form of ATP and reducing equivalents in the form of NAD(P)H to catalyze the synthesis of cellular components (the product) from simpler materials, e.g. synthesis of amino acids from small dicarboxylic acids (components of the the citric acid cycle). If the cell has plenty of the product already (in the presence of the product), the the enzymes catalyzing its synthesis are not needed, and the operon encoding them is repressed. In the absence of the product, when the cell needs to make more, the biosynthetic operon is induced. E.g., the trpoperon encodes the enzymes that catalyze the conversion of chorismic acid to tryptophan. When the cellular concentration of Trp (or Trp-tRNAtrp) is high, the operon is not expressed, but when the levels are low, the operon is expressed.
Inducible versus repressible Operons
Inducible operons are turned on in reponse to a metabolite (a small molecule undergoing metabolism) that regulates the operon. E.g. the lac operon is induced in the presence of lactose (through the action of a metabolic by-product allolactose). Repressible operons are switched off in reponse to a small regulatory molecule. E.g., the trpoperon is repressed in the presence of tryptophan. Note that in this usage, the terms are defined by the reponse to a small molecule. Although lac is an inducible operon, we will see conditions under which it is repressed or induced (via derepression).
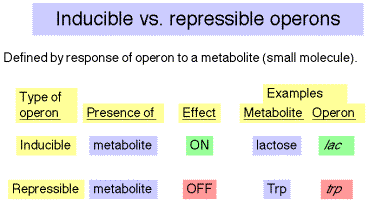
Table 4.1.3.
Map of the E. colilac operon
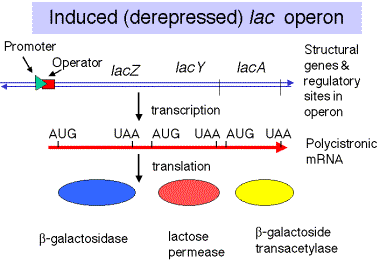
- Promoters = p= binding sites for RNA polymerase from which it initiates transcription. There are separate promoters for the lacIgene and the lacZYAgenes.
- Operator = o = binding site for repressor; overlaps with the promoter for lacZYA.
- Repressor encoded by lacIgene
- Structural genes: lacZYA
lacZ encodes b-galactosidase, which cleaves the disccharide lactose into galactose and glucose.
lacYencodes the lactose permease, a membrane protein that faciltitates uptake of lactose.
lacAencodes b-galactoside transacetylase; the function of this enzymes in catabolism of lactose is not understood (at least by me)
C. Negative control
The lac operon is under both negative and positive control. The mechanisms for these will be considered separately.
1. In negative control, the lacZYAgenes are switched off by repressor when the inducer is absent (signalling an absence of lactose). When the repressor tetramer is bound to o, lacZYAis not transcribed and hence not expressed.
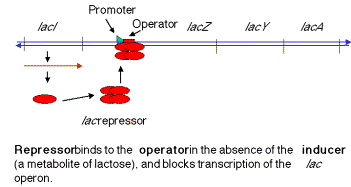
2. When inducer is present (signalling the presence of lactose), it binds the repressor protein, thereby altering its conformation, decreasing its affinity for o, the operator. The dissociation of the repressor-inducer complex allows lacZYAto be transcribed and therefore expressed.
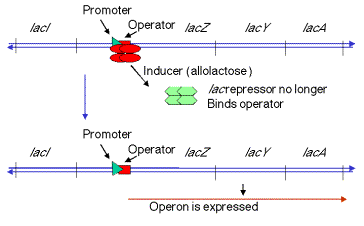
Inducers
The natural inducer (or antirepressor), is allolactose, an analog of lactose. It is made as a metabolic by-product of the reaction catalyzed by b-galactosidase. Usually this enzyme catalyzes the cleavage of lactose to galactose + glucose, but occasionally it will catalyze an isomerization to form allolactose, in which the galacose is linked to C6 of glucose instead of C4.
A gratuitous inducer will induce the operon but not be metabolized by the encoded enzymes; hence the induction is maintained for a longer time. One of the most common ones used in the laboratory is a synthetic analog of lactose called isopropylthiogalactoside (IPTG). In this compound the b-galactosidic linkage is to a thiol, which is not an efficient substrate for b-galactosidase.
E. Regulatory mutants
Regulatory mutations affect the amount of all the enzymes encoded by an operon, whereas mutations in a structural gene affects only the activity of the encoded (single) polypeptide.
Repressor mutants
- a. Wild-type strains (lacI+) are inducible.
- b. Most strains with a defective repressor (lacI-) are constitutive, i.e. they make the enzymes encoded by the lac operon even in the absence of the inducer.
- c. Strains with repressor that is not able to interact with the inducer (lacIS) are noninducible. Since the inducer cannot bind, the repressor stays on the operator and prevents expression of the operon even in the presence of inducer.
- d. Deductions based on phenotypes of mutants
| b-galactosidase | transacetylase | ||||
|---|---|---|---|---|---|
| Genotype | -IPTG | +IPTG | -IPTG | +IPTG | Conclusion |
| I+Z+A+ | <0.1> | 100 | <1> | 100 | Inducible |
| I+Z-A+ | <0.1> | <0.1> | <1> | 100 | lacZencodes b-galactosidase |
| I-Z+A+ | 100 | 100 | 90 | 90 | Constitutive |
| I+Z-A+ /F' I-Z+A+ | <0.1> | 100 | <1> | 200 | I+ >I- in trans |
| IsZ+A+ | <0.1> | <1> | <1> | <1> | Noninducible |
| IsZ+A+ /F' I+Z+A+ | <0.1> | 1 | <1> | 1 | Is>.I+ in trans |
- The wild-type operon is inducible by IPTG.
- A mutation in lacZaffects only b-galactosidase, not the transacetylase (or other products of the operon), showing that lacZis a structural gene.
- A mutation in lacIaffects both enzymes, hence lacIis a regulatory gene. Both are expressed in the absence of the inducer, hence the operon is constitutively expressed (the strain shows a constitutive phenotype).
- In a merodiploid strain, in which one copy of the lac operon is on the chromosome and another copy is on an F' factor, one can test for dominance of one allele over another. The wild-type lacI+is dominant over lacI-intrans. In a situation where the only functional lacZgene is on the same chromosome as lacI-, the functional lacI still causes repression in the absence of inducer.
- The lacISallele is noninducible.
- In a merodiploid, the lacISallele is dominant over wild-type in trans.
e. The fact that the product of the lacIgene is trans-acting means that it is a diffusible molecule that can be encoded on one chromosome but act on another, such as the F' chromosome in example (d) above. In fact the product of the lacIgene is a repressor protein.
2. Operator mutants
a. Defects in the operator lead to constitutive expression of the operon, hence one can isolate operator constitutive mutations, abbreviated oc. The wild-type o+is inducible.
b. Mutations in the operator are cis-acting; they only affect the expression of structural genes on the same chromosome.
(1)The merodiploid I+ocZ+/I+o+Z- [this is an abbreviation for lacI+oclacZ+/lacI+o+lacZ-] expresses b-galactosidase constitutively. Thus oc is dominant to o+ when oc is in cisto lacZ+.
(2)The merodiploid I+ocZ-/I+o+Z+ is inducible for b-galactosidase expression. Thus o+ is dominant to oc when o+ is in cisto lacZ+.
(3)The allele of othat is in cisto the active reporter gene (i.e., on the same chromosome as lacZ+ in this case) is the one whose phenotype is seen. Thus the operator is cis-acting, and this property is referred to as cis-dominance. As in most cases of cis-regulatory sequences, these are sites on DNA that are required for regulation. In this case the operator is a binding site for the trans-acting repressor protein.
Interactions between Operator and Repressor
Sequence of operator
The operator overlaps the start the site of transcription and the promoter. It has a dyad symmetry centered at +11. Such a dyad symmetry is commonly found within binding sites for symmetrical proteins (the repressor is a homotetramer). The sequence of DNA that consititutes the operator was defined by the position of oC mutations, as well as the nucleotides protected from reaction with, e.g. DMS, upon binding of the repressor.
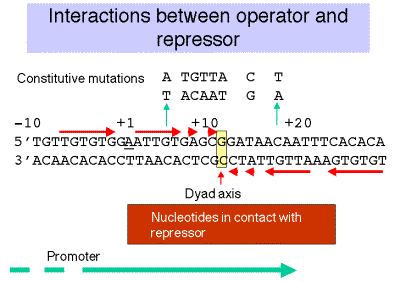
2. Repressor
a. Purification
(1)Increase the amount of repressor in the starting material by over-expression.
A wild-type cell has only about 10 molecules of the repressor tetramer. Isolation and purification of the protein was greatly aided by use of mutant strain with up-promoter mutations for lacI, so that many more copies of the protein were present in each cell. This general strategy of over-producing the protein is widely used in purification schemes. Now the gene for the protein is cloned in an expression vector, so that the host (bacteria in this case) makes a large amount of the protein - often a substantial fraction of the total bacterial protein.
(2)Assays for repressor
[1]Binding of radiolabeled IPTG (gratuitous inducer) to repressor
[2]Binding of radiolabeled operator DNA sequence to repressor. This can be monitored by the ability of the protein-DNA complex to bind to nitrocellulose (whereas a radiolabeled mutant operator DNA fragement, oc, plus repressor will not bind). Electrophoretic mobility shift assays would be used now in many cases.
[3]This ability of particular sequences to bind with high affinity to the desired protein is frequently exploited to rapidly isolate the protein. The binding site can be synthesized as duplex oligonucleotides. These are ligated together to form multimers, which are then attached to a solid substrate in a column. The desired DNA-binding protein can then be isolated by affinity chromatography, using the binding site in DNA as the affinity ligand.
b. The isolated, functional repressor is a tetramer; each of the four monomers is the product of the lacI gene (i.e. it is a homotetramer).
c. The DNA-binding domainof the lac repressor folds into a helix-turn-helixdomain. We will examine this structural domain in more in Chapter III. It is one of the most common DNA-binding domains in prokaryotes, and a similar structural domain (the homeodomain) is found in some eukaryotic transcriptional regulators.
3. Contact points between repressor and operator
a. Investigation of the contact points between repressor and the operator utiblized the same techniques that we discussed previously for mapping the binding site of RNA polymerase on the promoter, e.g. electrophoretic mobility shift assays (does the DNA fragment bind?), DNase footprints (where does the protein bind?) and methylation interference assays (methylation of which purines will prevent binding?). Alternative schemes will allow one to identify sites at which methylation is either prevented or enhanced by the binding of the repressor. These techniques provide a biochemical defintion of the operator = binding site for repressor.
b. The key contact points (see Figure 4.1.4.):
(1)are within the dyad symmetry.
(2)coincide (in many cases) with nucleotides that when mutated lead to constitutive expression. Note that the latter is a genetic definition of the operator, and it coincides with the biochemically-defined operator.
(3)tend to be distributed symmetrically around the dyad axis (+11).
(4)are largely on one face of the DNA double helix.
c. The partial overlap between the operator and the promoter initially suggested a model of steric interference to explain the mechanism of repression. As long a repressor was bound to the operator, the polymerase could not bind to the promoter. But, as will be explored in the next chapter, this is notthe case. RNA polymerase canbind to the lacpromoter even when repressor is boudn to the lac operator. However, the polymerase cannot initiatetranscription when juxtaposed to the repressor.
4. Conformational shift in repressor when inducer binds
a. The repressor has two different domains, one that binds to DNA ("headpiece" containing the helix-turn-helix domain) and another that binds to the inducer (and other subunits) (called the "core). These are connected by a "hinge" region.
b. These structural domains can be distinguished by the phenotypes of mutations that occur in them.
lacI-dprevents binding to DNA, leads to constitutive expression.
lacISprevents binding of inducer, leads to a noninducible phenotype.
c. Binding of inducer to the "core" causes an allosteric shift in the repressor so that the "headpiece" is no longer able to form a high affinity complex with the DNA, and the repressor can dissociate (go to one of the many competing nonspecific sites).
Positive control: "catabolite repression"
1. Catabolite repression
a. Even bacteria can be picky about what they eat. Glucose is the preferred source of carbon for E. coli; the bacterium will consume the available glucose before utilizing alternative carbon sources, such as lactose or amino acids.
b. Glucose leads to repression of expression of lacand some other catabolic operons. This phenomenon is called catabolite repression.
2. Two components are needed for this form of regulation
a. cAMP
[1]In the presence of glucose, the [cAMP] inside the cell decreases from 10-4 M to 10-7 M. A high [cAMP] will relieve catabolite repression.
[2]cAMP synthesis is catalyzed by adenylate cyclase (product of the cyagene)
ATP ® cAMP + PPi
b. Catabolite Activator Protein = CAP
[1]Product of the capgene, also called crp(cAMP receptor protein).
[2]Is a dimer
[3]Binds cAMP, and then the cAMP-CAP complex binds to DNA at specific sites
3. Binding site for cAMP-CAP
a. In the lac operon, the binding site is a region of about 20 bp located just upstream from the promoter, from -52 to -72.
b. The pentamer TGTGA is an essential element in recognition. For the lac operon, the binding site is a dyad with that sequence in both sides of the dyad.
c. Contact points betwen cAMP-CAP and the DNA are close to or coincident with mutations that render the lacpromoter no longer responsive to cAMP-CAP.
d. cAMP-CAP binds on one face of the helix.
Figure 4.1.5. Binding site for cAMP-CAP
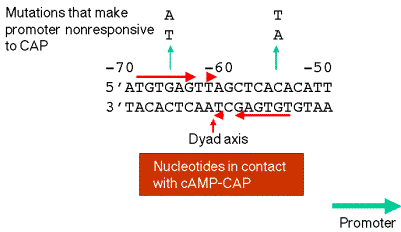
4. Binding of cAMP-CAP to its site will enhance efficiency of transcription initiation at promoter
a. The lacpromoter is not a particularly strong promoter. The sequence at -10, TATGTT, does not match the consensus (TATAAT) at two positions.
b. In the presence of cAMP-CAP, the RNA polymerase will initiate transcription more efficiently.
c. The lacUV5 promoter is an up-promoter mutation in which the -10 region matches the consensus. The lac operon driven by the UV5 promoter will achieve high level induction without cAMP-CAP, but the wild-type promoter requires cAMP-CAP for high level induction.
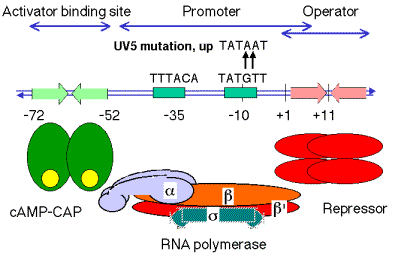
5. Mode of action of cAMP-CAP
a. Direct positive interaction with RNA polymerase. The C-terminus of the a subunit is required for RNA polymerase to be activated by cAMP-CAP. This will be explored in more detail in Chapter 16.
b. cAMP-CAP bends the DNA about 90o.
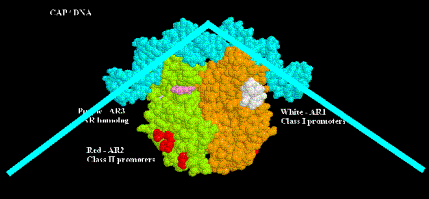
Some generalities
- Repressors, activators and polymerases interact primarily with one face of the DNA double helix.
- Regulatory proteins, such as activators and repressors, are frequently symmetrical and bind symmetrical sequences in DNA.
- RNA polymerases are not symmetrical, and the promoters to which they bind also are asymmetrical. This confers directionality on transcription.


