8.7: Double-strand-break model for Recombination
- Page ID
- 354
Several lines of evidence, primarily from studies of recombination in yeast, required changes to the reciprocal exchange of DNA chains initiated at single-strand nicks. As just mentioned, one DNA duplex tended to be the donor of information and the other the recipient, in contrast to the equal exchange predicted by the original Holliday model. Also, in yeast, recombination could be initiated by double-strand breaks. For instance, both DNA strands are cleaved (by the HO endonuclease) to initiate recombination between the MATand HML(R) loci in mating type switching in yeast. Using plasmids transformed into yeast, it was shown that a double-strand gap in the “aggressor” duplex could be used to initiate recombination, and the gap was repaired during the recombination (this experiment is explored in problem 8.___). In this case, the gap in one duplex was filled by DNA donated from the other substrate. All this evidence was incorporated into a major new model for recombination from Jack Szostak and colleagues in 1983. It is called the double-strand-break model. New features in this model (contrasting with the Holliday model) are initiation at double-strand breaks, nuclease digestion of the aggressor duplex, new synthesis and gap repair. However, the fundamental Holliday junction, branch migration and resolution are retained, albeit with somewhat greater complexity because of the additional numbers of Holliday junctions. Although many aspects of the recombination mechanism differ
The steps in the double-strand-break model up to the formation of the joint molecules are diagrammed in Figure 8.9.
- An endonuclease cleaves both strands of one of the homologous DNA duplexes, shown as thin blue lines in Figure 8.9. This is the aggressor duplex, since it initiates the recombination. It is also the recipientof genetic information, as will be apparent as we go through the model.
- The cut is enlarged by an exonuclease to generate a gap with 3' single-stranded termini on the strands.
- One of the free 3' ends invades a homologous region on the other duplex (shown as thick red lines), called the donor duplex. The formation of heteroduplex also generates a D-loop(a displacement loop), in which one strand of the donor duplex is displaced.
- The D-loop is extended as a result of repair synthesis primed by the invading 3' end. The D-loop eventually gets large enough to cover the entire gap on the aggressor duplex, i.e. the one initially cleaved by the endonuclease. The newly synthesized DNA uses the DNA from the invaded DNA duplex (thick red line) as the template, so the new DNA has the sequence specified by the invaded DNA.
- When the displaced strand from the donor (red) extends as far as the other side of the gap on the recipient (thin blue), it will anneal with the other 3' single stranded end at that end of the gap. The displaced strand has now filled the gap on the aggressor duplex, donating its sequence to the duplex that was initially cleaved. Repair synthesiscatalyzed by DNA polymerase converts the donor D-loop to duplex DNA. During steps 4 and 5, the duplex that was initially invaded serves as the donorduplex; i.e. it provides genetic information during this phase of repair synthesis. Conversely, the aggressor duplex is the recipient of genetic information. Note that the single strand invasion models predict the opposite, where the initial invading strand is the donor of the genetic information.
- DNA ligase will seal the nicks, one on the left side of the diagram in Figure 8.9 and the other on the right side. Although the latter is between a strand on the bottom duplex and a strand on the top duplex, it is equivalent to the ligation in the first nick (the apparent physical separation is an artifact of the drawing). In both cases, sealing the nick forms a Holliday junction.
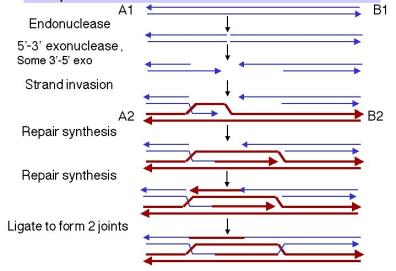
At this point, the recombination intermediate has two recombinant joints (Holliday junctions). The original gap in the aggressor duplex has been filled with DNA donated by the invaded duplex. The filled gap is now flanked by heteroduplex. The heteroduplexes are arranged asymmetrically, with one to the left of the filled gap on the aggressor duplex and one to the right of the filled gap on the donor duplex. Branch migration can extend the regions of heteroduplex from each Holliday junction.
The recombination intermediate can now be resolved. The presence of two recombination joints adds some complexity, but the process is essentially the same as discussed for the Holliday model. Each joint can be resolved horizontally or vertically. The key factor is whether the joints are resolved in the same mode or sense (both horizontally or both vertically) or in different modes.
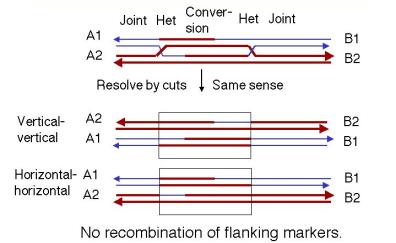
If both joints are resolved the same sense (Figure 8.10), the original duplexes will be released, each with a region of altered genetic information that is a "footprint" of the exchange event. That region of altered information is the original gap, plus or minus the regions covered by branch migration. For instance, if both joints are resolved by cutting the originally cleaved strands ("horizontally" in our diagram of the Holliday model), then you have no crossover at either joint. If both joints are resolved by cleaving the strands not cut originally ("vertically" in our diagram of the Holliday model), then you have a crossover at both joints. This closely spaced double crossover will produce no recombination of flanking markers.
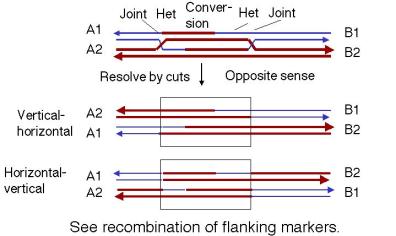
In contrast, if each joint is resolved in opposite directions (Figure 8.11), then there will be recombination between flanking markers. That is, one joint will not give a crossover and the other one will.
Several features distinguish the double-strand-break model from the single-strand nick model initially proposed by Holliday. In the double-strand-break model, the region corresponding to the original gap now has the sequence of the donor duplex in both molecules. This is flanked by heteroduplexes at each end, one on each duplex. Hence the arrangement of heteroduplex is asymmetric; i.e. there is a different heteroduplex in each duplex molecule. Part of one duplex molecule has been converted to the sequence of the other (the recipient, initiating duplex has been converted to the sequence of the donor). In the single strand invasion model, each DNA duplex has heteroduplex material covering the region from the initial site of exchange to the migrating branch, i.e. the heteroduplexes are symmetric. In variations of the model (Meselson-Radding) in which some DNA is degraded and re‑synthesized, the initiating chromosome is the donor of the genetic information.
These models also have many important features in common. Steps that are common to all the models include the generation of a single strand of DNA at an end, a search for homology, strand invasion or strand exchange to form a joint molecule, branch migration, and resolution. Enzymes catalyzing each of these steps have been isolated and characterized. This is the topic of the rest of this chapter.


