8.6: Holliday Model for General Recombination - Single Strand Invasion
- Page ID
- 358
In 1964, Robin Holliday proposed a model that accounted for heteroduplex formation and gene conversion during recombination. Although it has been supplanted by the double-strand break model (at least for recombination in yeast and higher organisms), it is a useful place to start. It illustrates the critical steps of pairing of homologous duplexes, formation of a heteroduplex, formation of the recombination joint, branch migration and resolution.
The steps in the Holliday Model are illustrated in Figure 8.6.
- Two homologous chromosomes, each composed of duplex DNA, are paired with similar sequences adjacent to each other.
- An endonuclease nicks at corresponding regions of homologous strands of the paired duplexes. This is shown for the strands with the arrow to the right in the figure.
- The nicked ends dissociate from their complementary strands and each single strand invades the other duplex. This occurs in a reciprocal manner to produce a heteroduplex regionderived from one strand from each parental duplex.
- DNA ligase seals the nicks. The result is a stable joint molecule, in which one strand of each parental duplex crosses over into the other duplex.This X-shaped joint is called a Holliday intermediate or Chi structure.
- Branch migration then expands the region of heteroduplex. The stable joint can move along the paired duplexes, feeding in more of each invading strand and extending the region of heteroduplex.
- The recombination intermediate is then resolved by nicking a strand in each duplex and ligation.
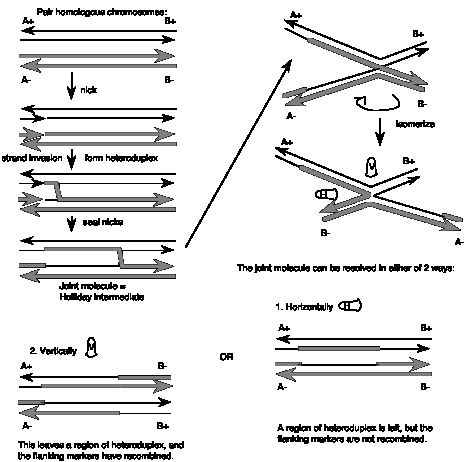
Resolution can occur in either of two ways, only one of which results in an exchange of flanking markers after recombination. The two modes of resolution can be visualized by rotating the duplexes so that no strands cross over each other in the illustration (Figure 8.6). In the “horizontal” mode of resolution, the nicks are made in the same DNA strands that were originally nicked in the parental duplexes. After ligation of the two ends, this produces two duplex molecules with a patch of heteroduplex, but no recombination of flanking regions. In contrast, for the “vertical” mode of resolution, the nicks are made in the other strands, i.e. those not nicked in the original parental duplexes. Ligation of these two ends also leaves a patch of heteroduplex, but additionally causes recombination of flanking regions. Note that “horizontal” and “vertical” are just convenient designations for the two modes based on the two-dimensional drawings that we can make. The important distinction in terms of genetic outcome is whether the resolution steps target the strands initially cleaved or the other strand.
The steps in this model of general recombination can be viewed in a dynamic form by visiting a web site maintained by geneticists at the University of Wisconsin (URL is www.wisc.edu/genetics/Holliday/index.html). This shows the steps in the Holliday model in a movie, illustrating the actions much more vividly than static diagrams.
The recombinant joint proposed by Holliday has been visualized in electron micrographs of recombining DNA duplexes (Figure 8.7A). It has the proposed X shape. Although this joint is drawn with some distance between the duplexes in illustrations, in fact the two duplexes are juxtaposed, and only a very few base pairs are broken in the Holliday intermediate (Figure 8.7B). The structure is symmetrical , and it is likely that the choice between “horizontal” and “vertical” resolution is a random event by the resolving nuclease. It chooses two strands, but it cannot tell which were initially cleaved and which were not.
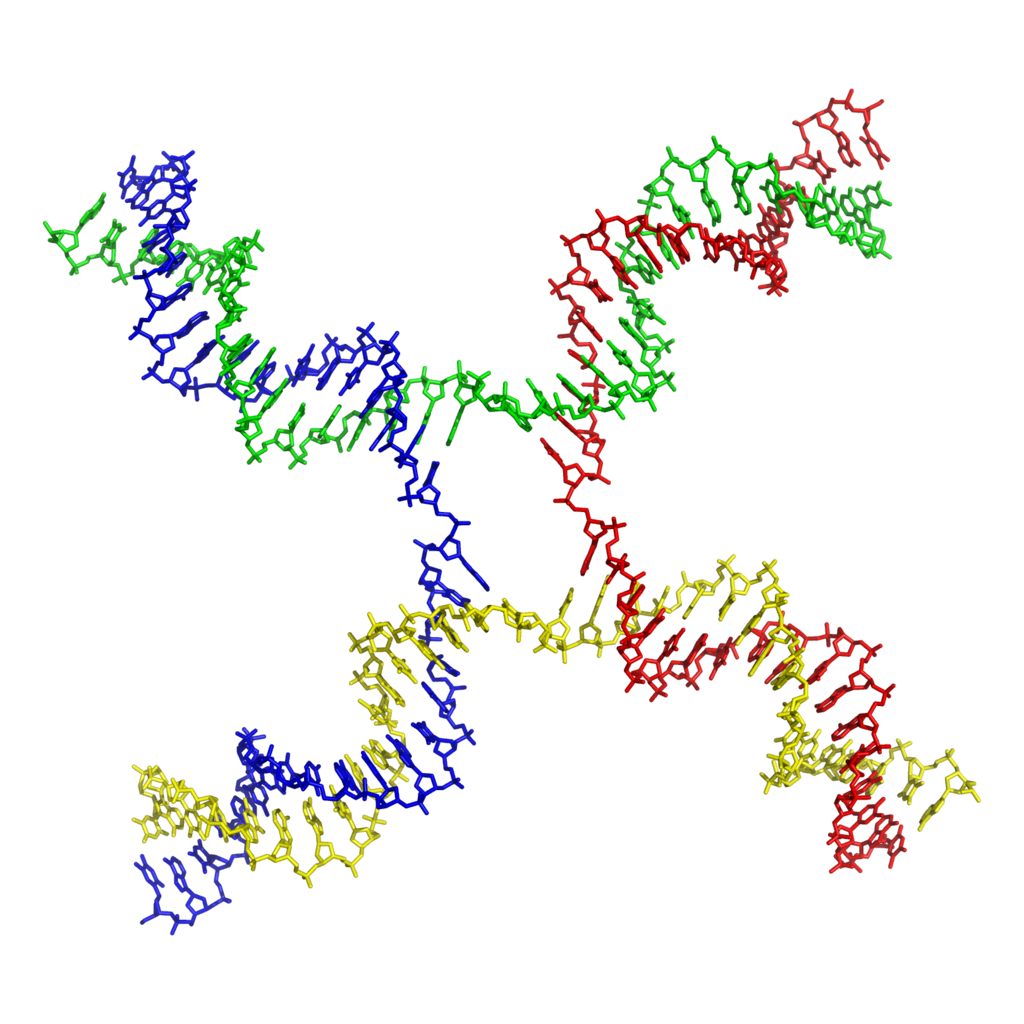

Studies of recombination between chromosomes with limited homology have shown that the minimum length of the region required to establish the connection between the recombining duplexes is about 75 bp. If the homology region is shorter than this, the rate of recombination is substantially reduced.
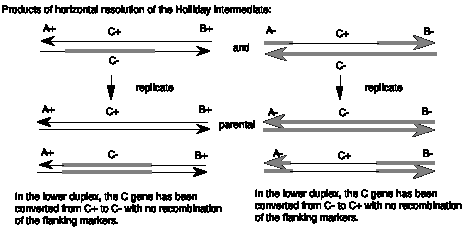
The patch of heteroduplex can be replicated (Figure 8.8) or repaired to generate a gene conversion event. As shown in Figure 8.8, replication through the products of horizontal resolution (from Figure 8.6) will generate a duplex from each strand of the heteroduplex. If we consider the parental chromosomes to be A+C+B+ and A-C-B-, and the heteroduplex to be in gene C, the products of replication can have a the parental C+ converted to a C- but still flanked by A+ and B+ or C- converted to C+ but still flanked by A- and B-. In either case, gene C has changed to a new allele without affecting the flanking markers.
Although the original Holliday model accounted for many important aspects of recombination (all that were known at the time), some additional information requires changes to the model. For instance, the Holliday model treats both duplexes equally; both are the invader and the target of the strand invasion. Also, no new DNA synthesis is required in the Holliday model. However, subsequent work showed that one of the duplex molecules is the used preferentially as the donor of genetic information. Hence additional models, such as one from Meselson and Radding, incorporated new DNA synthesis at the site of the nick to make and degradation of a strand of the other duplex to generate asymmetry into the two duplexes, with one the donor the other the recipient of DNA. These ideas and others have been incorporated into a new model of recombination involving double strand breaks in the DNAs.


