5.03: A. Oxygen-Binding Proteins and Allosterism
- Page ID
- 158384
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)-
Structural Comparison and Analysis
- Identify the structural features of myoglobin versus hemoglobin, including the number and arrangement of α‑helices, subunit organization (monomeric vs. tetrameric), and the positioning of the heme group.
- Analyze how the protein fold (globin fold) contributes to the functions of oxygen binding and storage.
-
Ligand Binding Mechanisms
- Explain the nature of ligand binding to heme, including the differences between covalent, coordinate (dative) covalent, and noncovalent interactions.
- Describe how small gaseous ligands (e.g., O₂, CO₂, H⁺) interact with the Fe²⁺ ion in the heme via coordinate bonds and how this contrasts with other ligand types (such as substrates and inhibitors).
-
Electronic and Molecular Orbital Considerations
- Interpret the electronic configuration of Fe²⁺ in heme and understand how its d orbitals are involved in ligand binding.
- Apply crystal field and ligand field theories to explain the changes in electronic structure (e.g., high spin vs. low spin states) and the resulting effects on protein color and function.
-
Cooperative Binding and Allosterism
- Define cooperative binding and allosteric regulation in hemoglobin, and contrast these with the binding properties of myoglobin.
- Describe the effects of allosteric ligands (H⁺, CO₂, and bisphosphoglycerate) on the oxygen binding curve of hemoglobin and explain the Bohr effect.
-
Mathematical Modeling of Ligand Binding
- Derive and analyze the hyperbolic binding equation for myoglobin and understand why its oxygen binding curve is hyperbolic.
- Explain the derivation and application of the Hill equation for hemoglobin, including the significance of the Hill coefficient (n) in modeling cooperativity.
- Compare and contrast the Hill model with more mechanistic models such as the Monod-Wyman-Changeux (MWC) and the Koshland-Nemethy-Filmer (KNF) models, discussing how these models account for the sigmoidal binding curves observed in hemoglobin.
-
Conformational Changes and Quaternary Structure
- Examine the structural transitions between the T (tense/deoxy) and R (relaxed/oxy) states of hemoglobin and how these transitions affect oxygen affinity.
- Evaluate the role of subunit interfaces, salt bridges, and hydrogen bonding in stabilizing these conformational states.
-
Clinical and Evolutionary Implications
- Analyze the molecular basis of hemoglobin variants, particularly the mutation causing sickle cell disease, and understand how a single amino acid substitution can alter protein behavior and lead to pathology.
- Explore the evolutionary relationship between hemoglobin mutations and diseases (e.g., the protective effect against malaria) and discuss current therapeutic approaches (including gene editing strategies).
-
Integration of Interactive Structural Models
- Utilize interactive 3D molecular visualization tools (such as iCn3D) to explore the structures of myoglobin and hemoglobin, focusing on the spatial arrangement of the heme group, ligand-binding sites, and key residues involved in protein function.
By achieving these goals, students will not only reinforce their understanding of protein structure and function but also connect molecular details with physiological and clinical phenomena.
Myoglobin, Hemoglobin, and Their Ligands
Almost all biochemistry textbooks describe proteins' biological functions by using myoglobin and hemoglobin as exemplars. This approach seems rational, as these proteins have become model systems for describing the binding of simple ligands, such as dioxygen (O2), CO2, and H+, and how the binding of ligands determines and influences a protein's structure.
Yet in most ways, these globin-binding "ligands" are dissimilar to the majority of both small ligands, such as substrates (for enzymes), inhibitors, and activators, as well as large "ligands", such as other proteins, nucleic acids, carbohydrates, and lipids that bind to proteins through noncovalent interactions (described in detail in Chapter 2.4). In contrast, dioxygen (O2), CO2, and H+ bind reversibly but through covalent interactions. Dioxygen binds to a heme Fe2+ transition metal through a coordinate covalent or dative bond, and protons bind covalently to proton acceptors (Lewis bases like histidine). In contrast, CO2 binds covalently as it forms a carbamate with the N terminus of a hemoglobin chain. In typical covalent bonds, each bonded atom contributes to and shares the two electrons. In coordinate or dative covalent bonds, the ligand, a Lewis base, contributes both electrons in the bond. For simple analyses and counting electrons, both electrons can be considered "owned" by the ligand and not by the transition metal ion, a Lewis acid, unless you analyze the interactions using molecular orbital (ligand field) theory. Hence, the ligand can readily dissociate from the metal ion, like a ligand bound through classical noncovalent interactions. This analogy can be extended to protons, which are also Lewis acids (with no contributing electrons), as they react with Lewis bases (lone pair donors) on atoms such as nitrogen on a histidine side chain. H+ readily leaves (ionizes) from a Lewis acid if the pH of the microenvironment is conducive to ionization.
Even though we disagree with starting the discussion of protein structure and function with the covalent binding of small gaseous and marginally soluble ligands to myoglobin and hemoglobin, we will do so to make the book consistent with most other texts and allow easier use without shuffling the order of chapters.
Let's start with myoglobin (Mb), a monomeric protein containing 8 α−helices (A-H), and with hemoglobin, a heterotetramer with two α -and two β−subunits, each of which also contains eight α−helices. Both are oxygen-binding proteins. Both contain heme (one in myoglobin and four in the four subunits of hemoglobin). Each heme has a central Fe2+ ion, which forms a coordinate covalent bond with dioxygen. Dioxygen is transported from the animals' lungs, gills, or skin to the capillaries, where it can be delivered to respiring tissue. O2 has a low solubility in blood (0.1 mM). Whole blood contains 150 g Hb/L and can achieve a dissolved oxygen concentration of 10 mM. Invertebrates can have alternative proteins for oxygen binding, including hemocyanin, which contains Cu, and hemerythrin, a non-heme protein. On binding dioxygen, solutions of Hb change color to bright red. Solutions of hemocyanin and hemerythrin change to blue and burgundy colors, respectively, on binding dioxygen. Some Antarctic fish don't require Hb since dioxygen is more soluble at low temperatures. Myoglobin is found in the muscles and serves as a storage protein for oxygen transported by hemoglobin.
Figure \(\PageIndex{1}\) shows the structure of heme in both myoglobin and hemoglobin.
The heme group contains protoporphyrin IX, with four tetrapyrrole rings linked by methene bridges. Four methyl, two vinyl, and two propionate groups are attached to the tetrapyrrole structure. These can be arranged in 15 ways, of which only one (IX) occurs in biological systems. Protoporphyrin IX with bonded ferrous (Fe2+) iron is called heme and has a nitrogen atom on each of the four pyrrole rings that form a coordinate covalent bond to Fe2+. The heme fits into a hydrophobic crevice in heme-binding proteins with the propionate groups exposed to solvent.
Myoglobin (Mb)
Myoglobin is an extremely compact protein with 75% alpha-helical structure. It has eight alpha helices labeled A-H. A proline, a helix breaker, terminates four. The interior amino acids are almost entirely nonpolar. The only polar amino acids found completely buried are two histidines. One is called the proximal His, nearer the heme, and serves as the 5th ligand to the heme Fe2+. The other is called the distal His, which is too far to coordinate the heme Fe2+. This last potential 6th ligand-binding site forms a coordinate covalent bond with O2 in oxy-myoglobin.
Figure \(\PageIndex{2}\) shows an interactive iCn3D model of deoxymyoglobin from wild boar. The heme is shown in sticks along with the proximal and distal histidines.
.png?revision=1&size=bestfit&width=242&height=238)
Hemoglobin
Hemoglobin has an illustrious history. It is the first protein whose molecular weight was determined and the first to be assigned a specific function (dioxygen transport). It was the first protein in which a mutation in a single amino acid caused by a single base pair change in the DNA coding sequence was shown to cause a disease (sickle cell trait and disease). The mathematical theories developed to model dioxygen binding are used to explain enzyme activity. It also binds H+, CO2, and bisphosphoglycerate at sites (allosteric) distant from the oxygen binding site. This binding then regulates dioxygen binding affinity.
As with myoglobin, the Fe2+ ion is coordinated to 4 Ns on the four pyrrole rings. The 5th ligand is supplied by the proximal His, the 8th amino acid on helix F of the protein. In the absence of dioxygen, the 6th ligand is missing and the geometry of the complex is somewhat square pyramidal, with the Fe slightly above (0.2 Å) the plane of the heme ring. A distal His (E7) is on the opposite side of the heme ring but too far to coordinate with the Fe2+. When dioxygen binds, it occupies the 6th coordination site and pulls the Fe into the plane of the ring, leading to octahedral geometry. The animated image below shows these changes that occur on oxygenation.
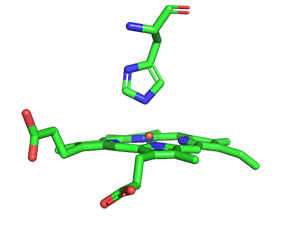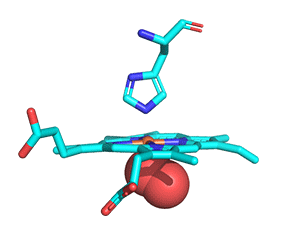
The proximal histidine that provides the imidazole nitrogen ligand is shown. Dioxygen is shown as red spheres. The Fe2+ ion is shown as a small orange sphere. Its size has been dramatically reduced in this image, so its movement can be more readily observed. Note that Fe2+ is centered a small distance above the heme plane in deoxyHb but in the plane in oxyHb.
Figure \(\PageIndex{3}\) shows interactive iCn3D models of a single A chain of deoxyhemoglobin (2HHB) and oxyhemoglobin (1HHO) in the top row, and just the heme, distal and proximal histidines, along with O2 for oxyhemoglobin in the bottom row.
| A chain of human deoxyhemoglobin (2HHB) |
A chain of human oxyhemoglobin (1HHO) |
|
(Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...odtS7K7Raj3EZ7 |
(Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...nm2Am9A5SKa9w9 |
|
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...hUd6hfDKpk5t98 |
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...7nJTQ24QA5JLKA |
Figure \(\PageIndex{3}\): interactive iCn3D models of a single A chain of deoxyhemoglobin (2HHB) and oxyhemoglobin (1HHO) in the top row, and just the heme, distal and proximal histidines, along with O2 for oxyhemoglobin in the bottom row.
Rotate the models in the bottom row to see the subtle changes in the heme and relative position of the Fe2+ ion in the heme ring.
Carbon monoxide (CO), nitric oxide (NO), and hydrogen sulfide (H2S) also bind to the sixth coordination site, but with higher affinity than dioxygen, which can lead to CO poisoning, for example. The distal histidine causes these ligands (including dioxygen) to be bound in a bent, non-optimal geometry. This minimizes the chances of CO poisoning.
Figure \(\PageIndex{4}\) shows an interactive iCn3D model of human oxy-hemoglobin (2dn1)

Fe2+ ion ligand interactions
When the 6th ligand, dioxygen, binds to heme Fe2+, the geometry of the complex becomes octahedral. The Fe2+ ion has 6 electrons in d orbitals. The electronic configuration of atomic Fe is 3d64s2, while the Fe2+ ion has a 3d6 configuration, as shown in Figure \(\PageIndex{5}\). Each of the orbitals would have the same energy except for the doubly occupied one, which would have slightly higher energy due to the extra repulsion of the two electrons in the orbital. This effect is minimal, so to a first approximation, the orbitals are considered to have the same energies (degenerate). The figure below shows them having the same energy
From introductory chemistry classes, you will remember that transition metal complexes and their solutions are highly colored. Since oxygenated hemoglobin (found in arteries) appears bright red/orange, it must absorb blue/green light more than deoxyhemoglobin, which is a darker red (but still reddish). These absorbed wavelengths are removed from the spectrum, making hemoglobin shades of red. Veins contain more deoxygenated blood returning to the heart to be reoxygenated. The visible veins in your arms and legs appear blue, but not because of the spectral properties of deoxyhemoglobin. Rather, blue light doesn't penetrate the tissue as far as red, so red is preferentially removed from the remaining light, which is reflected, making veins appear blue. Figure \(\PageIndex{6}\) shows a partial absorbance spectrum of deoxy- and oxyhemoglobin from 280-1000 nm.
The dashed light blue vertical line shows the approximate wavelength (around 450 nm) of the largest molar extinction coefficient difference between the two forms. Note that the y-axis is a log scale. At 450 nm, oxyhemoglobin has an extinction coefficient of about 600,000, while deoxyhemoglobin is about 60,000, so much more blue light is removed from a solution of oxyhemoglobin, making it appear bright red. Note also that the same differences appear in the red region of the spectra, where the extinction coefficients only vary from 3000 for oxyhemoglobin to 200 for deoxyhemoglobin. Hence, absorption in this region has little effect on the visible color of blood.
The spectrum shown in Figure \(\PageIndex{6}\) also shows the near-infrared region (denoted by a rectangle) of the spectra. Many may use inexpensive pulse oximeters (some built into watches) to measure their oxygenation status during the COVID-19 pandemic. These use two pulsed LEDs, one at 660 nm, where oxyhemoglobin has a higher extinction coefficient, and one at 940 nm (infrared region), where deoxyhemoglobin has a higher extinction coefficient. Inexpensive pulse oximeters overestimate the oxygenation state of people with darker skin tones and put such people's health at risk from overestimated values.
Binding of ligand to the heme Fe2+: crystal and ligand fields
Most biochemistry books offer minimal coverage of bioinorganic chemistry, even though many proteins bind metal ions. Since biochemistry is an interdisciplinary field, it is important to use past learning in other biology and chemistry courses and apply it to biochemistry. Most students study transition metal chemistry in introductory chemistry courses. You should be familiar with transition metal ions, their electronic configuration, crystal field theory, high and low spin states, paramagnetism, and diamagnetism. Hence, it is appropriate and important to bring these ideas into biochemistry and extend them when necessary. The basis of the material below is modified from Structure & Reactivity in Organic, Biological, and Inorganic Chemistry by Chris Schaller (Creative Commons Attribution-NonCommercial 3.0 Unported License).
Let's look at the electronic structure of Fe2+ in oxyhemoglobin. The Fe2+ is coordinated to six ligands (4 pyrrole rings of the heme in a plane, one axial imidazole ring each from the proximal histidine and bonded dioxygen). The geometry of the electron clouds around the Fe2+ is octahedral. The geometries, shapes, and orientations of the five Fe2+ d atomic orbitals are illustrated in Figure \(\PageIndex{7}\).
Two orbitals, dz2 and dx2-y2, appear different because they are oriented directly along the x, y, and z axes. The other three orbitals are oriented in between the axes. Now imagine six anionic ligands with lone pairs approaching along the axes of the Fe2+ atomic orbitals so they could directly interact with the dz2 and dx2-y2 orbitals, a postulate of crystal field theory used to explain bonding in transition metal complexes. The energy of the Fe electrons in the dz2 and dx2-y2 atomic orbitals would increase due to electron-electron repulsion. This is illustrated in Figure \(\PageIndex{8}\). Two different outcomes can result from these interactions.
1 (left panel): If the electrostatic interaction between the atomic orbitals of the Fe ion and the incoming ligands is weak, the energy of the dz2 and dx2-y2 orbitals would be a bit higher (by an amount Δ) due to the repulsion along those axes from anions oriented along them. This is illustrated in the left part of Figure \(\PageIndex{8}\). When filling the orbitals with the six Fe2+ 3d6 electrons, add one electron to each of the five orbitals and then pair one for the sixth orbital. In that case, there would be four unpaired electrons, and the complex would be paramagnetic. We call this a high-spin state.
2 (right panel): If, however, the electrostatic interactions of the incoming ligands with the d atomic orbital electrons are strong, the Δ would be large. When filling these orbitals with the six Fe2+ 3d6 electrons, the electrons would be paired in the three lower energy orbitals. There would be no electrons unpaired, so the ion is diamagnetic. This is the low-spin case.
In either case, if light of energy equal to the Δ interacts with the metal ion, electrons would be promoted to the higher energy level. For the low spin case (larger axial electrostatic interactions, Δ is large, and the energy of the required photon is larger (more blue shifted) than in the high spin case. This could make a solution of the molecule in the low spin case appear redder than in the high spin case (as blue light is removed on absorption). This is the case for dioxygen, which interacts strongly with the Fe2+ axially oriented orbitals. Hence, oxy-Fe2+ heme complex is low spin and diamagnetic. A solution would be red due to the removal of blue photons by absorption.
You studied basic crystal field theory in introductory chemistry courses. The theory is simplistic in several ways. The ligands might not be anions. More importantly, bonding is best described by molecular orbitals. A more comprehensive ligand field theory considers the effect of the donor electrons and the d orbitals of the transition metal ion. These atomic orbitals combine to produce molecular orbitals (MOs), better describing bonding.
Take the simple case of a covalent bond between two singly occupied adjacent p atomic orbitals between the two carbons of ethylene (C2H4). Simplistically, you could imagine two carbon atoms with their two p orbitals approaching each other. The two p orbitals (or, more accurately, their wave functions) could combine constructively or destructively to form two new molecular orbitals (MO). One is a pi bonding MO, π, which is lower in energy (promoting bond formation) than the atomic p orbitals. The other is a pi antibonding MO, π*, which is higher in energy (antagonizing bond formation), as shown in Figure \(\PageIndex{9}\). Two atomic orbitals form from two MOs!
Let's use ligand field theory with MOs to describe ligands binding to the heme Fe2+ d orbitals. Figure \(\PageIndex{10}\) illustrates this in the MO diagram.
Figure \(\PageIndex{10}\): Ligand field molecular orbtials for the d6 Fe2+ ion for two axial ligands
For simplicity, for Fe2+, we will consider only d orbitals and focus on the dz2 and dx2-y2 (also called the eg) orbitals, since these are most affected by the ligands as described in crystal field theory. Let's assume these interact with lone pairs (a simplistic assumption as well) in two ligand orbitals, one on the dioxygen and one on the proximal histidine imidazole N. Seven atomic orbitals (five Fe d and two ligand orbitals) combine to produce 7 MOs. Since the orbitals (in sp2- or sp3-like orbitals) on the ligands are closer in energy to the lower energy bonding MOs, their electrons would go there. Since O2 interacts strongly with the Fe2+ d orbitals, the system is in a low-spin state, with the unoccupied dz2 and dx2-y2 (eg orbitals) now considered antibonding orbitals. The fully occupied dxy, dxz, and dyz orbitals are considered nonbonding since they have the same relative energy as atomic orbitals.
We will return to molecular orbitals occasionally throughout this book when they offer the best explanation for biological events.
Here are some important things to note. When dioxygen binds to the heme iron, the oxidation state of the Fe2+ ion does not change, even though dioxygen is a great oxidizing agent. Hence, the Fe2+ ion is a reversible carrier of dioxygen and not electrons. Free heme in solution is oxidized by dioxygen, forming a complex with water, which occupies the 6th position, with the iron in the Fe3+ state. An intermediate in this process is the formation of a dimer of 2 hemes linked by one dioxygen. This can't occur readily when the heme is in Hb or Mb. Other heme proteins (like Cytochrome C), which we will explore in future chapters, are designed to be carriers of electrons. A small amount of the Fe2+ ion is oxidized to Fe3+ ion in myoglobin and hemoglobin, resulting in met-Hb and met-Mb. The brown color of old meat results in large part from Met-Mb. An enzyme is required to reduce the iron back to the Fe2+ state.
The differences between hemoglobin and myoglobin are equally important. Hemoglobin is a heterotetramer of two α and two β subunits held together by noncovalent interactions (an example of quaternary protein structure), with four bound hemes, each of which can bind a dioxygen. In a fetus, two other subunits make up hemoglobin (two zeta - ζ and two epsilon - ε subunits -analogous to the two α and two β subunits, respectively). These subunits change during development to two α and two γ subunits. Fetal hemoglobin has a higher affinity for dioxygen than adult hemoglobin. Myoglobin is a single polypeptide chain with a higher affinity for dioxygen than hemoglobin.
The α and β chains of hemoglobin are similar to those of myoglobin, which is unexpected since only 24 of 141 residues in the α and β chains of Hb are identical to amino acids in myoglobin. This suggests that different sequences can fold into similar structures. The globin fold of myoglobin and each chain of hemoglobin is common to vertebrates and must be nature's design for dioxygen carriers. A comparison of the sequence of hemoglobin from 60 species shows much variability of amino acids, with only nine identical amino acids found. These must be important for structure/function. All internal changes are conservative (e.g., changing a nonpolar for a nonpolar amino acid). Not even prolines are conserved, suggesting there are different ways to break helices. The two active site histidines are conserved, as is glycine B6 (required for a reverse turn). http://www.umass.edu/molvis/tutorials/hemoglobin/
Normal and Cooperative Binding of Dioxygen - Structural Analyses
Plots of Y (fractional saturation) vs L (pO2) are hyperbolic for myoglobin (Mb), but sigmoidal for Hb, suggesting cooperative binding of oxygen to Hb (binding of the first oxygen facilitates binding of the second, etc). Figure \(\PageIndex{11}\) shows fractional saturation (Y) binding curves vs dioxygen concentration (PO2) for both myoglobin and hemoglobin.
Note that hemoglobin is saturated with O2 at the high concentration found in the lungs, but it releases much of its bound O2 in respiring tissues in which O2 is much lower. In contrast, myoglobin only releases significant bound oxygen at much lower O2 concentrations. Hence, myoglobin is designed for dioxygen storage.
In another difference, the affinity of Hb for dioxygen, but not Mb, depends on pH. This is called the Bohr effect, after the person who discovered it, Christian Bohr (who was the father of Niels Bohr). Figure \(\PageIndex{12}\) shows binding curves for hemoglobin in the presence of increasing and decreasing concentrations of H+ (pH) as well as for CO2 and another ligand, 2,3-diphosphoglycerate (2,3-DPG).
Michał Komorniczak (Poland). https://commons.wikimedia.org/wiki/F...ohr_Effect.svgCreative Commons 3.0. Attribution-ShareAlike (CC BY-SA 3.0).
Protons (decreasing pH), carbon dioxide, and bisphosphoglycerate, all allosteric ligands that bind distal to the oxygen binding sites on the heme, shift the binding curves of Hb for oxygen to the right, lowering the apparent affinity of Hb for oxygen. The same effects do not occur for Mb. These ligands regulate the binding of dioxygen to Hb.
From these clues, we wish to discern the
- molecular and mathematical bases for the sigmoidal binding curves
- mechanism for the exquisite regulation of O2 binding by allosteric ligands.
The two obvious features that differ between Mb and Hb are the tetrameric nature of Hb and its multiple (4) binding sites for oxygen. Regulation of dioxygen binding is associated with conformational changes in hemoglobin.
Based on crystallographic structures, two main conformational states appear to exist for Hb: the deoxy (or T-taut) state and the oxy (or R-relaxed) state. The major shift in conformation occurs at the alpha-beta interface, where contacts with helices C and G and the FG corner are shifted on oxygenation. Figure \(\PageIndex{13}\) shows conformation changes on O2 binding to deoxyhemoglobin (files aligned with DeepView, displayed with Pymol). Dioxygen is shown as red spheres.
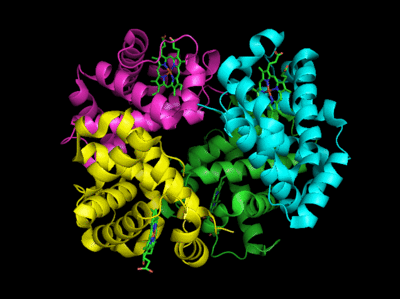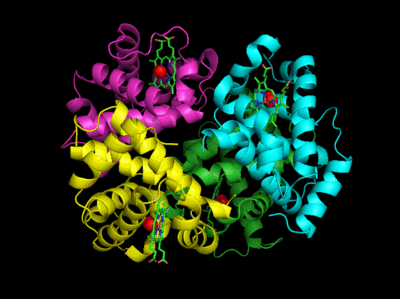
The deoxy or T form is stabilized by eight salt bridges, broken in the transition to the oxy or R state. This is illustrated in Figure \(\PageIndex{14}\).
6 of the salt bridges are between different subunits (as expected from the above analysis), with 4 involving the C- or N-terminus.
In addition, crucial H-bonds between Tyr 140 (alpha chain) or 145 (on the beta chain) and the carbonyl O of Val 93 (alpha chain) or 98 (beta chain) are broken. Crystal structures of oxy and deoxy Hb show that the major conformational shift occurs at the interface between the α and β subunits. When the heme Fe binds oxygen, it is pulled into the plane of the heme ring, causing a shift of about 0.2 nm. This small shift leads to larger conformational changes since the subunits are packed so tightly that compensatory changes in their arrangement must occur. The proximal His (coordinated to the Fe2+) is pulled toward the heme, which causes the F helix to shift, causing a change in the FG corner (the sequence separating the F and G helices) at the alpha-beta interface as well as the C and G helices at the interface, which all slide past each other to the oxy-or R conformation.
Decreasing pH shifts the oxygen binding curves to the right (to decreased oxygen affinity). Increased [proton] will cause protonation of basic side chains. In the pH range for the Bohr effect, the most likely side chain to get protonated is His (pKa around 6), which then becomes charged. The most likely candidate for protonation is His 146 (on the β chain - HC3), which can form a salt bridge with Asp 94 of the β(FG1) chain. This salt bridge stabilizes the positive charge on the His and raises its pKa compared to the oxyHb state. Carbon dioxide binds covalently to the N-terminus to form a negatively charged carbamate, which forms a salt bridge with Arg 141 on the alpha chain. Bisphosphoglycerate (BPG), a strongly negatively charged ligand, binds in a pocket lined with Lys 82, His 2, and His 143 (all on the beta chain). It fits into a cavity between the β subunits of the Hb tetramer in the T state. Notice that all these allosteric effectors lead to the formation of more salt bridges, which stabilize the T or deoxy state. The central cavity where BPG binds between the β subunits becomes much smaller on oxygen binding and the shift to the oxy or R state. Hence, BPG is extruded from the cavity.
The binding of H+ and CO2 helps shift the equilibrium to the deoxyHb form, facilitating the release of oxygen to the tissues. It is in respiring tissues that CO2 and H+ levels are high. CO2 is produced from glucose oxidation through glycolysis and the Krebs cycle. In addition, high levels of CO2 increase H+ levels through the following equilibrium:
\begin{equation}
\mathrm{H}_{2} \mathrm{O}+\mathrm{CO}_{2} \leftrightarrow \mathrm{H}_{2} \mathrm{CO}_{3} \leftrightarrow \mathrm{H}^{+}+\mathrm{HCO}_{3}{ }^{-}
\end{equation}
In addition, H+ increases from weak acids such as pyruvic acid produced in the central metabolic pathway (glycolysis) to produce energy from glucose oxidation.
The binding of CO2 and H+ to hemoglobin serves an additional function: it removes excess CO2 and H+ from the tissues where they build up. When deoxyHb with bound H+ and CO2 reaches the lungs, they dissociate as O2 builds. In the lungs, deoxyHb is converted to oxyHb.
Hemoglobin exhibits allosterism. Allosterism occurs when a regulatory ligand (like CO2 and H+) binds to a site distal to the binding site of a main ligand (like O2) and changes the affinity for the main ligand. We will define two kinds of allosterism:
Homotropic allosterism (or Type I) occurs when a ligand, such as dioxygen, binds to multiple ligand binding sites on the same protein and gives a sigmoidal binding plot as a function of ligand concentration. Multiple binding sites for a main ligand can be found on a multimeric protein with identical or similar subunits (as in the case of hemoglobin, but not myoglobin). In the case of hemoglobin, the main ligand, O2, binds to the same active site in each monomer. This site is called the orthosteric binding site. For the hemoglobin tetramer, the orthosteric site, of course, is the heme Fe2+ ion. Homotropic or Type I allosterism occurs when an allosteric ligand binds at the active sites of the monomeric subunits of the protein. Most texts call this homotropic allosterism, and the ligands homotropic ligands. Hence, O2 and CO are homotropic ligands for hemoglobin.
Heterotropic (or Type II) allosterism occurs when a chemical modulator binds to a site different from the binding site for the main ligand. In doing so, it modulates (activates or inhibits) the binding affinity of the main ligand for the active or orthosteric site and shifts the binding curve for ligand binding. In addition, binding plots at a fixed main ligand concentration with varying modulator concentrations are also sigmoidal. The modulator binds at an allosteric (other) binding site. In the case of hemoglobin, the allosteric modulators are H+, CO2, and BPG. H+ and CO2 binding shift the O2 binding curve in ways that lower the affinity for O2, leading to its release. Most texts call this type of allosterism heterotropic. Protons (H+) and CO2 are heterotropic ligands. We prefer Type I and Type II over the more jargonistic, less intuitive terms homo- and heterotropic.
Mathematical Analysis of Cooperative Binding
How do the sigmoidal dioxygen binding curves for Hb arise? Mathematics can offer clues that complement and extend structural information. At least three models (Hill, MWC, and KNF) have been developed that give rise to sigmoidal binding curves. Remember, sigmoidal curves imply cooperative binding of oxygen to hemoglobin. As oxygen binds, the next oxygen seems to bind with higher affinity (lower KD). We will discuss the mathematics behind two of the models. Both models are routinely applied to binding phenomena that give sigmoidal curves.
Previously, we have shown that chemical and mathematical equations can describe oxygen binding to myoglobin.
\begin{equation}
\mathrm{M}+\mathrm{L} \leftrightarrow \mathrm{ML}
\end{equation}
\begin{equation}
Y=\frac{L}{K_D+L}
\end{equation}
These mathematical equations are those of a hyperbola, where Y is fractional saturation. Let's now explore two models that give sigmoidal curves.
Hill Model
In this model, we base our mathematical analysis on the fact that the stoichiometry of binding is not 1:1, but rather 4 to 1. Perhaps a more useful equation to express the equilibrium would be M + 4L ↔ ML4. We can derive an equation analogous to the one above:
\begin{equation}
Y=\frac{L^4}{K_D+L^4}
\end{equation}
For any given L and KD, a corresponding Y can be calculated. A plot of Y vs L is not hyperbolic but sigmoidal (see the next link below). Hence, we're getting closer to modeling that actual data. However, there is one problem. This sigmoidal curve does not fit the actual oxygen binding curve for Hb well. A better fit may be achieved by altering the exponents in the equation. A more general equation for binding might be M + nL ↔ MLn, which gives the Hill equation:
\begin{equation}
Y=\frac{L^n}{K_D+L^n}
\end{equation}
Some books/sources offer a different Hill equation. The one above is correct. Click the derivation below, which requires only a background from the preceding chapter sections and maybe a bit from Chapter 6.1.
- Derivation
-
Let's consider this reaction:
For a minute, assume n=2, and the reaction was written as M + 2L ↔ ML2. Also, assume that the reaction occurs when M, L, and a second L all collide simultaneously (unlikely) to form the product ML2. From introductory chemistry, you would write the rate equation for just the forward "ternary" reaction as:
ratef = d[ML2]/dt = kf[M][L][L] = kf[M][L]2
The rate of just the reverse reaction would be:
rater = d[ML2]/dt = kr[ML2]
Subtract the two rates to get the net rate. Now let's switch from n= 2 back to just n and proceed without further explanation.
At equilibrium, the forward and reverse rates are equal, so
\begin{equation}
\frac{d\left[M L_n\right]}{d t}=k_f[M][L]^n=k_f[M][L]^n-k_r\left[M L_n\right]=0 \text { at equilibrium }
\end{equation}KD is equal to the ratio of kr/kf, so solving for KD gives
\begin{equation}
K_D=\frac{k_r}{k_f}=\frac{[M][L]^n}{\left[M L_n\right]}
\end{equation}Solving for [MLn] gives
\begin{equation}
\left[M L_n\right]=\frac{[M][L]^n}{K_D}
\end{equation}The fractional saturation is given by
\begin{equation}
Y=\frac{\left[M L_n\right]}{[M]+\left[M L_n\right]}
\end{equation}Plug in the previous equation for [MLn] to obtain the correct form of the Hill Equation.
\begin{equation}
Y=\frac{\frac{[M][L]^n}{K_D}}{[M]+\frac{[M][L]^n}{K_D}}\left(\frac{K_D}{K_D}\right)=\frac{[M][L]^n}{K_D[M]+[M][L]^n}=\frac{[L]^n}{K_D+[L]^n}
\end{equation}QED!
If n is set to 2.8, the theoretical curve of Y vs L gives the best, but still not perfect, fit to the experimental data. Changing the exponent, which reflects the binding stoichiometry, must seem arbitrary. What molecular interpretation could you give to 2.8?
Consider another meaning of the equilibrium described above: M + 4L ↔ ML4.
One interpretation is that all four oxygens bind at once to Hb. Alternatively, the first one binds with some low affinity, which, through associated conformational changes, alters the remaining three sites to very high-affinity sites. This model implies what is described as infinitely cooperative binding of oxygen.
(Notice that the Hill equation becomes: Y = L/[KD + L] when n =1 (as in the case with myoglobin, and in any equilibrium expression of the form M+L↔ML. Remember plots of ML vs L or Y vs L give hyperbolas, with KD = L at Y = 0.5.)
Does KD = L at Y = 0.5? The oxygen concentration at which Y = 0.5 is defined as P50. We can substitute this value into equation 3, which gives an operational definition of KD in terms of P50.
\begin{equation}
\mathrm{Y}=0.5=\frac{\mathrm{P}_{50}^{\mathrm{n}}}{\mathrm{K}_{\mathrm{D}}+\mathrm{P}_{50}^{\mathrm{n}}}
\end{equation}
multiply both sides by 2 give
\begin{equation}
\begin{gathered}
1=\frac{2 \mathrm{P}_{50}^{\mathrm{n}}}{\mathrm{K}_{\mathrm{D}}+\mathrm{P}_{50}^{\mathrm{n}}} \\
\mathrm{K}_{\mathrm{D}}+\mathrm{P}_{50}^{\mathrm{n}}=2 \mathrm{P}_{50}^{\mathrm{n}} \\
\mathrm{K}_{\mathrm{D}}=\mathrm{P}_{50}^{\mathrm{n}}
\end{gathered}
\end{equation}
Note that KD in this case is not the ligand concentration at half-saturation for this equation, as we saw with hyperbolic binding curves.
This gives a modified version of the Hill equation for hemoglobin binding of dioxygen:
\begin{equation}
Y=\frac{L^n}{P_{50}^n+L^n}
\end{equation}
The Hill equation and Hill coefficient, which is empirically determined to obtain the best fit to the binding data, might seem a bit contrived, especially after seeing the more mechanistically and chemically intuitive MWC equation described below. Is it useful in any other circumstance? Yes, it is often used to model more complex, interconnected binding and kinetic pathways that exhibit similar "exquisite" sensitivity to concentrations, resulting in sigmoidal binding and kinetic plots. We will see its use in Chapter 30.13 when we model protein kinases that regulate the cell cycle!
The interactive graph below allows you to use the sliders to explore the effect of changes in KD and n on fractional saturation.
The Hill equation for hemoglobin gives a sigmoidal dioxygen binding curve that fits the actual binding data.
MWC Symmetry Model
In the MWC (Monod, Wyman, and Changeux) model, in the absence of ligand (oxygen), hemoglobin is assumed to exist in two distinct conformations, the T state (equivalent to the crystal structure of deoxyHb) and the R state (equivalent to the crystal structure of oxyHb without the oxygen). Without dioxygen, the T state (T0) is greatly favored over the unliganded R state (R0) at equilibrium. In the presence of increasing oxygen, the R state is favored.
A constant (somewhat equivalent to a dissociation constant) can be defined.
\begin{equation}
\mathrm{L}=\mathrm{T}_{0} / \mathrm{R}_{0}
\end{equation}
(Note: L is not the ligand concentration, so don't get confused.) In addition, let us assume that hemoglobin can not exist with some of the monomers in the tetramer in the T state while others in the same tetramer are in the R state. Hence, this model is often called the symmetry model. Finally, let's assume that each oxygen can bind to the T or R state with the dissociation constants KT and KR, respectively. These constants do not depend on the number of dioxygens bound to the tetramer. Hence
\begin{equation}
\mathrm{K}_{\mathrm{R}}=\frac{\left[\mathrm{R}_{0}\right][\mathrm{S}]}{\left[\mathrm{R}_{1}\right]}=\frac{\left[\mathrm{R}_{1}\right][\mathrm{S}]}{\left[\mathrm{R}_{2}\right]}=\ldots \frac{\left[\mathrm{R}_{\mathrm{n}}\right][\mathrm{S}]}{\left[\mathrm{R}_{\mathrm{n}+1}\right]}
\end{equation}
and
\begin{equation}
\mathrm{K}_{\mathrm{T}}=\frac{\left[\mathrm{T}_{0}\right][\mathrm{S}]}{\left[\mathrm{T}_{1}\right]}=\frac{\left[\mathrm{T}_{1}\right][\mathrm{S}]}{\left[\mathrm{T}_{2}\right]}=\ldots \frac{\left[\mathrm{T}_{\mathrm{n}}\right][\mathrm{S}]}{\left[\mathrm{T}_{\mathrm{n}+1}\right]}
\end{equation}
The subscript on R and T refers to the number of dioxygens bound to that form of R or T. A cartoon representation of the T and R forms and accompanying dioxygen binding is shown in Figure \(\PageIndex{15}\).
Now define two new parameters:
\begin{equation}
\alpha=\frac{\mathrm{pO}_{2}}{\mathrm{~K}_{\mathrm{R}}}=\frac{[\mathrm{S}]}{\mathrm{K}_{\mathrm{R}}}
\end{equation}
where α is a normalized ligand concentration describing how many times the KR the ligand concentration is, and
\begin{equation}
\mathrm{c}=\frac{\mathrm{K}_{\mathrm{R}}}{\mathrm{K}_{\mathrm{T}}}
\end{equation}
is the ratio of the dissociation constants for the R and T forms.
If oxygen binds preferentially to the R form of hemoglobin, c would be a small fractional number. In the limiting case, when oxygen didn't bind to the T form, KT would be infinite, and c = 0.
Using these definitions and equations, the following equation for Y, fractional saturation, vs α can be derived, with n, the number of binding sites per molecule, = 4 for Hb.
\begin{equation}
\mathrm{Y}=\frac{\alpha(1-\alpha)^{\mathrm{n}-1}+\operatorname{Lc} \alpha(1+\mathrm{c} \alpha)^{\mathrm{n}-1}}{(1+\alpha)^{\mathrm{n}}+\mathrm{L}(1+\mathrm{c} \alpha)^{\mathrm{n}}}
\end{equation}
Figure \(\PageIndex{16}\) shows how fractional saturation (Y) vs alpha varies with L and c for the MWC model.
When L is set at 9000, and c = 0.014, the Y vs. α curve fits the experimental oxygen binding data well. Figure \(\PageIndex{17}\) shows the best experimental dioxygen binding data that we could find (obtained from a graph, not from a table), the best fit of the Y vs L data using a Hill coefficient of n=2.8 (fitting equation 3 above), and the best fit of Y vs L using the MWC model, with L=9000, c=0.014, and Kr = 2.8 torr.
Use the sliders in the interactive graph below to explore the effect of changes in L and c on fractional saturation.
Hence, the MWC equation gives sigmoidal dioxygen binding curves like the Hill equation. It does not require an empirical Hill-like coefficient, which has no clear physical meaning!
Another way to think about the MWC Model
The MWC model assumes that oxygen binds to either the T or R form of Hb in a non-cooperative fashion. Hence, KT and KR are constant, independent of the number of dioxygens bound to that form. If so, what is the basis of the cooperative oxygen binding curves? The answer can be seen below. The cyan curve might reflect the binding of a ligand to the T form of a macromolecule, with KD = 100 uM (low affinity), for example. The binding curve looks linear, but it is just the initial part of a hyperbolic binding curve. Likewise, the magenta curve reflects the binding of a ligand to the R form of the macromolecule with KD = 10 uM. The T and R forms are linked through the T↔ R equilibrium. That equilibrium will be shifted to the tighter binding (lower KD) R form with increasing ligand concentration, assuming the ligand binds preferentially to the R form. This shifts the actual binding curve from that resembling the T form at low ligand (cyan) to one resembling the R form (magenta) as the ligand increases, imparting sigmoidal characteristics to the "observed" binding curve (gray). Figure \(\PageIndex{18}\) shows how a sigmoidal binding curve could arise from a switch from a low-affinity to a high-affinity form.
KNF Sequential Model
The KNF (Koshland, Nemethy, and Filmer) Sequential model was developed to address concerns with the concerted model. One of the major problems with the concerted model is that it seems unrealistic to expect all the subunits to change conformation together. Why shouldn't there be some differences in subunit conformation? The KNF model also fits the experimental data well. Figure \(\PageIndex{19}\) shows the linked equilibria in the KNF model. Data suggest that the MWC model better explains the transition in proteins on ligand binding and that there is an all‑or‑none interconversion between the two states.
Hemoglobin variants and disease
Many diseases have been associated with alterations in the amino acid sequence of hemoglobin. Around 300,000 babies are born each year with a genetic disorder causing an aberrant hemoglobin structure. Over 80% of these are in low/middle-income countries. The worst is sickle cell anemia, followed by β-thalassaemias and the less serious α-thalassaemias. We will only focus on sickle cell disease. In this disease, red blood cells become distorted in shape from a normal, smooth discoid shape to a crescent-like shape under low oxygen concentrations found in capillaries. These impede blood flow, as shown in Figure \(\PageIndex{25}\), and lead to symptoms ranging from anemia, episodic pain, swelling of hands and feet, and vision problems. Complications often lead to premature death.

Linus Paul and colleagues showed that the isoelectric point (pI) of oxy-Hb and deoxy-Hb in normal blood was 6.87 and 6.68, respectively, but were higher (7.09 and 6.91, respectively) in sickle cell disease. This was the first evidence that a molecular alteration of a protein caused a disease. Eventually, we learned that a single negatively charged amino acid, Glu 6, on the β-chain of hemoglobin, was mutated into a nonpolar one, valine. This causes hemoglobin at a concentration of 150 g/L in blood to self-aggregate into a long polymer. This distorts the cell into a sickle shape. Humans have two genes for the beta chain of hemoglobin, one from the egg donor and the other from the sperm donor. If only one allele is mutated, the disease is called sickle cell trait. If both are mutated, sickle cell disease is observed.
The hydrophobic Val 6 is on the surface of both beta chains in sickle cell disease. It can bind to a hydrophobic patch of Ala 70, Phe 85, and Leu 88 on another β-chain on another hemoglobin tetramer. Hence, hemoglobin has two Val 6s and two hydrophobic patches, allowing the formation of a sickle cell hemoglobin "dimer" of tetramers first, followed by the elongation of the growing fibril. This is a disease of aberrant induced dipole-induced dipole interactions and the "hydrophobic effect."
Figure \(\PageIndex{27}\) shows a "dimer" aggregate of two hemoglobin S tetramers, in which each β-chain has the D6V mutation. α chains are shown in grey, β chains in cyan, Val 6 in red spacefill, and the surface hydrophobic patch of A70, F85, and L88 in orange spacefill (1hbs). Note that the binding of the two tetramers is mediated by the interaction of the red Val 6 on the right tetramer with the orange surface hydrophobic patch on the left tetramer. The "dimer" aggregate has three more exposed Val 6 (red sphere) and three more hydrophobic patches. This would allow the extension of the dimer and the formation of long fibril-like polymers, with binding mediated by noncovalent interactions.
Figure \(\PageIndex{28}\) shows an interactive iCn3D model of one tetramer of hemoglobin S. As in Figure \(\PageIndex{21}\), the α chains are shown in grey, the β chains are shown in cyan, and Val 6s are shown in red spacefill. The surfaces of the hydrophobic pockets where the Val 6 on other HbS tetramer binds, comprised of amino acids A70, F85, and L88, are shown in orange.

Sickle cell disease and trait are endemic in Sub-Saharan Africa, where over 4 million have the disease, and over 40 million have the trait. Figure \(\PageIndex{29}\) shows its geographic distribution along with malaria.
The Plasmodium parasite reproduces in red blood cells. Their ability to reproduce is compromised as red blood cells with sickle cell hemoglobin rupture more frequently. Also, the parasite uses hemoglobin as a source of amino acids. They endocytose it and hydrolyze it to amino acids in digestive organelles. Sickle cell hemoglobin is more resistant to this process. Hence, evolution appears to have maintained the sickle cell gene in these areas to protect against malaria.
Sickle cell disease is a systemic problem, so treating many symptoms is important. Figure \(\PageIndex{30}\) describes some of these treatments.
Of course the cure would be to use DNA editing to change the base pair for the mutated Val 6 gene back to the wild-type Glu 6 as shown in Figure \(\PageIndex{31}\). The amino acid sequence encoded by the DNA and RNA shown below is Pro-Glu-Glu for the normal and Pro-Val-Glu for the sickle cell chain.

Crisper gene editing trials are now underway to attempt a cure for this dreadful disease.
Summary
This chapter explores the structural and functional paradigms of oxygen-binding proteins, using myoglobin and hemoglobin as central examples to illustrate fundamental biochemical concepts. Although myoglobin and hemoglobin are often used to introduce protein-ligand interactions, their binding mechanisms and regulatory features also provide a basis for understanding more complex systems.
Structural Features and Ligand Binding
Myoglobin is a compact, monomeric protein composed of eight α‑helices, designed primarily for oxygen storage in muscle. In contrast, hemoglobin is a heterotetramer made up of two α and two β subunits, each containing a heme group. The heme prosthetic group—consisting of protoporphyrin IX and a centrally coordinated Fe²⁺ ion—is essential for binding dioxygen (O₂) through a coordinate covalent bond. This chapter emphasizes how these protein architectures not only facilitate oxygen transport but also serve as models for studying ligand interactions that range from small gaseous molecules to large biomolecules.
Electronic Structure and Binding Interactions
A detailed discussion of the electronic configuration of Fe²⁺ (a 3d⁶ transition metal ion) provides insight into the binding mechanisms at the molecular level. By applying crystal field and ligand field theories, the chapter explains how the d orbitals of iron interact with the ligands, leading to differences in spin states (high-spin versus low-spin) and affecting the color and functionality of the heme complex. The unique properties of coordinate covalent bonding—where the ligand supplies both electrons—contrast with the noncovalent interactions observed in many other protein-ligand systems.
Cooperativity and Allosteric Regulation
Hemoglobin’s ability to bind oxygen cooperatively is a hallmark of its function. Unlike the hyperbolic binding curve of myoglobin, hemoglobin displays a sigmoidal oxygen binding curve. This behavior is explained through models such as the Hill equation, the Monod-Wyman-Changeux (MWC) model, and the Koshland-Nemethy-Filmer (KNF) model. These models illustrate how the binding of one oxygen molecule increases the affinity of the remaining sites (homotropic allosterism) and how allosteric effectors—like H⁺, CO₂, and bisphosphoglycerate (BPG)—modulate oxygen affinity (heterotropic allosterism) via structural transitions between the T (tense) and R (relaxed) states. The Bohr effect, where a decrease in pH shifts the binding curve to favor oxygen release, is highlighted as a critical regulatory mechanism in oxygen delivery to tissues.
Conformational Changes and Disease Implications
The transition from the deoxy (T) to the oxy (R) conformation in hemoglobin is central to its function. Small shifts in the heme iron position trigger larger conformational changes at the subunit interfaces, altering salt bridges and hydrogen bonds that stabilize the protein’s quaternary structure. This finely tuned regulation of oxygen binding is not only a triumph of molecular design but also a vulnerability. The chapter concludes with a discussion of hemoglobin variants, particularly sickle cell disease, where a single amino acid substitution (Glu → Val) in the β-chain leads to aberrant polymerization, distorted red blood cells, and significant clinical consequences. The evolutionary trade-offs, such as the partial protection against malaria conferred by the sickle cell trait, further underscore the interplay between molecular structure and biological function.
Overall, the chapter integrates structural biology, bioinorganic chemistry, and mathematical modeling to provide a comprehensive understanding of how myoglobin and hemoglobin function as paradigms for protein-ligand interactions, highlighting their central roles in physiology and disease.



.png?revision=1&size=bestfit&width=237&height=206)
.png?revision=1&size=bestfit&width=258&height=213)
.png?revision=1&size=bestfit&width=192&height=189)
.png?revision=1&size=bestfit&width=183&height=198)