7.2: Polysaccharides
- Last updated
- Save as PDF
- Page ID
- 102272
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Search Fundamentals of Biochemistry
Learning Goals
(Learning goals written by Claude, Sonnet 4.6, Anthropic)
α-Linked Homopolysaccharides: Structure and Function
- Compare the structures and biological roles of starch (amylose: unbranched α1→4 glucose polymer; amylopectin: α1→4 main chain with α1→6 branches every 24–30 residues) and glycogen (α1→4 main chain with far more frequent α1→6 branches), explaining how the helical conformation of α1→4-linked chains (analogous to the protein α-helix in having non-extended φ/ψ angles) creates a hollow central channel that accommodates iodine (I₃⁻) to produce the characteristic blue-purple color used in starch indicators, and why the high branching density of glycogen (with glucose released simultaneously from all non-reducing branch ends) allows rapid glucose mobilization during metabolic demand.
- Explain why storing glucose as glycogen or starch rather than as free monosaccharides is chemically advantageous in terms of colligative properties — noting that a single glycogen particle (anchored to the core protein glycogenin via Tyr 195) exerts negligible osmotic pressure compared to the millions of free glucose molecules it contains — and describe how the V-amylose structure (a left-handed single helix of 6 residues per turn, with an ~8 Å pitch and a hollow interior) can self-associate into a 4-helix bundle as a tertiary structural unit, analogous to the 4-helix bundle seen in some proteins.
β-Linked Homopolysaccharides: Structural Roles
- Explain why the β1→4 glycosidic linkage in cellulose and chitin produces extended, linear chains (with φ/ψ angles analogous to the extended protein β-strand) rather than helices — describing how every other glucose (in cellulose) or GlcNAc (in chitin) monomer is flipped 180° relative to its neighbor, how multiple cellulose chains align in parallel and are stabilized by extensive intrachain and interchain hydrogen bonds, and how the axial C–H bonds on the top and bottom faces of cellulose strands create flat hydrophobic surfaces that drive additional hydrophobic stacking interactions between adjacent layers — explaining why this combination of hydrogen bonding and hydrophobic interactions makes cellulose the most abundant biological molecule on Earth and an exceptionally strong structural polymer.
- Connect the chemical difference between cellulose (β1→4-linked glucose polymer) and chitin (β1→4-linked N-acetylglucosamine polymer) — noting that a single N-acetyl substituent at C2 distinguishes the two — to their distinct biological roles (cellulose as the primary plant cell wall structural polymer; chitin as the major component of arthropod exoskeletons and fungal cell walls) and to the reason humans can digest starch but not cellulose (lack of a β-glucosidase, while α-amylase cleaves α1→4 links).
Glycosaminoglycans: Heteropolysaccharides with Disaccharide Repeats
- Describe the structural features shared by all glycosaminoglycans (GAGs) — repeating disaccharide units containing one amino sugar (GlcNAc, GalNAc) and one uronic acid (glucuronate, iduronate) or galactose, with variable sulfation at multiple positions creating a highly negatively charged polyanion — and compare the disaccharide repeat units, sulfation patterns, and biological locations of hyaluronic acid (glucuronate-β1,3-GlcNAc repeats, no sulfate, synovial fluid and extracellular matrix), keratan sulfate (Gal-β1,4-GlcNAc-6-sulfate repeats, cornea), chondroitin sulfate (glucuronate-β1,4-GalNAc-4- or 6-sulfate repeats, cartilage), dermatan sulfate (iduronate-α1,3-GalNAc-4-sulfate, derived by epimerization of glucuronate), and heparin (highly trisulfated glucuronate/iduronate-GlcNSO₃ repeats, anticoagulant).
- Explain the mechanism of heparin's anticoagulant activity — describing how heparin simultaneously binds antithrombin III (inducing a conformational change that makes antithrombin III a more potent thrombin inhibitor) and transiently binds thrombin (a positively charged serine protease) electrostatically, converting the search for the inhibitor from a three-dimensional diffusional process to a one-dimensional sliding search along the polyanion — and connect this to the broader principle that highly sulfated GAGs like heparin create diverse, sequence-variable binding surfaces for positively charged proteins despite the absence of a genetic template specifying GAG sequence.
Polysaccharides contain many monosaccharides in glycosidic links and may have many branches. They serve as either structural components or energy storage molecules. Polysaccharides consisting of single monosaccharides are homopolymers. Starch, glycogen, dextran, cellulose, and chitin are the most common. We'll discuss based on whether the acetal link is alpha or beta.
α 1,4 main chain links
Starch and Glycogen: These polysaccharides are polymers of glucose linked in α 1,4 links with α 1,6 branches. Starch, found in plants, is subdivided into amylose, which has no branches, and amylopectin, which does. Starch granules consist of about 20% amylose and 80% amylopectin. Glycogen, the main CHO storage in animals, is found in muscle and liver and consists of glucose residues in α 1,4 links with lots of α 1,6 branches (many more branches than in starch).
Here are various ways to render in 2D the chemical structure of a branched glycogen and starch fragment, as shown in Figure \(\PageIndex{1}\).
The top part of the figure shows the Haworth structure. The bottom part shows two glucose units in red and blue in the more structurally clear chair and wedge/dash representations.
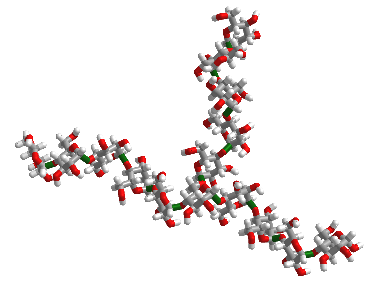
Figure \(\PageIndex{2}\) shows an interactive iCn3D model of 10 glucose monosaccharides in an α-(1,4) linkage with five glucose units with α-(1,4) linkages attached to the main chain through an α-(1,6) branch at glucose 6 of the main chain. The type of substructure would be found in starch (amylopectin) and glycogen.
_linked_glucose_with_an_%25CE%25B1-(1%252C6)_branch.png?revision=1&size=bestfit&width=336&height=259) I
I
Figure \(\PageIndex{3}\) shows the structure in the iCn3D model in a diagrammatic fashion in which glucose is represented as a blue circle with the acetal/glycosidic/glucosidic linkages between the monosaccharides written between the circles. The 14A label shows that the acetal linkage is an α-(1,4) link with a single α-(1,6) branch.

The linkages are written according to various conventions. These include 14A, 14α, 4A and 4α. The link between many sugars is often a 1,x link where x is 2,3, 4, 5, or 6. In those cases, the number 1 can be omitted. The program-generated images in this text use numbers and A or B.
What makes carbohydrates so complex is their 3D structures. Like proteins and nucleic acids, they can adopt a myriad of conformations. Because monomeric units are so homogeneous, especially in homopolymers, obtaining crystal structures can be difficult; computer models are often used instead.
Studies have shown that the simple starch fraction amylose, an α 1,4 polymer of glucose, often envisioned as a straight chain, can adopt three main conformations. They are double-helical A- (found chiefly in cereals), double helical B-(found primarily on tubers) amyloses, and single-helical V-amylose (or simply A, B, and V structures). The A and B do NOT represent alpha or beta in this classification system. The A and B forms consist of double helices aligned in parallel, with about 6 glucose residues per turn. The helices appear to be left or right-handed, and this ambiguity might arise from a lack of crystal structures.
In contrast, a well-defined structure of the V helix is known. It folds into a left-handed helix with six glucose residues per turn and a pitch of about 8Å. Unlike alpha helices of proteins and the double-stranded helix of DNA, the center of the helix is NOT packed tightly and can accommodate small molecules. One is iodine (triiodide, I3-), which exhibits a dark blue color when bound in amylose in starch. This is the basis for starch indicators that you may have used in titration reactions in biology and chemistry courses.
Alpha helices might self-associate during folding in proteins to form a 4-helix bundle. Likewise, the helices in V-amylose can associate into bundles. Figure \(\PageIndex{4}\) shows an interactive iCn3D model of the actual structure of a V-amylose, cycloamylose 26 (1C58). It consists of a linear cycloamylose strand of 26 glucose monomers, which has collapsed to form secondary structure with 6-residue helices packed together into a tertiary structure of 4-helix bundle. The blue-sphere "cartoon" color coding of each glucose residue corresponds to the blue circles in the diagram above.
.png?revision=1&size=bestfit&width=372&height=291)
Rotate the model to explore it. Trace the chains by following the blue sphere symbolic representation of glucose as you trace the main chain. Rotate it to view down the helix axes to see the four holes that can each accommodate an I3-. In the menu button (=), choose Style, Chemicals, Sphere to see a spacefilling model that shows the holes within each helix. Remember, NO unoccupied holes exist within either a protein alpha helix or a double-stranded DNA molecule.
The well-known macrocyclic compound cyclodextrins (for example, α-cyclodextrin) are structures equivalent to one turn of V-amylose. The V-amylose helix is partially stabilized by hydrogen bonds from donors and acceptors within the helix from the OH3 on the ith glucose and the OH2 on the i+1th glucose, as well as from the OH6 on the ith glucose and the OH2 on the i+6th glucose.
In vivo, glycogen is synthesized by attaching glucose monomers to a core protein called glycogenin. Figure \(\PageIndex{5}\) shows a model of a glycogen particle with glycogenin at its core.
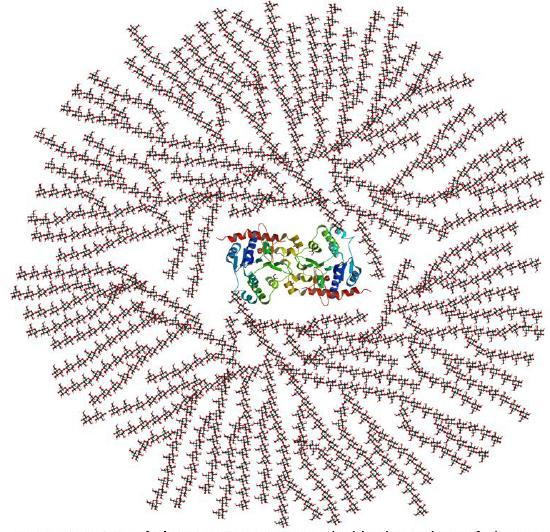
The dimeric protein glycogenin is an enzyme that autoglucosylates itself in a stepwise manner. The first glucose is added at Tyr 195. At some point, the active site must get buried, and the protein can no longer add more monomers.
Storing glucose residues as either glycogen or starch, one large molecule, makes chemical sense. A review of colligative properties would inform you that if glucose were stored as a monosaccharide, a great osmotic pressure difference would be found between the outside and inside of the cell. Glycogen, with its many branches, is a single molecule. When glucose is needed, it is cleaved one residue at a time from all the branches (at the nonreducing ends) of glycogen, producing a large amount of free glucose quickly.
Phi/Psi angles can also be used in Ramachandran plots to show the conformations around the acetal link of the starch/glycogen main chain, in a manner comparable to that for proteins (around the alpha carbon). The phi torsion angle describes rotation around the C1-O bond of the acetal link, and the psi angle describes rotation around the O-C4 bond of the same acetal link, with the glucopyranose ring considered as a rigid rotator (just as the six atoms in the planar peptide bond unit). The most extended form of a glucose polymer occurs when the glycosidic link is β-1,4 (as in cellulose), forming linear chains. This would be analogous to the more extended parallel beta strand (phi/psi angles of -1190, -1130) and antiparallel beta strands (phi/psi angles of -1390, +1350) of proteins. The α 1,4-linked main chain of glycogen and starch causes the chain to turn and form a large helix. Iodine (or I3-) can fit into the helix, which turns a solution/suspension of starch blue, which turns starch purple. The less extended structure is analogous to the less extended protein alpha helix, which has phi/psi angles of -570,-470.
Figure \(\PageIndex{6}\) shows phi/psi angles for acetal/glycosidic linkage in maltose, a disaccharide of glucose, as shown below.
α 1,6 main chain links
Dextran is a branched polymer of glucose in α 1,6 links with α 1,2, α 1,3, or α 1,4 linked side chains. This polymer is used in some chromatography resins. Figure \(\PageIndex{7}\) shows chair structures (A) and wedge/dash structures (B) for dextran, showing the main chain α 1,6 link with one α 1,3 branch.
Depending on its molecular weight, it is soluble in water (forming viscous solutions) and organic solvents. It is also used as a food thickener and stabilizer. It is synthesized by lactic acid-forming bacteria using sucrose as an energy source. Most uses are commercial.
β 1,4 links
Cellulose, a structural homopolymer of glucose in plants, has β 1,4 main chain links without branching. Multiple chains are held together by intra- and inter-chain H-bonds. It is the most abundant biological molecule in nature. Various renderings of 4 glucose residues in cellulose are shown in Figure \(\PageIndex{8}\). Haworth structures are not shown. Instead, more chemically informative chairs and wedge/dash structures are used. It's important to see the structures in various forms, since carbohydrate structures are represented differently across sources.
In A, the most common chair representation, the 2nd and 4th residues from the right-hand end are flipped versions of residues 1 and 3. Residues 1 and 2 are colored red and blue for clarity. This unit is repeated to generate the full chain. The top part of A shows a simplified version of the flip of the red ring to produce the blue ring to help you see that they are indeed identical structures.
The same structure as in A is shown in the left part of B in wedge/dash from (looking down on the ring). The right-hand side of B shows a variant of B's left-hand side generated by a simple 1800 rotation around the bond indicated on the left of B.
The simple repeat is shown in C without the chain flips in A and B. The acetal/glycosidic/glucosidic bond appears as a straight line in the chair structures (a bit confusing and structurally deceptive), but is shown more clearly in the adjacent wedge/dash structure.
All structures are correct, but the one shown in A is most often used.
A single long chain can interact with other chains, forming a structure stabilized by intrachain and interchain hydrogen bonds. Different sources display different hydrogen bonds. Some common ones are shown below. These chains align in parallel and twist to form larger cellulose fibers. Figure \(\PageIndex{9}\) shows an interactive iCn3D model of cellulose chains.

In addition, hydrophobic interactions occur between adjacent planes of cellulose strands. How can that be, considering the strongly polar nature of glucose and a single cellulose strand? Figure \(\PageIndex{9B}\): below, with two representations of cellulose, shows how axial hydrogens project vertically from the plane of the glucose monomer rings to form a weak but biologically significant hydrophobic surface.
Figure \(\PageIndex{9}\): Two representations showing the nonpolar axial surface of cellulose strands that contribute to hydrophobic interactions stabilizing the cellulose assemblies. The top panel shows how cellulose could interact with a nonpolar planar substance, such as a layer of graphite. Axial H atoms are shown in green/cyan in the bottom panel.
Top Panel: Yang G, Luo X and Shuai L (2021) Bioinspired Cellulase-Mimetic Solid Acid Catalysts for Cellulose Hydrolysis. Front. Bioeng. Biotechnol. 9:770027. doi: 10.3389/fbioe.2021.770027. Creative Commons Attribution License (CC BY).
Bottom Panel: Uusi-Tarkka, E.-K.; Skrifvars, M.; Haapala, A. Fabricating Sustainable All-Cellulose Composites. Appl. Sci. 2021, 11, 10069. https://doi.org/10.3390/app112110069. Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/
4.0/).
The glycan is the major component in the exoskeletons of anthropoids and mollusks. It is a β 1,4-linked polymer of N-acetylglucosamine (GlcNAc). Compare this with cellulose, a β-1,4-linked polymer of glucose. What a difference an N-acetyl substituent makes!
The basic chemical structure of chitin is shown in chair form in Figure \(\PageIndex{10}\) along with the symbolic nomenclature for glycans (SNFG).
Symbolic nomenclature for glycans (SNFG) -
Before we go further into the complexities of glycan structure, let's explore the symbolic nomenclature for glycan structures. The Consortium for Functional Glycomics (2005) proposed a scheme based on specific colored geometric shapes for each, as shown for the example glycan shown in Figure \(\PageIndex{11}\) for a complex glycan.
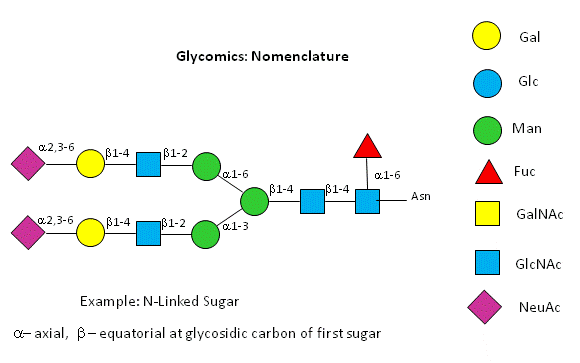
This nomenclature has recently been updated in Appendix 1B of Essentials of Glycobiology, 3rd Edition (Glycobiology 25(12): 1323-1324, 2015. doi: 10.1093/glycob/cwv091 (PMID 26543186) and is summarized in the Figure \(\PageIndex{12}\).
Glycosaminoglycans - Heteropolysaccharides with Disaccharide Repeating Units
Many polysaccharides consist of repeating disaccharide units. A major class of polysaccharides with disaccharide repeats includes the glycosaminoglycans (GAGs), all of which contain one amino sugar in the repeat, and in which one or both of the sugars contain negatively charged sulfate and/or carboxyl groups. The extent and position of sulfation vary widely between and within GAGs. GAGs are found in the vitreous humor of the eye and synovial fluid of joints, as well as in connective tissue like tendons, cartilage, etc., and skin. They are found in the extracellular matrix and are often covalently attached to proteins to form proteoglycans. From a bird's eye view, they are all elongated polyanions.
They and their structures are very complicated and exceedingly diverse. This makes it difficult for those who want unambiguous structures. From a biological perspective, they present an incredibly diverse array of potential ligand-binding sites in their local environment (both small and large). Because of these, they also play a role in cell signaling. In addition, some GAGs are free-standing, while others are covalently attached to proteins (like glycogen is attached to glycogenin). These large molecules are called proteoglycans. We will discuss this later in Chapter 7.4 when we discuss the "carbohydrate code."
Here are the ring structures and descriptions of important GAGs. The common disaccharide repeat unit is shown twice for each structure, with the knowledge that sulfation patterns may differ for the disaccharide repeats in the actual chains. Note also that the first member of each disaccharide repeat shows the ring flipped vertically (top to bottom) as was shown in the structures for other beta-linked glycans (cellulose, chitin) above.
In a long chain, selecting which is the repeating disaccharide unit is a bit relative, as shown in Figure \(\PageIndex{13}\) for the repeating disaccharide sequence of N-acetylglucosamine (blue square) and N-acetylgalactosamine (yellow square).

At the top, the repeating units (blue-yellow) are connected through beta 1,4 links, while at the bottom, the connection of the repeating unit (yellow-blue) is beta 1,3. The best way to annotate the repeating unit without knowing the full chain is elusive. What's most important, however, is to note the alternating acetal/glycosidic links throughout the whole sequence. In the figures below, different disaccharide repeats are highlighted.
Hyaluronic acid
This is a polymer of glucuronate (β1,3) linked to GlcNAc. It offers a backbone for the attachment of proteins and other GAGs. It's the only GAG without sulfate. Figure \(\PageIndex{14}\) shows a tetrasaccharide fragment with two disaccharide repeats. The illustrated disaccharide repeat's internal acetal/glycosidic link is β 1,3, while the disaccharide's connection is β 1,4. For one last time, the vertical flip of the glucuronic acid is shown to allow a better understanding of its flipped presentation in the actual GAG.
Hyaluronic acids are found in a variety of locations, including synovial fluid, the extracellular matrix, and skin, where they help control skin moisture. It is water-soluble and displays twin antiparallel left-oriented helices. Covalent conjugates of the chemotherapeutic drugs doxorubicin and camptothecin linked to hyaluronic acid, whose overall structure is similar to "worm-like micelles", have been used successfully to treat skin cancers.
Figure \(\PageIndex{15}\) shows an interactive iCn3D model of hyaluronic acid (4HYA). The dotted lines represent hydrogen bonds.
Three calcium ions are shown as well. The SNFG cartoon is also illustrated.
Keratan sulfate
This GAG contains repeats of N-acetyl-D-glucosamine-6-phosphate in β 1,3 link to D-galactose or D-galactose-6-sulfate. The link between Gal and the modified glucosamine is β 1,4. Keratin sulfate is most abundant in the cornea of the eye as well as in other connective tissues such as bone, cartilage, and tendon, as well as in the central and peripheral nervous system.
Figure \(\PageIndex{16}\) shows a tetrasaccharide containing two repeating disaccharides.
Chondroitin sulfate
The repeat disaccharide unit is D-glucuronate β(1,4) GalNAc-4 or 6-sulfate. It's found in the connective tissue matrix, at the cell surface (as proteoglycans), in basement membranes, and in intracellular granules. Figure \(\PageIndex{17}\) shows a tetrasaccharide showing two disaccharide repeats.
Figure \(\PageIndex{18}\) shows an interactive iCn3D model of chondroitin-4-sulfate (1C4S). The dotted lines represent hydrogen bonds.
Dermatan sulfate
This glycosaminoglycan is similar to chondroitin sulfate. It is first formed as a polymer of the disaccharide units D-gluconic and N-acetyl-D-galactosamine. Gluconic acid is epimerized to L-iduronic acid, which is then sulfated. Its structure is shown in Figure \(\PageIndex{19}\).
Heparin
This GAG contains a highly trisulfated disaccharide repeat, as shown in Figure \(\PageIndex{20}\). Note that the molecule can contain glucuronate or iduronate, and the degree of sulfation of the chains varies. Remember, no genetic code specifies the sequence or sulfation pattern of these polymers.
Figure \(\PageIndex{21}\) shows an interactive iCn3D model of the solution structure of an 18-mer of heparin (3IRI). The dotted lines represent hydrogen bonds.
Most people are familiar with the anti-clotting properties of heparin administered as a drug. Heparin acts as a "catalyst" to accelerate the inhibition of the enzyme thrombin, which cleaves fibrinogen and activates platelets to form clots by the blood protein antithrombin III. Heparin works in two ways to facilitate thrombin inactivation. It has a specific binding site for antithrombin III, which causes a conformational change in the protein, making it a more effective inhibitor. Thrombin, a positively-charged serine protease, can nonspecifically bind heparin, a polyanion. When it does, it diffuses along the heparin chain, where it can bind to bound antithrombin III much more quickly than if the inhibitor were free in the blood. Heparin effectively redirects thrombin's search from 3D to 1D.
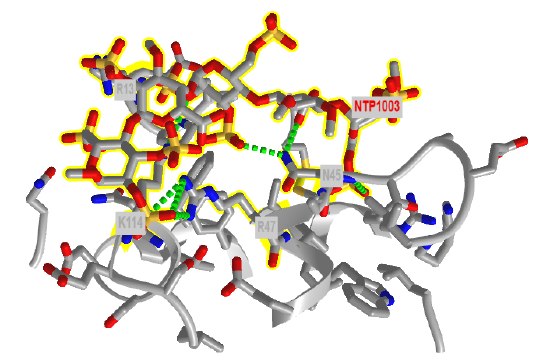
Figure \(\PageIndex{22}\) shows an interactive iCn3D model of the amino acids in antithrombin III within seven angstroms of a bound heparin 5mer (1NQ9). Dotted lines represent hydrogen bonds and salt bridges between the two. Heparin is highlighted in yellow
Agarose
Agarose is the main polysaccharide component derived from red algae. Agarose is a polymer of a disaccharide repeat of (1,3)-β-D-galactopyranose-(1,4)-3,6-anhydro-α-L-galactopyranose, is often used for a gelable solid phase for electrophoresis of nucleic acid and as a component of chromatography beads. As with starch, a mixture of amylose and amylopectin, agarose is often found with agaropectin, a sulfated galactan. A tetrasaccharide fragment with two disaccharide repeats is shown in Figure \(\PageIndex{23}\).
Figure \(\PageIndex{24}\) shows an interactive iCn3D model of the agarose double helix (1AGA). The dotted lines represent hydrogen bonds within the polymer.
Figure \(\PageIndex{24}\): Agarose double helix (1AGA). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...5nnzm9XcPogMz7
In the iCn3D model, choose Style, Glycan, Show Cartoon to see the yellow sphere for galactose.
Summary
(Summary written by Claude, Sonnet 4.6, Anthropic)
This chapter surveys the major classes of polysaccharides — homopolymers of glucose or glucose derivatives linked through α or β glycosidic bonds, and heteropolymers with disaccharide repeat units — illustrating how the type of glycosidic linkage, the degree and pattern of branching, and chemical modifications of the monosaccharide building blocks determine the three-dimensional structure and biological function of these carbohydrates.
α-Linked homopolysaccharides — starch and glycogen — serve as the primary carbohydrate energy stores in plants and animals, respectively. Both are polymers of glucose joined by α1→4 main-chain glycosidic bonds with α1→6-linked branches at intervals. Starch exists as a mixture of amylose (unbranched, ~20% of starch) and amylopectin (branched at every 24–30 residues, ~80%). Glycogen (the animal equivalent) is much more highly branched than amylopectin, enabling the simultaneous release of glucose from all non-reducing ends when glycogen phosphorylase cleaves the chain — a critical feature for rapid glucose mobilization during exercise or fasting. The α1→4 linkage imparts a helical conformation because its φ/ψ angles (analogous to protein α-helix φ/ψ values) correspond to a non-extended, turning structure. Amylose adopts three main conformations: double-helical A (cereals) and B (tubers) forms and the single-helical V form. The V-amylose helix has six glucose residues per turn with a pitch of ~8 Å and, unlike the protein α-helix or DNA double helix, contains a hollow core capable of accommodating small molecules. Triiodide (I₃⁻) binding within this channel produces the characteristic dark blue color used in starch indicator tests. V-amylose helices can further self-associate into 4-helix bundle quaternary structures analogous to those in proteins. The biological logic of glucose storage as polymers rather than free monosaccharides is rooted in colligative chemistry: a single glycogen particle (containing up to 55,000 glucose residues anchored through the core protein glycogenin) exerts essentially no osmotic pressure, whereas free glucose at equivalent concentration would create a dangerous osmotic imbalance. Glycogenin itself is an autoglucosylating enzyme that initiates glycogen synthesis by attaching the first glucose to its own Tyr 195, continuing until the active site is buried.
β-Linked homopolysaccharides — cellulose and chitin — serve structural rather than energy storage roles. Cellulose (β1→4-linked glucose) is the most abundant biological molecule in nature and the primary structural polymer of plant cell walls. The β1→4 linkage forces each glucose ring to be rotated 180° relative to its neighbors, producing a fully extended chain with φ/ψ angles analogous to the protein β-strand. Multiple parallel cellulose chains form extensive intrachain and interchain hydrogen bond networks. Crucially, the equatorial C–H bonds on cellulose project perpendicular to the plane of the ring on top and bottom faces, creating flat hydrophobic surfaces that drive additional hydrophobic stacking interactions between adjacent cellulose layers — an unexpected source of stability in an otherwise very polar molecule. The combination of dense hydrogen bonding and hydrophobic stacking gives cellulose exceptional tensile strength. Chitin (β1→4-linked N-acetylglucosamine) is structurally analogous to cellulose, differing only in the N-acetyl substituent at C2 that replaces the C2 hydroxyl of glucose; this single modification dramatically alters the mechanical and biochemical properties of the polymer, making it the dominant structural material in arthropod exoskeletons and fungal cell walls. The contrast between cellulose/chitin (β-linked, structural, indigestible by humans) and starch/glycogen (α-linked, energy storage, digestible by α-amylase) illustrates the profound functional consequences of anomeric configuration. Dextran (α1→6 main chain with various branch linkages) is a bacterial polymer of commercial interest as a chromatography matrix, food thickener, and experimental tool.
Glycosaminoglycans (GAGs) are a structurally and functionally diverse class of heteropolysaccharides built from repeating disaccharide units, each containing one amino sugar (GlcNAc or GalNAc) and one uronic acid (glucuronate or iduronate) or galactose. Variable sulfation at multiple positions of both sugar residues — patterns that are not specified by a genetic template but are instead controlled by the spatial and temporal expression of sulfotransferase and epimerization enzymes — creates an extraordinarily diverse array of polyanions. Hyaluronic acid (glucuronate-β1,3-GlcNAc repeats, unsulfated) forms twin antiparallel left-handed helices and is found in synovial fluid, the vitreous humor, the extracellular matrix, and skin; it can be conjugated to chemotherapeutic drugs for targeted cancer therapy. Keratan sulfate (Gal-β1,4-GlcNAc-6-sulfate) predominates in the cornea and nervous tissue. Chondroitin sulfate (glucuronate-β1,4-GalNAc-4- or 6-sulfate) and dermatan sulfate (its 5-epimer, containing iduronate derived by post-synthetic epimerization of glucuronate) are major components of connective tissue, cartilage, and basement membranes. Heparin contains the most highly sulfated disaccharide repeating unit in biology (trisulfated, with variable glucuronate/iduronate content), and its potent anticoagulant activity illustrates a remarkable mechanism: heparin simultaneously binds antithrombin III (inducing a conformational change that dramatically accelerates thrombin inhibition) and transiently binds thrombin electrostatically, reducing the dimensionality of the thrombin-antithrombin encounter from a 3D diffusion problem to a 1D sliding search along the polyanion — effectively acting as a catalytic scaffold rather than a stoichiometric inhibitor. Agarose (a disaccharide polymer of β-D-galactose and 3,6-anhydro-α-L-galactose from red algae) forms double helices and gels, making it indispensable for nucleic acid electrophoresis and affinity chromatography matrices. The SNFG (Symbolic Nomenclature for Glycans) system uses colored geometric shapes to represent monosaccharides, providing a standardized visual language for the complex, branched structures of glycans that would be impractical to draw in full chemical detail.




.png?revision=1&size=bestfit&width=526&height=202)



.png?revision=1&size=bestfit&width=202&height=333)