19.2C: Antibiotics
- Page ID
- 5972
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Humans, and our domestic animals, can serve as hosts to a wide variety of disease-causing organisms (pathogens) including bacteria, viruses, fungi, protozoans, helminths (worms). This page will examine only those chemical agents that are used to combat bacterial pathogens.
The Problem
There are many chemicals that are lethal to bacteria - cyanide does a good job — but they cannot be used to cure infections because they are lethal to the host as well. The problem, then, is to find substances that attack a metabolic pathway found in the bacterium but not in the host. This is not an insurmountable problem for bacterial pathogens because they differ in many respects from eukaryotes.
The Solution
Natural products. A number of natural products, penicillin for example, have been discovered that are antibiotics suitable for therapy. They were originally discovered as secretions of fungi or soil bacteria. Soils are complex ecosystems, and it is not surprising that its inhabitants have evolved chemical defenses against each other.
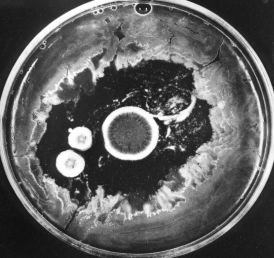
The photo (courtesy of Merck & Co., Inc.) shows how the growth of bacteria on the agar in a culture dish has been inhibited by the three circular colonies of the fungus Penicillium notatum. The antibiotic penicillin, diffusing outward from the colonies, is responsible for this effect. Today, penicillin is made from cultures of Penicillium chrysogenum that has been specially adapted for high yields.
- Semi-synthetic products. These are natural products that have been chemically modified in the laboratory (and pharmaceutical facility) to
- improve the efficacy of the natural product
- reduce its side effects
- circumvent developing resistance by the targeted bacteria
- expand the range of bacteria that can be treated with it
- Completely synthetic products. The sulfa drugs are examples.
Sulfa Drugs and Folic Acid Analogs
Sulfa Drugs
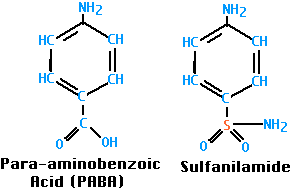
Sulfanilamide was the first antibacterial agent. Many other sulfa drugs (such as sulfamethoxazole) have since come into use.
Both bacteria and their human hosts require folic acid for nucleic acid synthesis (it is converted into purines and thymidine) as well as protein synthesis (precursor of the amino acids methionine and glycine). However, bacteria synthesize their folic acid starting with para-aminobenzoic acid (PABA), while we must ingest our folic acid already formed; that is, for us it is a vitamin.
Sulfanilamide, and the other sulfa drugs, are analogs of PABA; they compete with PABA and, when chosen, block the synthesis of folic acid. Mammals ignore PABA and its analogs and thus can tolerate sulfa drugs.
Folic Acid Analogs
These synthetic molecules block the final step in the conversion of PABA to folic acid so they, too, block nucleotide and protein synthesis in bacteria but not in mammals. Trimethoprim is one of several in current use. These folic acid analogs are often used in combination with a sulfa drug.
The Beta-Lactams
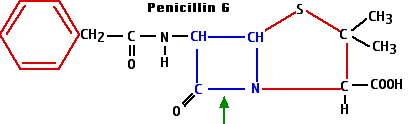
The beta-lactams include the
- penicillins such as
- penicillin G (a natural product) produced by the fungus Penicillium chrysogenum
- ampicillin (a semi-synthetic)
- amoxicillin (semi-synthetic)
- cephalosporins There are over two dozen of them in current use. Most are semi-synthetics derived from the secretion of the mold Cephalosporium. Some examples:
- cephalexin (e.g., Keflex®)
- cefaclor (e.g., Ceclor®)
- cefixime (e.g., Suprax®)
- carbapenems such as
- meropenem (Merrem®)
- ertapenem (Invanz®)
The beta-lactams all work by interfering with the synthesis of the bacterial cell wall — a structure that is not found in eukaryotes. The walls of bacteria are made of a complex polymeric material called peptidoglycan.
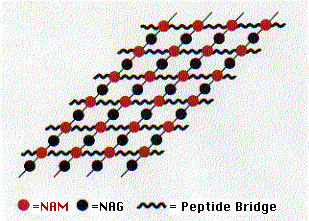
Peptidoglycan contains both amino acids and amino sugars. The amino sugars are of two kinds
- N-acetylglucosamine (NAG) and its close relative
- N-acetylmuramic acid (NAM).
These two form a linear polymer of NAG alternating with NAM. They are linked by a glycosidic bond between the #1 and #4 carbons (this is the linkage attacked by lysozyme) and are oriented in the same way they are in cellulose. Side chains containing 4 or 5 amino acids are attached to each NAM. These form covalent bonds with amino acids in adjacent chains. The bonds may be direct to the next chain or include additional peptide cross bridges (e.g., 5 glycine residues) which extend to chains in the same plane (shown here) as well as to chains above and below.
This elaborate, covalently cross-linked structure provides the great strength of the cell wall. It also leads to the remarkable conclusion that the bacterial cell wall meets the definition of a single molecule!
The beta-lactam antibiotics bind to and inhibit enzymes needed for the synthesis of the peptidoglycan wall. While they have little effect on resting bacteria, they are lethal to dividing bacteria as defective walls cannot protect the organism form bursting in hypotonic surroundings.
Aminoglycosides
These are products of actinomycetes (soil bacteria) or semi-synthetic derivatives of the natural products.
Examples are:
- streptomycin
- kanamycin
- neomycin
- gentamycin
The 70S bacterial ribosome differs in several ways from the 80S eukaryotic ribosome. The aminoglycosides bind to the 30S subunit of the bacterial ribosome and interfere with the formation of the initiation complex. They also cause misreading of the mRNA. Although the eukaryotic ribosome in the cytosol is relatively unaffected by these drugs, ribosomes in the mitochondria are 70S and sensitive to their effects.
Tetracyclines
These are natural products derived from soil actinomycetes or their semi-synthetic derivatives. Examples:
- chlortetracycline (aureomycin®)
- oxytetracycline (terramycin®)
- doxycycline
- tigecycline (Tygacil®)
Tetracyclines bind to the 30S subunit of the bacterial ribosome. They prevent the transfer of activated amino acids to the ribosome so protein synthesis is halted.
Macrolides, Lincosamides, Streptogramins, and Ketolides
The Chink in the Armor = the bacterial ribosome
These antibiotics bind to the large (50S) subunit of the bacterial ribosome where they block the growing peptide chain from exiting the ribosome thus severely hindering protein synthesis. Because of their similar action, the development of antibiotic resistance to one usually extends to all the others.
Macrolides
Macrolides are also products of actinomycetes (soil bacteria) or semi-synthetic derivatives of them. Erythromycin, azithromycin (Zithromax®), and clarithromycin (Biaxin®) are a commonly-prescribed macrolides.
Lincosamides
The first member of this group was also isolated from a soil actinomycete (found near Lincoln, Nebraska). A semi-synthetic derivative, called clindamycin (Cleocin®), is now widely used against Gram-positive bacteria.
Streptogramins
Quinupristin and dalfopristin are examples. As of 1 October 1999, they will be sold as a mixture under the trade name Synercid. Combined, they show great promise in treating certain infections resistant to vancomycin- currently the antibiotic of last resort for some hospital-acquired infections.
Ketolides
Ketolides are derivatives of macrolides. Telithromycin (Ketek®) is an example. Ketolides also bind to the 50S subunit of the bacterial ribosome. In so doing they cause it to induce frameshifts during translation. Bacteria that have become resistant to macrolides, lincosamides, and streptogramins are still susceptible to ketolides.
Fluoroquinolones
Ciprofloxacin (Cipro®), levofloxacin and norfloxacin are examples. Cipro is the preferred antibiotic for people who have been intentionally exposed to anthrax, although some other antibiotics appear to be equally effective.
The Chink in the Armor = DNA topoisomerases
The fluoroquinolones block the action of two bacterial topoisomerases - enzymes that relieve the coils that form in DNA when the helix is being opened in preparation for replication or transcription or repair. The topoisomerases in eukaryotes are not affected.
Polypeptides
The most common of these is polymixin E (also known as colistin). It behaves as a detergent, increasing the permeability of the membranes that encase bacteria and causing the contents of the bacterial cell to leak out.
Rifampin
This semi-synthetic antibiotic binds to the bacterial RNA polymerase and prevents it from carrying out its role in transcription. Its affinity for the equivalent eukaryotic enzyme is much lower. Rifampin is also known as rifampicin.
Mupirocin
This antibiotic blocks the action of the bacterial isoleucine tRNA synthetase, the enzyme responsible for attaching the amino acid isoleucine (Ile) to its tRNA in preparation for protein synthesis, so protein synthesis is inhibited. It spares the equivalent eukaryotic enzyme.
Cycloserine
Cycloserine inhibits synthesis of the bacterial cell wall but by a different mechanism than the beta-lactam antibiotics discussed above. Cycloserine is an analog of D-alanine and blocks the incorporation of D-alanine into the peptide bridges in the bacterial cell wall. It is derived from an actinomycete.
Aminocyclitols
These products of another actinomycete achieve their effect by interfering with the 30S subunit of the bacterial ribosome. Spectinomycin (trade name = Trobicin®) is an example. It is particularly effective against the gonococcus, the bacterium that causes the sexually-transmitted disease (STD) gonorrhea.
Glycopeptides
Glycopeptides also interfere with the synthesis of the bacterial cell wall but by a different mechanism than the beta-lactams.
Vancomycin is a widely-used glycopeptide in the U.S. It binds to the D-alanines on the precursors of the peptidoglycan cross bridges preventing their cross-linking. It has become the antibiotic of last resort as resistance to the other antibiotics has become more and more common.
Oxazolidinones
The first of these new antibiotics, linezolid (Zyvox®), was approved by the U.S. Food and Drug Administration on 19 April 2000. It is effective against many Gram-positive bacteria that have developed resistance to the older antibiotics.
Linezolid attacks a previously-unexploited chink in the bacterium's armor: the proper assembly of the two ribosomal subunits (30S and 50S). It does not affect eukaryotic ribosomes — and thus translation of mRNAs in the cytosol. However, it does affect the bacterial-like mitochondrial ribosomes and can interfere with the synthesis of those mitochondrial proteins synthesized by them.
Lipopeptides
These are natural compounds derived from a species of Streptomyces. The one now in clinical use is daptomycin (Cubicin®). It is effective against Gram-positive bacteria. It attacks another previously-unexploited chink in the bacterial armor — the integrity of its cell membranes.
So far there is no evidence of bacteria developing resistance against it.
Resistance to Antibiotics
None of the antibiotics discussed above is effective against all bacterial pathogens.
Intrinsic resistance
Some bacteria are intrinsically resistant to certain of the antibiotics. Example: Gram-positive bacteria are much less susceptible to polymixins than Gram-negative bacteria. [The "Gram" designations refer to the behavior of the bacteria when stained with the Gram stain; this behavior is a reflection of the very different organization of their cell walls.]
Acquired resistance
Many bacteria acquire resistance to one or more of the antibiotics to which they were formerly susceptible.
Example: In the U.S. in the decade from 1985–1995, resistance of Shigella (which causes gastrointestinal illness) to ampicillin grew from 32% to 67%. And, while only 7% of these isolates were resistant to the combination of sulfamethoxazole and trimethoprim at the start of the decade, that figure had grown to 35% by the end of the decade.
Bacteria develop resistance by acquiring genes encoding proteins that protect them from the effects of the antibiotic. In some cases the genes arise by mutation; in others, they are acquired from other bacteria that are already resistant to the antibiotic. The genes are often found on plasmids which spread easily from one bacterium to another - even from one species of bacterium to another.
Examples:
- Synthesis of the enzyme penicillinase - or other beta-lactamases - provides protection from the beta-lactam antibiotics. These enzymes break the beta-lactam ring at the position shown with the green arrow in the diagram of penicillin G.
- Likewise synthesis of cephalosporinases defeats the cephalosporins.
- Defeating quinolones:
- Some bacteria do this by modifying their DNA gyrase.
- Others, e.g., Mycobacterium tuberculosis, develop quinolone resistance by synthesizing a protein that resembles a short length of DNA. This protein binds the gyrase so it cannot form the DNA/gyrase complex that is the target of quinolone action.
- Some bacteria synthesize "pumps" in their plasma membrane through which they remove antibiotics like tetracyclines from the interior of the cell.
- Bacteria may methylate their ribosomes obscuring the target of antibiotics (e.g., erythromycin) that ordinarily bind to and inactivate the ribosome — or conversely
- they may enzymatically modify the antibiotic (e.g., kanamycin) so it can no longer "see" its ribosomal target.
- Bacteria may modify the structure of their peptidoglycan wall and thus avoid the inhibitory effects of antibiotics like cycloserine.
An alarming number of human pathogens have acquired genes to combat all the presently-used antibiotics except vancomycin and recently vancomycin-resistant bacteria have appeared. These multidrug-resistant strains are particularly common in hospitals where antibiotic use is heavy, and the patients often have weakened immune systems.
Measuring Antibiotic Resistance
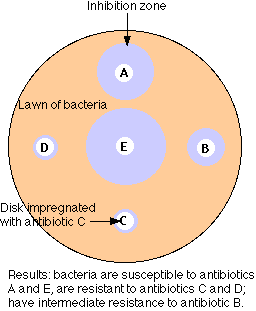
The figure illustrates the simplest method of the several available for measuring antibiotic resistance.
- A suspension of the bacteria to be tested (e.g. cultured from the infected patient) is spread over the surface of a petri dish containing a solid culture medium.
- Disks of several different antibiotics are pressed on the surface of the agar. The concentration of antibiotic in each type of disk is standardized.
- Incubate overnight.
- The bacteria will grown into a "lawn" except where an antibiotic to which they are sensitive has diffused out from its disk.
- Measure the diameter of any zones of inhibition that are formed.
What can you do to delay the spread of antibiotic resistance?
- Don't ask your doctor for an antibiotic to treat a viral disease (e.g., a cold) for which antibiotics are useless. (However, your doctor may prescribe an antibiotic if you are infected by an influenza virus - not to fight the virus but to protect you against a secondary bacterial infection of your damaged lungs.)
- Stay the course. Use all doses prescribed even though you are feeling better. This will minimize the opportunity to select for resistance among the bacteria that remain late in the infection.
- Don't save unused antibiotics for later self-medication.
Farmers can help as well by avoiding the use of antibiotics in their livestock that are similar to those used in humans. Antibiotics are widely used in healthy livestock to improve their growth rate (by an unknown mechanism).
An article in the 20 May 1999 issue of The New England Journal of Medicine documents the recent development of quinolone resistance in Campylobacter jejuni, the most frequent bacterial cause of gastroenteritis in humans. The rise coincides with the approval in 1995 of the use of quinolones by U. S. poultry farmers (chickens also become infected by C. jejuni). Similar recent increases in fluoroquinolone-resistant C. jejuni have been reported in the Netherlands and also in Spain (where as many as 50% of human infections are now caused by bacteria resistant to the antibiotic). In each country, the appearance of resistant strains followed the widespread introduction of quinolone treatment for animals.
Future Prospects
Drug companies - after many years of complacency - are now responding to the threat of antibiotic-resistant bacteria. Over a dozen new antibiotics are being developed and some have already reached clinical trials. Many of these are semi-synthetic modifications of already-existing antibiotics, including new
- beta-lactams
- macrolides
- glycopeptides
- quinolones
- modifications of vancomycin
Others are entirely new, attacking previously-unexploited chinks in the bacterial armor.
- Urea hydroxamates, that block the enzyme (peptide deformylase) that removes fMet from the finished protein so it can begin its work. (Eukaryotes do not begin translation with fMET.)
- Heteroaromatic polycycles (HARP) that bind to bacterial promoters preventing gene transcription.


