Meiosis
- Page ID
- 23891
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)1. Description of Meiosis
Meiosis is the type of cell division used to produce gametes (sperm and eggs). Meiosis assures that genetic diversity is achieved during sexual reproduction. Meiosis consists of 2 cell divisions: Meiosis I and Meiosis II. Meiosis starts with a diploid (2n) parent cell that divides to make 4 haploid (n) cells. In sexual reproduction, haploid gametes from two different individuals combine to produce a diploid zygote. The resulting offspring is genetically different from both parents.
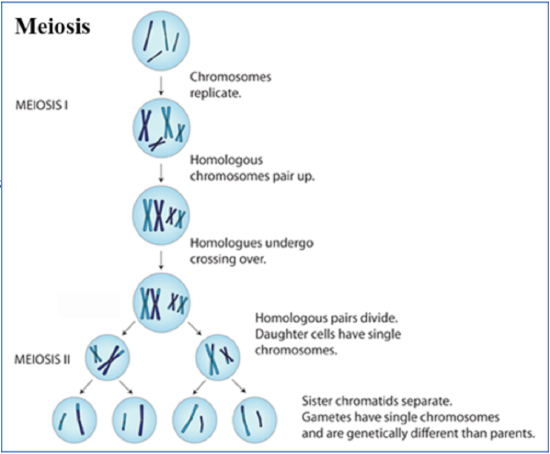
Figure \(\PageIndex{1}\). (CC BY-NC-SA)
Chromosome characteristics:
Haploid (n) = one set of chromosomes
Diploid (2n) = two sets of chromosomes
Eggs and sperm (gametes) are haploid
Diploid set for humans: 2n = 46
Interphase before Meiosis: During the interphase preceding meiosis, DNA replication takes place.
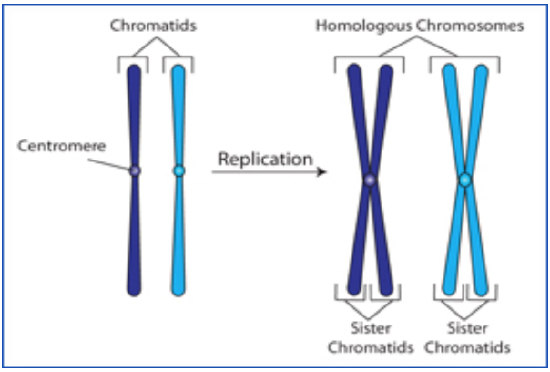
Figure \(\PageIndex{2}\). (CC BY-NC-SA)
Meiosis I
Prophase I: Homologous chromosomes pair up and form tetrads. This pairing is known as synapsis. While paired, the homologous chromosomes exchange genetic material in a process called crossing over. Crossing over contributes to the genetic variation of sexual reproduction. While all this is occurring, the nuclear envelope and nucleoli begin to disappear. Spindle fibers attach to the chromosomes and begin moving them to the equatorial plate.
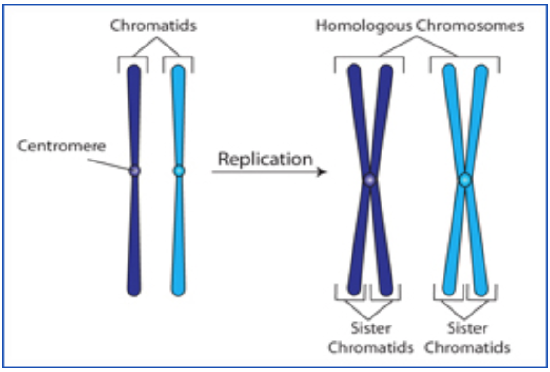
Figure \(\PageIndex{3}\). (CC BY-NC-SA)
Metaphase I: Homologous chromosomes, in a pair-wise fashion, have lined up on the equatorial plate. One homologue is positioned on each side of the equatorial plate. The orientation is random, which means that there is a 50-50 chance for the daughter cells to get either the maternal or paternal homologue for each chromosome. This is known as independent assortment.
Anaphase I: Chromosomes from each pair move to opposite poles of the cell. Each chromosome still consists of two sister chromatids.
Telophase I: Nuclear envelopes may reform, or the cell may immediately start meiosis II. DNA replication does NOT take place. There are now only a haploid number of chromosomes in each cell.
Summary of Meiosis I: Crossing over occurs between homologous chromosomes. Homologous chromosomes separate from each other and 2 haploid cells are formed.
Meiosis II
Prophase I: Chromatin once again condenses into discrete chromosomes. The spindle apparatus forms.
Metaphase II: Chromosomes are lined up along the equatorial plate, similar to metaphase in mitosis. Due to crossing over in meiosis I, the two sister chromatids of each chromosome are no longer genetically identical. Microtubules from opposite poles attach to each sister chromatid of a chromosome.
Anaphase II: Sister chromatids separate and move toward opposite poles as individual chromosomes.
Telophase II: Chromosomes decondense and nuclear envelopes reform. Meiotic division has produced 4 daughter cells, each with a haploid set of chromosomes and each chromosome has only one chromatid. Each of the 4 daughter cells is genetically distinct from each other and the parent cell.
Summary of Meiosis II: Sister chromatids separate from each other (similar to mitosis) and 4 haploid gamete cells are formed.
Mitosis vs. Meiosis
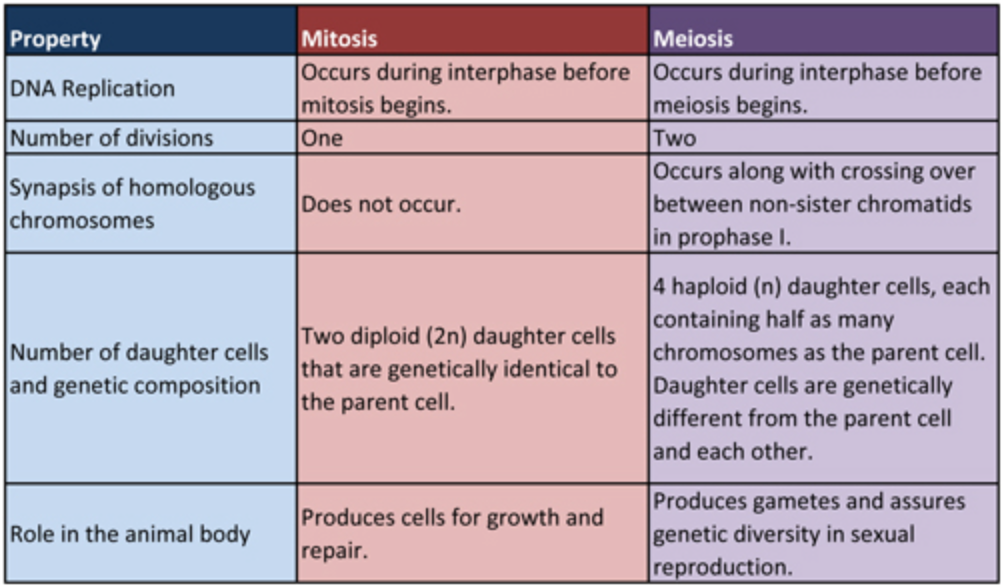

Figure \(\PageIndex{4}\). (CC BY-NC-SA)
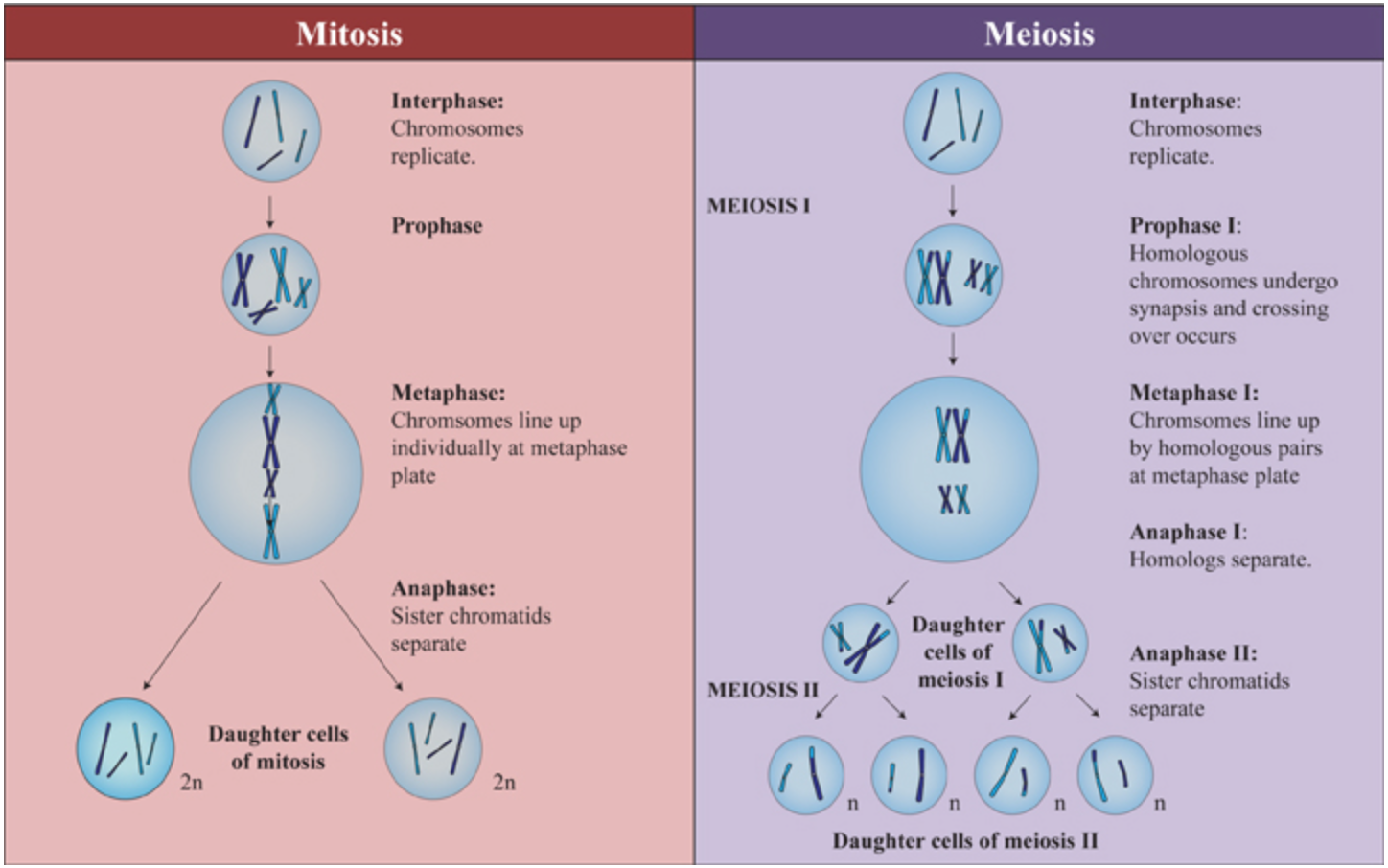
Figure \(\PageIndex{5}\). (CC BY-NC-SA)

Meiosis Tutorial by Dr. Katherine Harris is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License.
Funded by the U.S. Department of Education, College Cost Reduction and Access (CCRAA) grant award # P031C080096.


