DNA Profiling
- Page ID
- 2861
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)DNA fingerprinting is a technique that is used to identify patterns that occur in DNA. No two organisms have identical DNA so this procedure can be used to identify if a sample of DNA came from a particular individual. We will use this procedure in lab to identify whether a sample of DNA found at a crime scene belongs to one of three suspects. The technique has a variety of other uses, e.g., it can be used to identify whether individuals carry genes for certain genetic diseases.
Restriction Enzymes
The technique of DNA fingerprinting requires that the DNA be cut up into small fragments. Restriction enzymes are used to perform this digestion. Restriction enzymes were discovered in bacteria, which use them as a defense mechanism to cut up the DNA of viruses or other bacteria. Hundreds of different restriction enzymes have been isolated. Each one cuts DNA at a specific base sequence. For example, EcoRI always cuts DNA at GAATTC as indicated below.
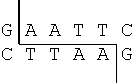
The sequence GAATTC appears three times in the DNA strand below. As a result, the strand is cut into four pieces.
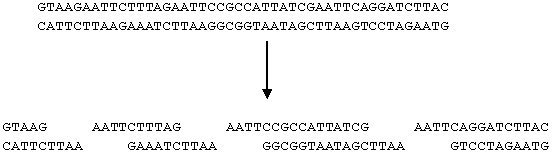
Other restriction enzymes cut at different sites, some examples are listed below.
| Enzyme | Cutting Site |
| Bam HI | GGATCC |
| Hae III | GGCC |
| Pst I | CTGCAG |
| Hinf I | GANTC |
In RFLP analysis, the DNA of an organism is cut up into fragments using restriction enzymes. A large number of short fragments of DNA will be produced.
Restriction enzymes always cut at the same base sequence. Because no two individuals have identical DNA, no two individuals will have the same length fragments. For example, the enzyme EcoRI always cuts DNA at the sequence GAATTC. Different people are going to have different numbers of this particular sequence and will therefore have different fragment lengths. In addition, some of them will be at different locations on the chromosome.
Gel Electrophoresis
Electrophoresis is a technique used to separate the DNA fragments according to their size. They are placed on a sheet of gelatin and an electric current is applied to the sheet. DNA is charged and will move in an electric field toward the positive pole. In the diagram below, holes (wells) in the gelatin can be seen. DNA samples placed in these wells will migrate through the gelatin toward the + side after an electric current is applied.
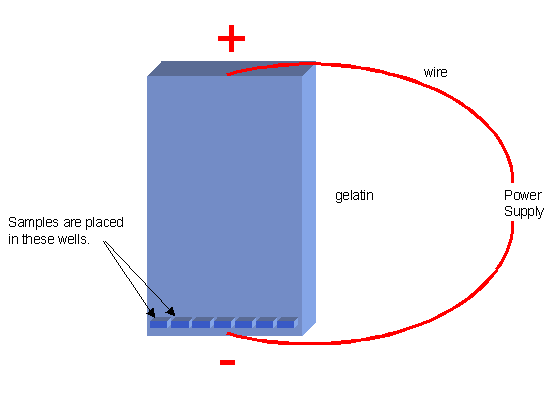
The smallest fragments will move the fastest because they are able to move through the pores in the gelatin faster. Bands will be produced on the gelatin where the fragments accumulate. The shortest fragments will accumulate near one end of the gelatin and the longer, slower-moving ones will remain near the other end.
In the diagram below, four samples of DNA were placed on the gelatin. After an electric current was applied for a period of time, the fragments separated. Notice that sample D on the right does not match the other three samples.
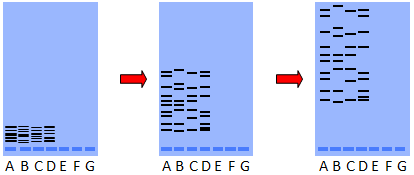
The DNA bands must be stained to make them visible. Ethidium bromide-stained DNA will fluoresce when illuminated with UV light. PCR techniques are used to produce sufficient quantities of DNA for this technique.
Procedure - Day 1
Setting Up the Apparatus
For this exercise, you will be assigned to investigate either a cancer gene (see Table A below) or DNA collected at a crime scene (Table B). In either case, the DNA has been cut using restriction enzymes.
Table A. The table below will be used for investigating a gene for cancer that occurs in families. All of the DNA was cut with the same restriction enzyme.
| Component (tube) | Source of DNA |
| A | Standard DNA fragments |
| B | Control DNA |
| C | Patient blood DNA |
| D | Patient tumor DNA |
| E | Patient breast normal DNA |
Table B. The table below will be used for investigating DNA found at a crime scene and DNA from two suspects.
| Component (tube) | Source of DNA | Enzyme used |
| A | Crime scene | Enzyme 1 |
| B | Crime scene | Enzyme 2 |
| C | Suspect 1 | Enzyme 1 |
| D | Suspect 1 | Enzyme 2 |
| E | Suspect 2 | Enzyme 1 |
| F | Suspect 2 | Enzyme 2 |
- Remove the tape from the end of the gel trays if it has not been done already.
- A plastic comb was used to create the wells for the samples. Carefully remove this comb.
- Place the gel tray in the electrophoresis apparatus. The wells should be placed nearest the negative (black) electrode.
- Add enough buffer solution so that the gel is completely submerged.
- The gel has eight lanes but you will need five or six for your samples. The two outside lanes can be used to practice loading samples in the wells. Use the practice loading solution for this purpose.
- Use a micropipette to load 25 ul of sample A (component A) into the second well from the left. Repeat this procedure, placing each of the remaining samples in a different well. Record what samples that you placed in each of the wells. This can be done by writing the sample letter and the well number for that sample.
Connecting the Power Supply
7. Place the lid on the apparatus. The red and black electrodes on the base should match the electrode connections on the lid.
8. Connect the apparatus to the transformer. This transformer can be used to run two different gels.
9. Switch on the transformer using the switch on the right side near the back.
10. Use the button to the right of the LED display on the transformer to select "V."
11. Use the up and down arrows to the left of the LED display to adjust the voltage to 125.
12. To start power flow to the gel, press the button on the right side of the front panel. This button shows a drawing of a person running. The green light next to this button indicates that the power is on. Check for the production of bubbles in the electrophoresis apparatus to confirm that the power is turned on.
Running the Gels
13. The gels should run for about 45 minutes. Each sample contains a marker dye that runs just ahead of the smallest DNA fragments. The electricity should be switched off when this dye approaches the end of the gel. Do not let the dye run off of the gel.
14. When the gels have finished running, switch off the power, disconnect the apparatus, and remove the lid.
Staining the Gels
15. Place a blue staining sheet on a plastic staining tray with the stain (blue) side facing upward.
16. Remove the plastic gel holder from the electrophoresis apparatus and and slide the gel onto a plastic staining tray. The gel should be directly on top of the staining paper. Ideally, the staining paper should remain on the gel at room temperature for at least one day.
17. If you plan to view the DNA in another lab period, wrap the tray containing the gel and staining paper with plastic wrap.
Day 2 - Results and Conclusions
After the gel has been allowed to stain for at least one day, remove the staining paper. The gel should be ready to view by placing the tray on a light source.
Draw your results or paste a photograph in your notebook. One of the tables below should also be included in your notebook.
Cancer Gene
For each lane on the gel, explain what is demonstrated and why it produced the results that it did.
| Why did this lane produce the number and size fragments that it did? | |
Lane 1- Standard DNA fragments |
|
Lane 2- Control DNA |
|
Lane 3- Blood DNA |
|
Lane 4- Tumor DNA |
|
Lane 5- Normal Breast Tissue DNA |
Crime Scene
In your lab notebook, explain which suspect's DNA was at the crime scene. Explain how you know. It is not enough to say that DNA from a suspect matches the crime scene. You must be more specific, for example, "DNA from the crime scene cut with enzyme 1 matches (or does not match) DNA from suspect 1 cut with enzyme 1" etc.
| Conclusion - Which crime scene DNA sample does each of the following match or not match? | ||
Suspect 1 |
Enzyme 1 |
|
Suspect 1 |
Enzyme 2 |
|
Suspect 2 |
Enzyme 1 |
|
Suspect 2 |
Enzyme 2 |

