14.3: Inhibitors of Cell Wall Biosynthesis
- Page ID
- 144195
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the mechanisms of action associated with drugs that inhibit cell wall biosynthesis.
- Explain the mode of action of β-lactams.
- Describe why β-lactamase (penicillinase) offers drug resistance for bacteria.
- Explain the mode of action of non-β-lactam cell wall inhibitors.
An important quality for an antimicrobial drug is selective toxicity, meaning that it selectively kills or inhibits the growth of microbial targets while causing minimal or no harm to the host. Most antimicrobial drugs currently in clinical use are antibacterial because the prokaryotic cell provides a greater variety of unique targets for selective toxicity, in comparison to fungi, parasites, and viruses. Each class of antibacterial drugs has a unique mode of action (the way in which a drug affects microbes at the cellular level), and these are summarized in Figure \(\PageIndex{1}\) and Table \(\PageIndex{1}\).
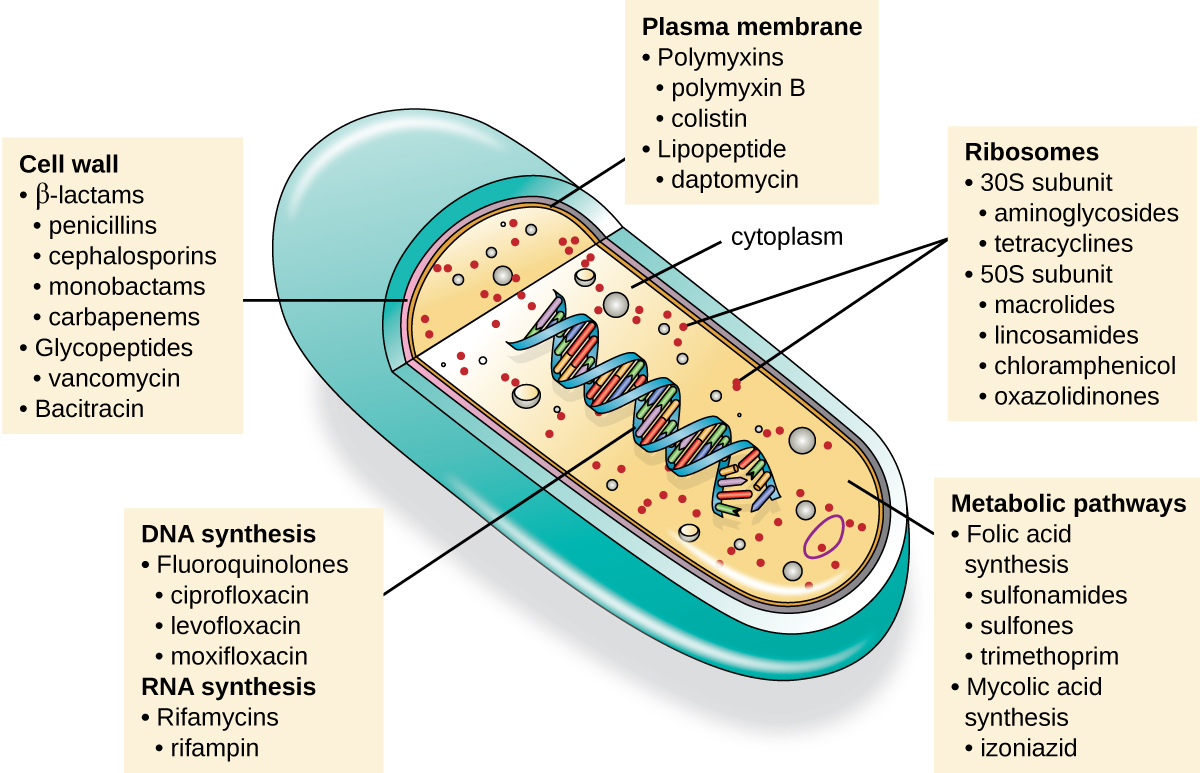
| Mode of Action | Target | Drug Class |
|---|---|---|
| Inhibit cell wall biosynthesis | Penicillin-binding proteins | β-lactams: penicillins, cephalosporins, monobactams, carbapenems |
| Peptidoglycan subunits | Glycopeptides | |
| Peptidoglycan subunit transport | Bacitracin | |
| Inhibit biosynthesis of proteins | 30S ribosomal subunit | Aminoglycosides, tetracyclines |
| 50S ribosomal subunit | Macrolides, lincosamides, chloramphenicol, oxazolidinones | |
| Disrupt membranes | Lipopolysaccharide, inner and outer membranes | Polymyxin B, colistin, daptomycin |
| Inhibit nucleic acid synthesis | RNA | Rifamycin |
| DNA | Fluoroquinolones | |
| Antimetabolites | Folic acid synthesis enzyme | Sulfonamides, trimethoprim |
| Mycolic acid synthesis enzyme | Isonicotinic acid hydrazide | |
| Mycobacterial adenosine triphosphate (ATP) synthase inhibitor | Mycobacterial ATP synthase | Diarylquinoline |
Inhibitors of Cell Wall Biosynthesis
Several different classes of antibacterials block steps in the biosynthesis of peptidoglycan, making cells more susceptible to osmotic lysis (Table \(\PageIndex{2}\)). Therefore, antibacterials that target cell wall biosynthesis are bactericidal in their action. Because human cells do not make peptidoglycan, this mode of action is an excellent example of selective toxicity.
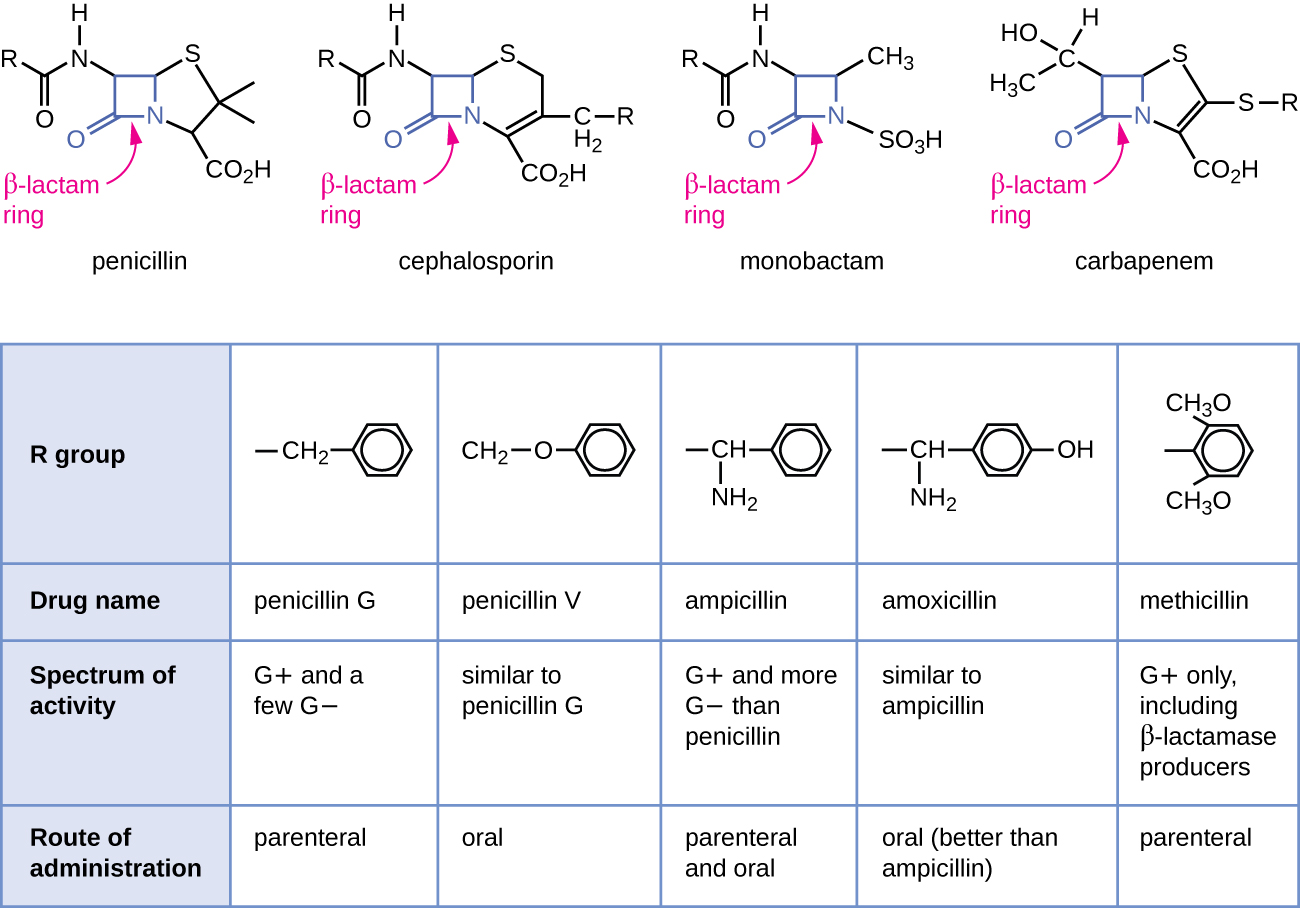
Penicillin, the first antibiotic discovered, is one of several antibacterials within a class called β-lactams. This group of compounds includes the penicillins, cephalosporins, monobactams, and carbapenems, and is characterized by the presence of a β-lactam ring found within the central structure of the drug molecule (Figure \(\PageIndex{2}\)). The β-lactam antibacterials block the crosslinking of peptide chains during the biosynthesis of new peptidoglycan in the bacterial cell wall. They are able to block this process because the β-lactam structure is similar to the structure of the peptidoglycan subunit component that is recognized by the crosslinking transpeptidase enzyme, also known as a penicillin-binding protein (PBP). Although the β-lactam ring must remain unchanged for these drugs to retain their antibacterial activity, strategic chemical changes to the R groups have allowed for development of a wide variety of semisynthetic β-lactam drugs with increased potency, expanded spectrum of activity, and longer half-lives for better dosing, among other characteristics.
Penicillin G and penicillin V are natural antibiotics from fungi and are primarily active against gram-positive bacterial pathogens, and a few gram-negative bacterial pathogens such as Pasteurella multocida. Figure \(\PageIndex{2}\) summarizes the semisynthetic development of some of the penicillins. Adding an amino group (-NH2) to penicillin G created the aminopenicillins (i.e., ampicillin and amoxicillin) that have increased spectrum of activity against more gram-negative pathogens. Furthermore, the addition of a hydroxyl group (-OH) to amoxicillin increased acid stability, which allows for improved oral absorption. Methicillin is a semisynthetic penicillin that was developed to address the spread of enzymes (penicillinases) that were inactivating the other penicillins. Changing the R group of penicillin G to the more bulky dimethoxyphenyl group provided protection of the β-lactam ring from enzymatic destruction by penicillinases, giving us the first penicillinase-resistant penicillin.
Query \(\PageIndex{1}\)
Similar to the penicillins, cephalosporins contain a β-lactam ring (Figure \(\PageIndex{2}\)) and block the transpeptidase activity of penicillin-binding proteins. However, the β-lactam ring of cephalosporins is fused to a six-member ring, rather than the five-member ring found in penicillins. This chemical difference provides cephalosporins with an increased resistance to enzymatic inactivation by β-lactamases. The drug cephalosporin C was originally isolated from the fungus Cephalosporium acremonium in the 1950s and has a similar spectrum of activity to that of penicillin against gram-positive bacteria but is active against more gram-negative bacteria than penicillin. Another important structural difference is that cephalosporin C possesses two R groups, compared with just one R group for penicillin, and this provides for greater diversity in chemical alterations and development of semisynthetic cephalosporins. The family of semisynthetic cephalosporins is much larger than the penicillins, and these drugs have been classified into generations based primarily on their spectrum of activity, increasing in spectrum from the narrow-spectrum, first-generation cephalosporins to the broad-spectrum, fourth-generation cephalosporins. A new fifth-generation cephalosporin has been developed that is active against methicillin-resistant Staphylococcus aureus (MRSA).
The carbapenems and monobactams also have a β-lactam ring as part of their core structure, and they inhibit the transpeptidase activity of penicillin-binding proteins. The only monobactam used clinically is aztreonam. It is a narrow-spectrum antibacterial with activity only against gram-negative bacteria. In contrast, the carbapenem family includes a variety of semisynthetic drugs (imipenem, meropenem, and doripenem) that provide very broad-spectrum activity against gram-positive and gram-negative bacterial pathogens.
Query \(\PageIndex{1}\)
The drug vancomycin, a member of a class of compounds called the glycopeptides, was discovered in the 1950s as a natural antibiotic from the actinomycete Amycolatopsis orientalis. Similar to the β-lactams, vancomycin inhibits cell wall biosynthesis and is bactericidal. However, in contrast to the β-lactams, the structure of vancomycin is not similar to that of cell-wall peptidoglycan subunits and does not directly inactivate penicillin-binding proteins. Rather, vancomycin is a very large, complex molecule that binds to the end of the peptide chain of cell wall precursors, creating a structural blockage that prevents the cell wall subunits from being incorporated into the growing N-acetylglucosamine and N-acetylmuramic acid (NAM-NAG) backbone of the peptidoglycan structure (transglycosylation). Vancomycin also structurally blocks transpeptidation. Vancomycin is bactericidal against gram-positive bacterial pathogens, but it is not active against gram-negative bacteria because of its inability to penetrate the protective outer membrane.
The drug bacitracin consists of a group of structurally similar peptide antibiotics originally isolated from Bacillus subtilis. Bacitracin blocks the activity of a specific cell-membrane molecule that is responsible for the movement of peptidoglycan precursors from the cytoplasm to the exterior of the cell, ultimately preventing their incorporation into the cell wall. Bacitracin is effective against a wide range of bacteria, including gram-positive organisms found on the skin, such as Staphylococcus and Streptococcus. Although it may be administered orally or intramuscularly in some circumstances, bacitracin has been shown to be nephrotoxic (damaging to the kidneys). Therefore, it is more commonly combined with neomycin and polymyxin in topical ointments such as Neosporin.
Query \(\PageIndex{1}\)
| Mechanism of Action | Drug Class | Specific Drugs | Natural or Semisynthetic | Spectrum of Activity |
|---|---|---|---|---|
| Interact directly with PBPs and inhibit transpeptidase activity | Penicillins | Penicillin G, penicillin V | Natural | Narrow-spectrum against gram-positive and a few gram-negative bacteria |
| Ampicillin, amoxicillin | Semisynthetic | Narrow-spectrum against gram-positive bacteria but with increased gram-negative spectrum | ||
| Methicillin | Semisynthetic | Narrow-spectrum against gram-positive bacteria only, including strains producing penicillinase | ||
| Cephalosporins | Cephalosporin C | Natural | Narrow-spectrum similar to penicillin but with increased gram-negative spectrum | |
| First-generation cephalosporins | Semisynthetic | Narrow-spectrum similar to cephalosporin C | ||
| Second-generation cephalosporins | Semisynthetic | Narrow-spectrum but with increased gram-negative spectrum compared with first generation | ||
| Third- and fourth-generation cephalosporins | Semisynthetic | Broad-spectrum against gram-positive and gram-negative bacteria, including some β-lactamase producers | ||
| Fifth-generation cephalosporins | Semisynthetic | Broad-spectrum against gram-positive and gram-negative bacteria, including MRSA | ||
| Monobactams | Aztreonam | Semisynthetic | Narrow-spectrum against gram-negative bacteria, including some β-lactamase producers | |
| Carbapenems | Imipenem, meropenem, doripenem | Semisynthetic | Broadest spectrum of the β-lactams against gram-positive and gram-negative bacteria, including many β-lactamase producers | |
| Large molecules that bind to the peptide chain of peptidoglycan subunits, blocking transglycosylation and transpeptidation | Glycopeptides | Vancomycin | Natural | Narrow spectrum against gram-positive bacteria only, including multidrug-resistant strains |
| Block transport of peptidoglycan subunits across cytoplasmic membrane | Bacitracin | Bacitracin | Natural | Broad-spectrum against gram-positive and gram-negative bacteria |
Key Concepts and Summary
- Antibacterial compounds exhibit selective toxicity, largely due to differences between prokaryotic and eukaryotic cell structure.
- Cell wall synthesis inhibitors, including the β-lactams, the glycopeptides, and bacitracin, interfere with peptidoglycan synthesis, making bacterial cells more prone to osmotic lysis.


