23.4: B Lymphocytes and Antibodies
- Page ID
- 146426
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the production and maturation of B cells
- Compare the structure of B-cell receptors and T-cell receptors
- Compare the maturation of B cells with the maturation of T cells
Humoral immunity refers to mechanisms of the adaptive immune defenses that are mediated by antibodies secreted by B lymphocytes, or B cells. This section will focus on B cells and discuss their production and maturation, receptors, and mechanisms of activation.
B Cell Production and Maturation
Like T cells, B cells are formed from multipotent hematopoietic stem cells (HSCs) in the bone marrow and follow a pathway through lymphoid stem cell and lymphoblast (see Figure 17.3.1). Unlike T cells, however, lymphoblasts destined to become B cells do not leave the bone marrow and travel to the thymus for maturation. Rather, eventual B cells continue to mature in the bone marrow.
The first step of B cell maturation is an assessment of the functionality of their antigen-binding receptors. This occurs through positive selection for B cells with normal functional receptors. A mechanism of negative selection is then used to eliminate self-reacting B cells and minimize the risk of autoimmunity. Negative selection of self-reacting B cells can involve elimination by apoptosis, editing or modification of the receptors so they are no longer self-reactive, or induction of anergy in the B cell. Immature B cells that pass the selection in the bone marrow then travel to the spleenfor their final stages of maturation. There they become naïve mature B cells, i.e., mature B cells that have not yet been activated.
Query \(\PageIndex{1}\)
B Cell Receptor (BCR)
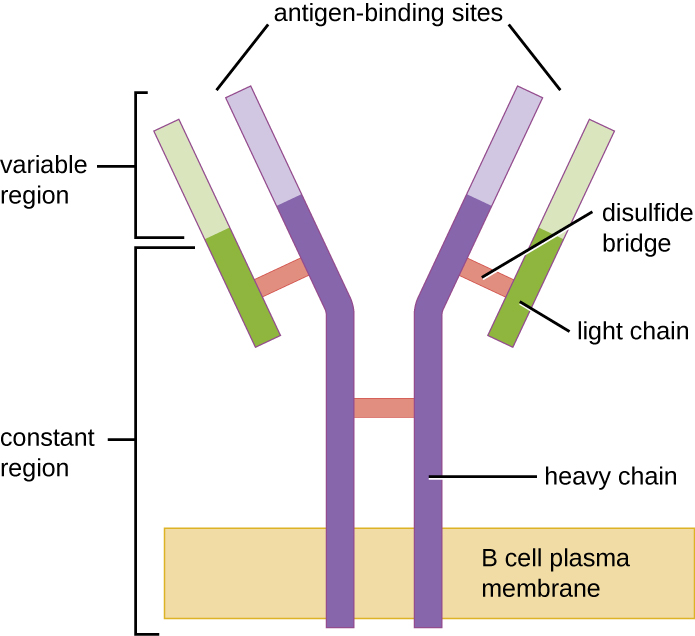
Like T cells, B cells possess antigen-specific receptors with diverse specificities. Although they rely on T cells for optimum function, B cells can be activated without help from T cells. B-cell receptors (BCRs) for naïve mature B cells are membrane-bound monomeric forms of IgD and IgM. They have two identical heavy chains and two identical light chains connected by disulfide bonds into a basic “Y” shape (Figure \(\PageIndex{1}\)). The trunk of the Y-shaped molecule, the constant region of the two heavy chains, spans the B cell membrane. The two antigen-binding sites exposed to the exterior of the B cell are involved in the binding of specific pathogen epitopes to initiate the activation process. It is estimated that each naïve mature B cell has upwards of 100,000 BCRs on its membrane, and each of these BCRs has an identical epitope-binding specificity.
In order to be prepared to react to a wide range of microbial epitopes, B cells, like T cells, use genetic rearrangementof hundreds of gene segments to provide the necessary diversity of receptor specificities. The variable region of the BCR heavy chain is made up of V, D, and J segments, similar to the β chain of the TCR. The variable region of the BCR light chain is made up of V and J segments, similar to the α chain of the TCR. Genetic rearrangement of all possible combinations of V-J-D (heavy chain) and V-J (light chain) provides for millions of unique antigen-binding sites for the BCR and for the antibodies secreted after activation.
One important difference between BCRs and TCRs is the way they can interact with antigenic epitopes. Whereas TCRs can only interact with antigenic epitopes that are presented within the antigen-binding cleft of MHC I or MHC II, BCRs do not require antigen presentation with MHC; they can interact with epitopes on free antigens or with epitopesdisplayed on the surface of intact pathogens. Another important difference is that TCRs only recognize protein epitopes, whereas BCRs can recognize epitopes associated with different molecular classes (e.g., proteins, polysaccharides, lipopolysaccharides).
Activation of B cells occurs through different mechanisms depending on the molecular class of the antigen. Activation of a B cell by a protein antigen requires the B cell to function as an APC, presenting the protein epitopes with MHC II to helper T cells. Because of their dependence on T cells for activation of B cells, protein antigens are classified as T-dependent antigens. In contrast, polysaccharides, lipopolysaccharides, and other nonprotein antigens are considered T-independent antigens because they can activate B cells without antigen processing and presentation to T cells.
Query \(\PageIndex{1}\)
Query \(\PageIndex{1}\)
Key Concepts and Summary
- B lymphocytes or B cells produce antibodies involved in humoral immunity. B cells are produced in the bone marrow, where the initial stages of maturation occur, and travel to the spleen for final steps of maturation into naïve mature B cells.
- B-cell receptors (BCRs) are membrane-bound monomeric forms of IgD and IgM that bind specific antigen epitopes with their Fab antigen-binding regions. Diversity of antigen binding specificity is created by genetic rearrangement of V, D, and J segments similar to the mechanism used for TCR diversity.
- Protein antigens are called T-dependent antigens because they can only activate B cells with the cooperation of helper T cells. Other molecule classes do not require T cell cooperation and are called T-independent antigens.


