20.5: Chemical Defenses - Complement System
- Page ID
- 146388
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Identify the three mechanisms that trigger complement activation
- Describe the outcome of complement activation
The Complement System
The complement system is a group of plasma protein mediators that can act as an innate nonspecific defense while also serving to connect innate and adaptive immunity (discussed in the next chapter). The complement system is composed of more than 30 proteins (including C1 through C9) that normally circulate as precursor proteins in blood. These precursor proteins become activated when stimulated or triggered by a variety of factors, including the presence of microorganisms. Complement proteins are considered part of innate nonspecific immunity because they are always present in the blood and tissue fluids, allowing them to be activated quickly. Also, when activated through the alternative pathway (described later in this section), complement proteins target pathogens in a nonspecific manner.
Complement Activation
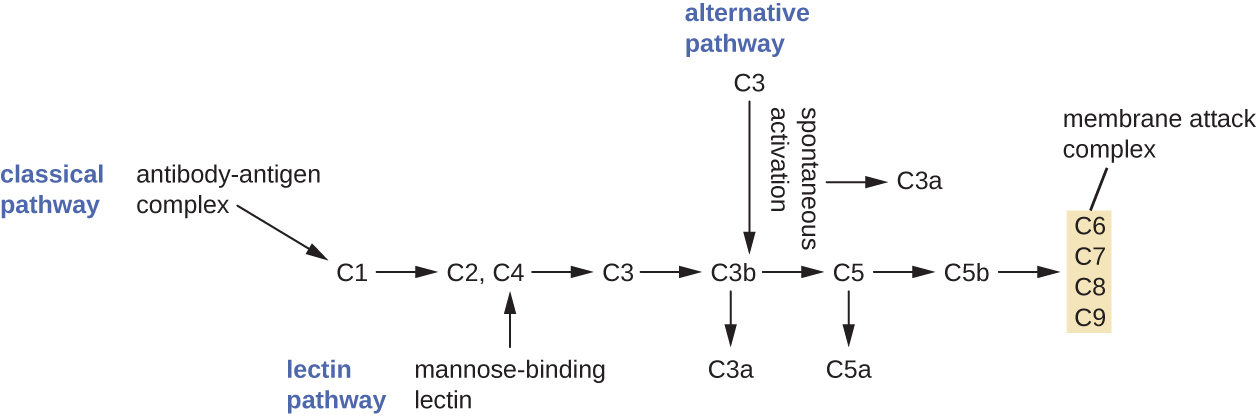
The process by which circulating complement precursors become functional is called complement activation. This process is a cascade that can be triggered by one of three different mechanisms, known as the alternative, classical, and lectin pathways.
The alternative pathway is initiated by the spontaneous activation of the complement protein C3. The hydrolysis of C3 produces two products, C3a and C3b. When no invader microbes are present, C3b is very quickly degraded in a hydrolysis reaction using the water in the blood. However, if invading microbes are present, C3b attaches to the surface of these microbes. Once attached, C3b will recruit other complement proteins in a cascade (Figure \(\PageIndex{2}\)).
The classical pathway provides a more efficient mechanism of activating the complement cascade, but it depends upon the production of antibodies by the specific adaptive immune defenses. To initiate the classical pathway, a specific antibody must first bind to the pathogen to form an antibody-antigen complex. This activates the first protein in the complement cascade, the C1 complex. The C1 complex is a multipart protein complex, and each component participates in the full activation of the overall complex. Following recruitment and activation of the C1 complex, the remaining classical pathway complement proteins are recruited and activated in a cascading sequence (Figure \(\PageIndex{2}\)).
The lectin activation pathway is similar to the classical pathway, but it is triggered by the binding of mannose-binding lectin, an acute-phase protein, to carbohydrates on the microbial surface. Like other acute-phase proteins, lectins are produced by liver cells and are commonly upregulated in response to inflammatory signals received by the body during an infection (Figure \(\PageIndex{2}\)).
Outcome of Complement Activation
Although each complement activation pathway is initiated in a different way, they all provide the same protective outcomes: opsonization, inflammation, chemotaxis, and cytolysis. The term opsonization refers to the coating of a pathogen by a chemical substance (called an opsonin) that allows phagocytic cells to recognize, engulf, and destroy it more easily. Opsonins from the complement cascade include C1q, C3b, and C4b. Additional important opsonins include mannose-binding proteins and antibodies. The complement fragments C3a and C5a are well-characterized anaphylatoxins with potent proinflammatory functions. Anaphylatoxins activate mast cells, causing degranulation and the release of inflammatory chemical signals, including mediators that cause vasodilation and increased vascular permeability. C5a is also one of the most potent chemoattractants for neutrophils and other white blood cells, cellular defenses that will be discussed in the next section.
The complement proteins C6, C7, C8, and C9 assemble into a membrane attack complex (MAC), which allows C9 to polymerize into pores in the membranes of gram-negative bacteria. These pores allow water, ions, and other molecules to move freely in and out of the targeted cells, eventually leading to cell lysis and death of the pathogen (Figure \(\PageIndex{2}\)). However, the MAC is only effective against gram-negative bacteria; it cannot penetrate the thick layer of peptidoglycan associated with cell walls of gram-positive bacteria. Since the MAC does not pose a lethal threat to gram-positive bacterial pathogens, complement-mediated opsonization is more important for their clearance.
Query \(\PageIndex{1}\)
Key Concepts and Summary
- The complement system involves numerous precursor proteins that circulate in plasma. These proteins become activated in a cascading sequence in the presence of microbes, resulting in the opsonization of pathogens, chemoattraction of leukocytes, induction of inflammation, and cytolysis through the formation of a membrane attack complex (MAC).


