13.6: Gene Regulation - Inducible Operon
- Page ID
- 159996
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the structure and components of the lac operon
- Differentiate between positive and negative regulation of the lac operon
- Identify the conditions under which the lac operon is activated or repressed
The lac Operon: An Inducible Operon
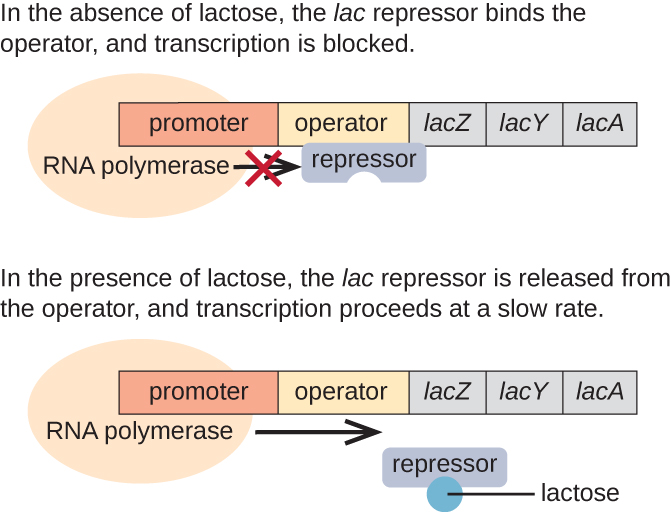
The lac operon is an example of an inducible operon that is also subject to activation in the absence of glucose (Figure \(\PageIndex{3}\)). The lac operon encodes three structural genes necessary to acquire and process the disaccharide lactose from the environment, breaking it down into the simple sugars glucose and galactose. For the lac operon to be expressed, lactose must be present. This makes sense for the cell because it would be energetically wasteful to create the enzymes to process lactose if lactose was not available.
In the absence of lactose, the lac repressor is bound to the operator region of the lac operon, physically preventing RNA polymerase from transcribing the structural genes. However, when lactose is present, the lactose inside the cell is converted to allolactose. Allolactose serves as an inducer molecule, binding to the repressor and changing its shape so that it is no longer able to bind to the operator DNA. Removal of the repressor in the presence of lactose allows RNA polymerase to move through the operator region and begin transcription of the lac structural genes.
Query \(\PageIndex{1}\)
Query \(\PageIndex{1}\)
The lac Operon: Activation by Catabolite Activator Protein
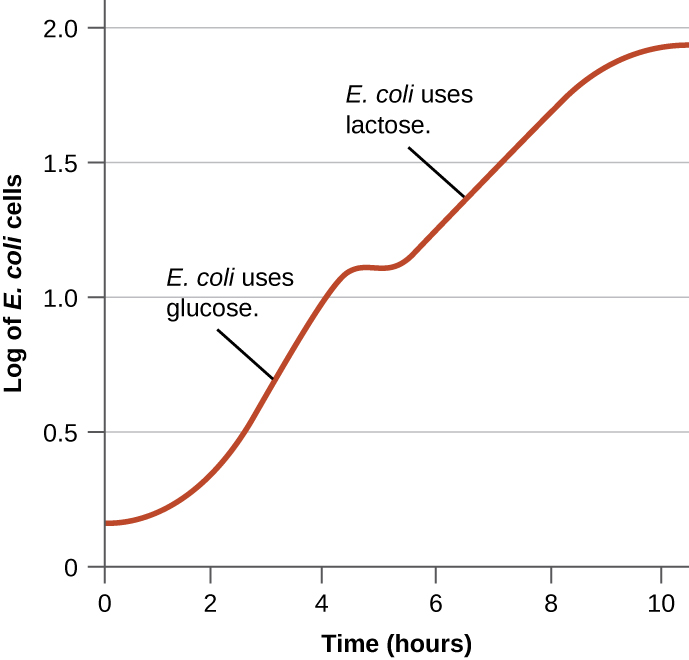
Bacteria typically have the ability to use a variety of substrates as carbon sources. However, because glucose is usually preferable to other substrates, bacteria have mechanisms to ensure that alternative substrates are only used when glucose has been depleted. Additionally, bacteria have mechanisms to ensure that the genes encoding enzymes for using alternative substrates are expressed only when the alternative substrate is available. In the 1940s, Jacques Monod was the first to demonstrate the preference for certain substrates over others through his studies of E. coli’s growth when cultured in the presence of two different substrates simultaneously. Such studies generated diauxic growth curves, like the one shown in Figure \(\PageIndex{4}\). Although the preferred substrate glucose is used first, E. coli grows quickly and the enzymes for lactose metabolism are absent. However, once glucose levels are depleted, growth rates slow, inducing the expression of the enzymes needed for the metabolism of the second substrate, lactose. Notice how the growth rate in lactose is slower, as indicated by the lower steepness of the growth curve.
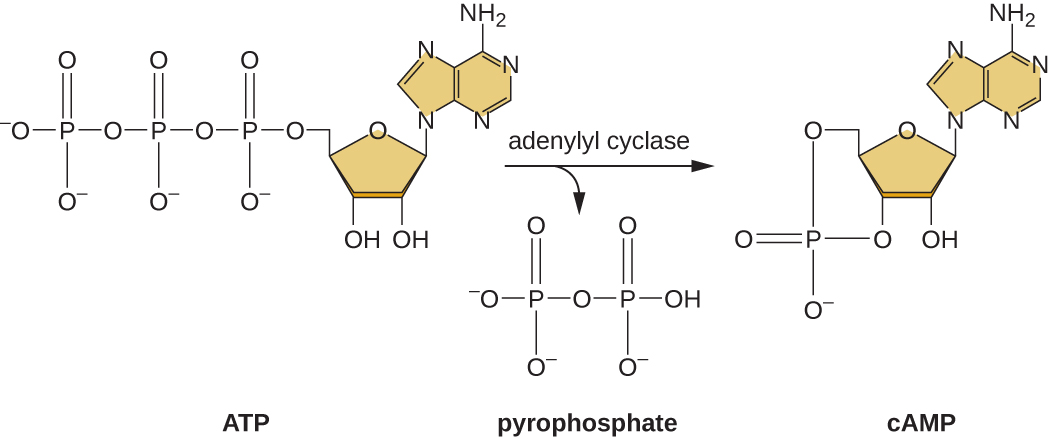
The ability to switch from glucose use to another substrate like lactose is a consequence of the activity of an enzyme called Enzyme IIA (EIIA). When glucose levels drop, cells produce less ATP from catabolism (see Catabolism of Carbohydrates), and EIIA becomes phosphorylated. Phosphorylated EIIA activates adenylyl cyclase, an enzyme that converts some of the remaining ATP to cyclic AMP (cAMP), a cyclic derivative of AMP and important signaling molecule involved in glucose and energy metabolism in E. coli. As a result, cAMP levels begin to rise in the cell (Figure \(\PageIndex{5}\)).
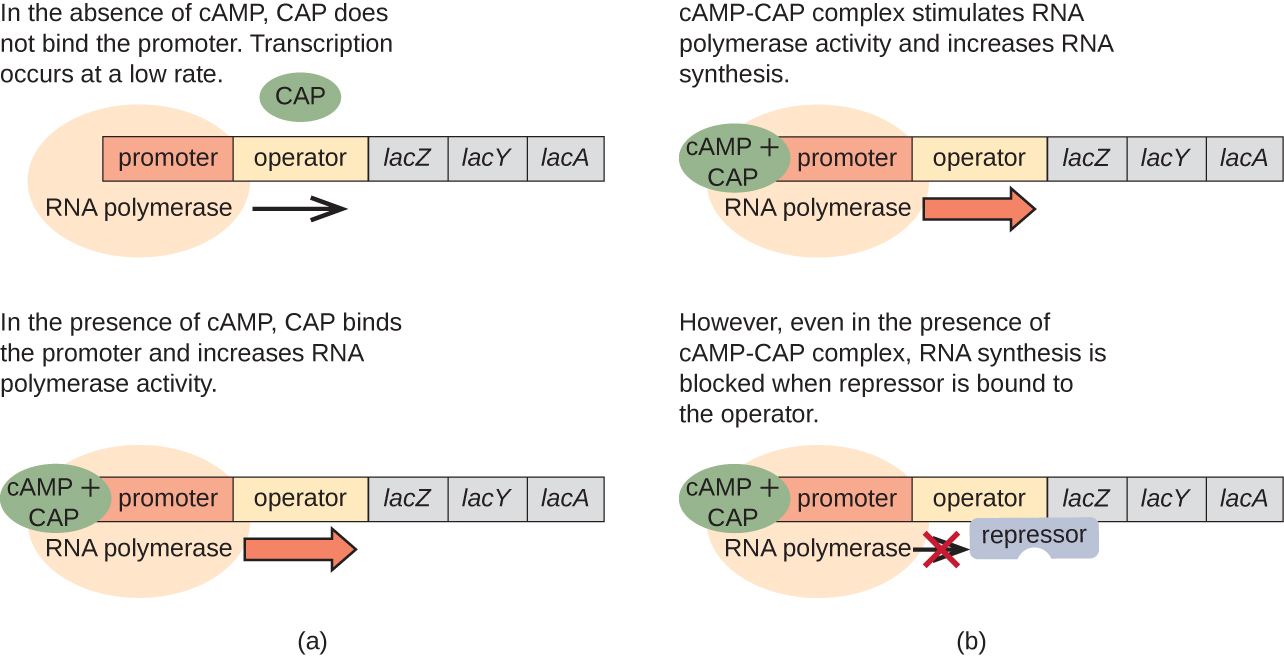
The lac operon also plays a role in this switch from using glucose to using lactose. When glucose is scarce, the accumulating cAMP caused by increased adenylyl cyclase activity binds to catabolite activator protein (CAP), also known as cAMP receptor protein (CRP). The complex binds to the promoter region of the lac operon (Figure \(\PageIndex{6}\)). In the regulatory regions of these operons, a CAP binding site is located upstream of the RNA polymerase binding site in the promoter. Binding of the CAP-cAMP complex to this site increases the binding ability of RNA polymerase to the promoter region to initiate the transcription of the structural genes. Thus, in the case of the lac operon, for transcription to occur, lactose must be present (removing the lac repressor protein) and glucose levels must be depleted (allowing binding of an activating protein). When glucose levels are high, there is catabolite repression of operons encoding enzymes for the metabolism of alternative substrates. Because of low cAMP levels under these conditions, there is an insufficient amount of the CAP-cAMP complex to activate transcription of these operons. See Table \(\PageIndex{1}\) for a summary of the regulation of the lac operon.
| Glucose | CAP binds | Lactose | Repressor binds | Transcription |
|---|---|---|---|---|
| + | – | – | + | No |
| + | – | + | – | Some |
| – | + | – | + | No |
| – | + | + | – | Yes |
Watch an animated tutorial about the workings of lac operon here.
Query \(\PageIndex{1}\)
Query \(\PageIndex{1}\)
Key Concepts and Summary
- The lac operon is a classic example an inducible operon. When lactose is present in the cell, it is converted to allolactose. Allolactose acts as an inducer, binding to the repressor and preventing the repressor from binding to the operator. This allows transcription of the structural genes.
- The lac operon is also subject to activation. When glucose levels are depleted, some cellular ATP is converted into cAMP, which binds to the catabolite activator protein (CAP). The cAMP-CAP complex activates transcription of the lac operon. When glucose levels are high, its presence prevents transcription of the lac operon and other operons by catabolite repression.


