4.1: Introduction: The Big Picture
- Page ID
- 21534
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
We already began examining Developmental Genetics and and got a taste of the complexity of genetic interactions. For example, the Drosophila gene Bicoid is transcribed into mRNA in maternal cells and pumped into the embryo. There its expression is regulated by localization signals on the mRNA, as well as degradation and localization of the protein. Bicoid protein itself acts as both a regulator of transcription and translation of downstream genes. The end effect is an embryo with different genes expressed along the A/P axis.
Now we will take a step back and look at a broader picture of Developmental Genetics, from the point of view of signaling pathways. A signaling pathway allows cells to communicate with their external environment. In Developmental Biology, this is usually cell-cell interactions. These kinds of interactions are incredibly important because each cell needs to follow its own developmental trajectory in coordination with all the cells around it. For example, imagine a growing mammalian limb bud. Each cell in the limb bud needs to know if it is on the thumb or pinky side of the limb and how close to the body it is. Not only that, it also needs to know how long development has progressed. Imagine if an osteogenic cell started forming bone matrix early in limb development. It would not undergo enough cell divisions to make the correct number of osteoblasts, leading to too little bone and an animal with a malformed limb.
The Genetic Toolkit
(An introductory PDF can be found here: Rediscovering Biology www.learner.org/courses/biology/pdf/7_gendev.pdf)
One of the major findings of Evo-Devo is that organisms share a common set of genes to build their very diverse bodies. In particular, animals share a suite of "body plan" genes that perform similar functions in development. Mixing and matching different gene expression patterns across tissue and cell types seems to be the primary mechanism for generating the unique bodies we see in each animal phylum. Novel evolutionary features (like spines, feathers, chambered hearts, tentacles, etc) appear to rely on the reuse of toolkit genes at new times and places in development (see Endless Forms for more details). Below, we will start to take a look at the genetics behind this genetic toolkit - how do these genes interact with each other?
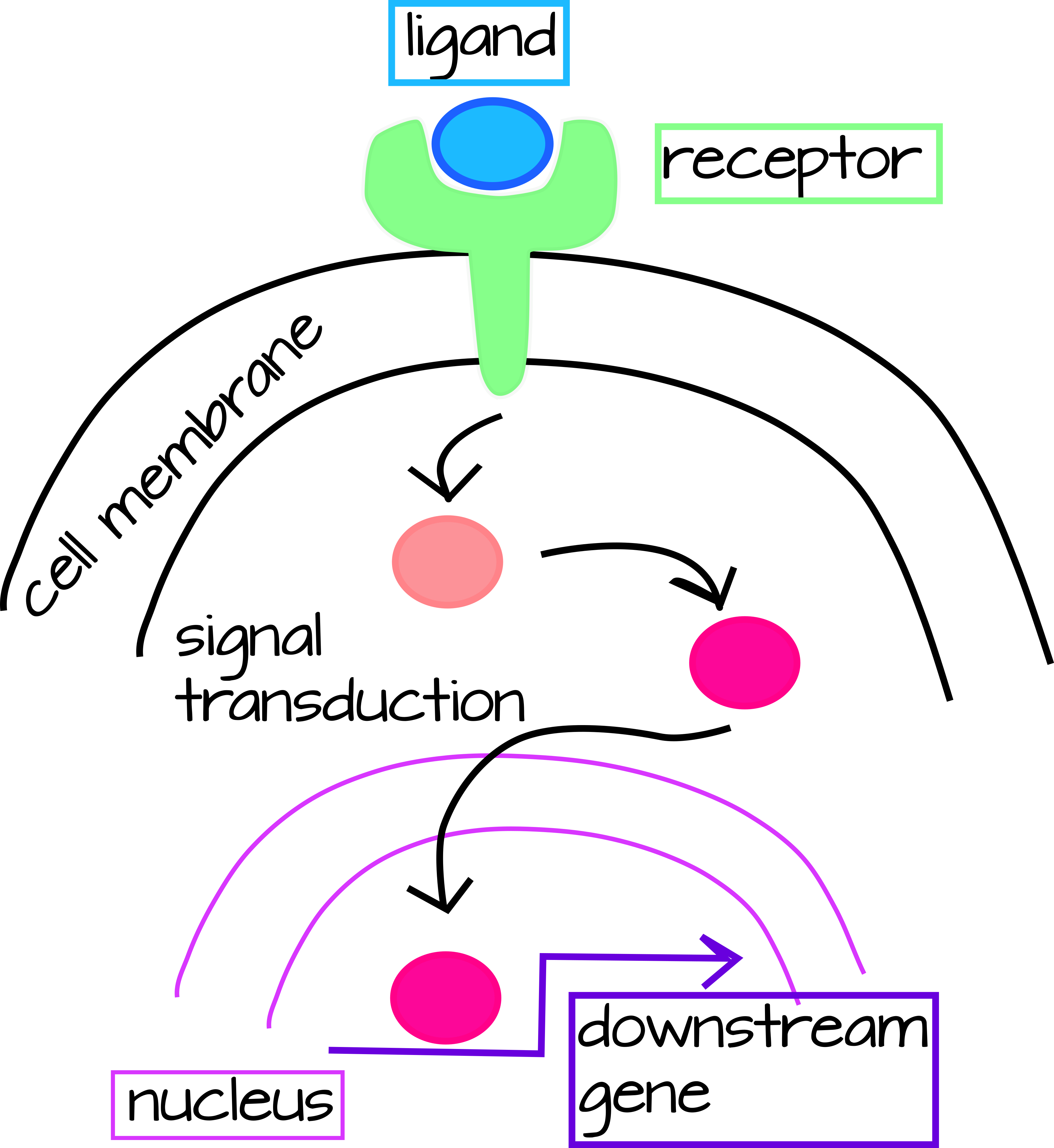
In Evo/Devo, there are two main ways we can think about genetic interactions, but since they are interconnected so we will consider both simultaneously. One is Signal Transduction Pathways, which have an external signal (in Developmental Biology this is usually a secreted protein) that is usually received by a cell-membrane receptor that only certain cells express. When the ligand (secreted protein) and receptor bind, this triggers a cascade of protein modifications that lead to transcriptional activation of downstream genes (Figure 1).
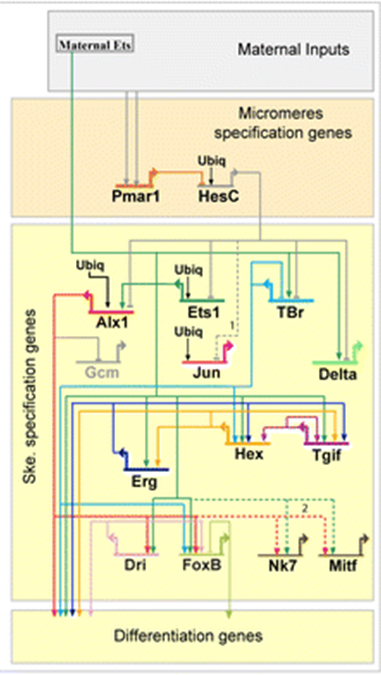
The second way Evo/Devo thinks about signaling is Gene Regulatory Networks (GRNs). This includes the signal transduction pathway and its downstream effects on gene expression. If we turn up the expression of one gene, what other genes does it affect? GRNs are often represented as wiring diagrams with arrows and bars showing the effect of proteins on the expression of downstream genes (Figure 2).
 |
.png?revision=1) |
| Figure 1: Generic signal transduction pathway. An extracellular ligand binds to a membrane-bound receptor triggering the activation of a signal tranduction cascade in the cytoplasm. This cascade may be simple (as in the case of the Notch pathway below) or very complex requiring multiple steps and types of input. The final step of the transduction cascade is translocation of a cytoplasmic transcription factor to the nucleus where it will activate or inhibit the transcription of a downstream gene. An editable svg file of this figure can be downloaded at https://scholarlycommons.pacific.edu/open-images/18/ | Figure 2: Partial Gene Regulatory Network wiring diagram for the specification of skeleton-forming mesoderm in the sea urchin, Strongylocentrotus purpuratus. Each gene is represented by a horizontal line attached to an arrow of the same unique color. The horizontal line indicates the cis-regulatory region of the gene and the arrow indicates transcription of the mRNA coding region. Lines emanating from the mRNA coding region indicate how the protein product of the gene regulates other downstream genes. Inputs onto the cis-regulatory regions by the protein products of other genes may be activating (arrow) or inhibitory (bar). “Ubiq” is unknown activators expressed in all cells. Figure from Dylus et al1 Published under a CC BY 4.0 license. Figure has been cropped from original. |
The big picture: Cell fates are progressively restricted as development proceeds
Over the course of development all organs, tissues, and cell-types must be specified. In animals this is accomplished by repeated fate restriction2. For example, the totipotent embryo undergoes several rounds of division to form a blastula (ball of cells). Some of these cells will take on a mesodermal fate. At this point, they can become any type of mesodermal cell. In some animals experiments have been performed to show that a generic mesodermal cell can be induced by its neighbors to develop into a huge variety of mesodermal cell types3. Over time, however, these mesodermal cells divide, undergo shape changes, migrate, and differentiate. As they receive signals from their environment (mostly the cells around them) they differentiate into more specific cell types. For example, Figure 3 shows a mesodermal cell differentiating into muscle, bone, and blood cells. As time progresses, different muscle progenitor (myogenic) cells will differentiate into different types of muscle, for example smooth and striated. Each of these cell-fate decisions is dependent on the gene expression pattern in the cell, which is dependent on the cell's history and its neighbors.

We call this type of progressive differentiation "hierarchical" with more generic cell types (like mesoderm, Figure 3) at the top of the hierarchy and more specific cell types (like neutrophils) at the bottom4. As development progresses, we move down the hierarchy. Development begin axis specification, these axes are soon read out into regions - for example head, trunk/abdomen, and tail. Within these different regions tissues get specified, first at the level of germ-layers and later at more specific levels. For example, ectoderm splits into neuroectoderm and epidermal ectoderm depending on the location of the tissue. Next, cell-types within the tissues are specified. For example, neuroectoderm can become neurons or glial cells. Both of these cell types later differentiate into the many specific types of neurons and glia. At this point, cells are more or less terminally differentiated - their adult fates are specified and they cannot return to pluripotent forms. However, most cells are still receptive to the environment and can respond to environmental signals even once they are terminally differentiated. We refer to this as cellular plasticity.
The GRN in Figure 2 mirrors this hierarchy, with early acting genes at the top specifying more general tissues and later acting genes at the bottom specifying specific cell types. However, it is important to note that the genes acting at these different hierarchical levels are reused at other levels of the hierarchy and in other developing tissues. In fact, core parts of GRNs and signal transduction pathways are reused throughout development and have different outputs depending on their context.


