4.5: Protein-Protein Recognition Probed Using a Yeast Transcriptional Activator System
- Page ID
- 18144
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Fields, S. and Ok-kyu, S. Nature (1989) 340:245-246
Background
The yeast genome contains Upstream Activator Sequences (UAS's)
UAS are located 5' to the coding region of a gene and are regulatory regions - they bind transcriptional activator proteins. These transcriptional activator proteins, when bound to UAS, stimulate transcription (i.e. they may contain binding sites for RNA polymerase
Yeast gene, protein nomenclature
Wild type genes are designated with capital letters in italics (e.g. GAL4). Mutant alleles of the genes are indicated in lowercase italic (e.g. gal4) and encoded proteins are indicated by roman type, with the first letter capitalized (e.g. Gal4)
Growth of yeast in galactose containing media
Incubation of wild type yeast in galactose results in >1,000 fold increase in the mRNA level for enzymes involved in galactose metabolism. This increase in mRNA levels is not observed in gal4 mutants. The Gal4 protein is not one of the enzymes involved in galactose metabolism, rather, it a appears to be a transcriptional activator protein which specifically activates the transcription of genes involved in the galactose metabolic pathway. Gal4 protein binds to a specific 17 nucleotide long region of DNA located in the 5' region of genes involved in galactose metabolism. These regions are termed UASGAL Binding of Gal4 to UASGAL increases transcription from the nearby promoter
Gal4 Protein
- Single polypeptide chain which contains two domains:
.png?revision=1&size=bestfit&width=455&height=280)
Figure 4.5.1: Gal4 protein domains
These domains are functionally separable in the polypeptide chain (Figure 4.5.2). The amino-terminal (1-74) domain by itself can bind to UASGAL sequences but cannot activate transcription. The carboxyl-terminal (738-881) domain contains the activating region but cannot activate transcription because it fails to localize to the UASGAL region (i.e. near the promoter region)
.png?revision=1&size=bestfit&width=583&height=184)
Figure 4.5.2: Gal4 polypeptide chain
Experimental Design
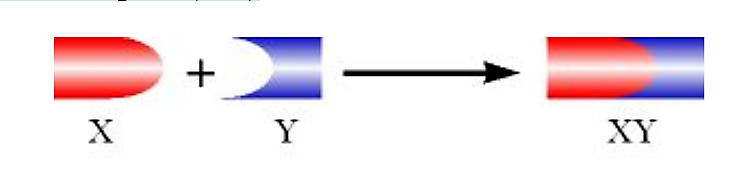
Imagine we have two proteins (X and Y), which normally form a stable complex (XY)
.png?revision=1&size=bestfit&width=442&height=107)
Figure 4.5.3: Complex
If we make a Gal4(1-74)- X fusion protein and a Y -Gal4(738-881) fusion protein we can potentially form a functional Gal4 transcription activator (i.e. a Gal4 activator region in complex with a UASGAL region
.png?revision=1&size=bestfit&width=502&height=151)
Figure 4.5.4: Formation of Gal4 transcription activator
This fusion protein complex can act as a functional transcriptional activator for the galactose metabolic enzymes
.png?revision=1&size=bestfit&width=521&height=313)
Figure 4.5.5: Protein complex acting as transcription activator
To test this hypothesis, the authors used the following system
- The protein complex involved the two proteins SNF1 and SNF4. The SNF1 protein is a serine-threonine specific protein kinase, and the SNF4 protein is known to be physically associated with SNF1 and is essential for function
- 5 different gene constructs involving Gal4 and SNF1 and SNF4 were made:
.png?revision=1&size=bestfit&width=571&height=261)
Figure 4.5.6: Gene constructs with Gal4, SNF1, SNF4
- These gene constructs were inserted into two different yeast plasmids:
- All constructs containing amino terminal fusions of GAL4 were put into plasmids containing the HIS3 gene (coding for a biosynthetic enzyme necessary for the production of the amino acid histidine).
- All constructs containing carboxy terminal fusions of GAL4 were put into plasmids containing the LEU2 gene (coding for a biosynthetic enzyme necessary for the production of the amino acid leucine)
- These plasmids were inserted into a host yeast (GGY1::171)which had the GAL4 gene deleted, the LACZ gene deleted, the HIS3 and LEU2 genes mutated to be non-productive and which also contained a GAL1-LACZ gene fusion (i.e. a LacZ protein driven from the GAL1 promoter which has a UASGAL. This host is thus DGAL4, DLACZ, his3, leu2, gal1-lacZ.
- Thus, selective pressure for the amino terminal GAL4 fusions can be maintained by eliminating histidine in the growth media (yeast relies on the HIS3 gene on the plasmid to make histidine)
- Likewise, selective pressure for the carboxy terminal GAL4 fusions can be maintained by eliminating leucine in the growth media (yeast relies on the LEU2 gene on the plasmid to make leucine)
- The host can harbor both kinds of plasmids (in which case the media would lack both histidine and leucine)
- The expression of the fusion constructs on the plasmid is actually regulated by galactose. Adding galactose results in expression of the fusion constructs. However, since the host is DGAL4 all the other galactose metabolic genes (e.g. GAL1) will not be up-regulated in response to added galactose.
The presence of a functional Gal4 protein in the GGY1::171 yeast host will result in the following:
- Expression of the LacZ protein (contained in the host GAL1-LACZ gene fusion) will occur. Thus, in the presence of XGAL substrate the yeast colony will turn blue
- All the host genes which contain an UASGAL will be up-regulated. This will include all the wild-type genes involved in galactose metabolism (e.g. GAL1)
Experimental Results
|
|
|
|---|---|
| 1. None |
|
| 2. GAL4(1-881) |
|
| 3. GAL4(1-147) |
|
| 4. GAL4(1-147)-SNF1 |
|
| 5. SNF4 |
|
| 6. SNF4- GAL4(768-881) |
|
| 7. GAL4(1-147)-SNF1; SNF4-GAL4(768-881) |
|
| 8. GAL4(1-147)-SNF1; SNF4 |
|
| 9. GAL4(1-147); SNF4-GAL4(768-881) |
|
- Functional Gal4 protein resulted in expression of the host GAL1-LACZ gene
- Co-expression of plasmids containing GAL4(1-147)-SNF1; SNF4-GAL4(768-881) fusion proteins resulted in expression of the host GAL1-LACZ gene, indicating the presence of functional Gal4 protein (i.e. complex formation of the Gal4 fusion proteins)
- The level of LacZ protein was less, however, indicating that the SFN1- and SFN4- Gal4 fusion protein complex was a less efficient as a transcription activator than wild type Gal4 protein
- There was some low level LacZ protein activity with the GAL4(1-147)-SNF1; SNF4 protein complex. This suggests that the SNF4 may have some minor affinity for RNA polymerase
Conclusions
This method may prove useful for identifying proteins which can form stable complexes (i.e. identifying specific protein:protein interactions). For example, if we want to know which proteins bind to a certain receptor we can make a GAL4(1-147)-receptor fusion protein and screen a cDNA library for potential binding proteins by constructing cDNA-GAL4(768-881) fusion constructs. Potential binding proteins from such a library can be screened by looking for blue colonies with XGAL substrate, or selected for by growing the yeast in media with galactose as the carbon source.


