6.7: Ribozymes - RNA Enzymes
- Page ID
- 21160
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Ribozymes
Any molecule that displays any of the catalytic motifs seen in the earlier chapters (general acid/base catalysis, electrostatic catalysis, nucleophilic catalysis, intramolecular catalysis, and transition state stabilization) can be a catalyst. So far we have examined only protein catalysts. These can fold to form unique 3D structures, which can have active sites with appropriate functional groups or nonprotein "cofactors" (metal ions, vitamin derivatives) that participate in catalysis. There is nothing special about the ability of proteins to do this. RNA also can form secondary and tertiary structures as we will see in Chapter 8. RNA molecules that act as enzymes are called ribozymes.
We are presenting the section before Chapter 8 for a few reasons. Most readers have encountered the structures of RNA and DNA before. They most likely know about three different types of RNA, ribosomal RNA (rRNA), transfer RNA (tRNA) and messenger RNA (mRNA). Likewise, they know from introductory biology classes the essential dogma of biology: (DNA, the holder of the genetic code) is transcribed into RNA which is translated into a protein sequence. Finally, most have studied (even at the high school level) that DNA of many species has exons and introns (intervening sequences), the latter that are spliced out of RNA transcripts to form mature RNA. In this section, we will discuss the catalytic properties of ribozymes, so the introductory background we just mentioned, although important, takes a "second" seat to the chemistry of catalysis, which is the main topic of Chapter 6.
There are 12 classes of ribozymes
- small self-cleaving RNAs (9 classes)
- Group I introns
- Group II introns
- Ribonuclease P
The large ribonucleoprotein nanoparticles, the spliceosome and ribosome, are also functionally ribozymes as well.
The term ribozyme is used for RNA that can act as an enzyme. Ribozymes are mainly found in selected viruses, bacteria, plant organelles, and lower eukaryotes. Ribozymes were first discovered in 1982 when Tom Cech’s laboratory observed Group I introns acting as enzymes. This was shortly followed by the discovery of another ribozyme, Ribonuclease P, by Sid Altman’s laboratory. Both Cech and Altman received the Nobel Prize in chemistry in 1989 for their work on ribozymes.
Ribozymes can be categorized based on size. Small ones, which usually don't require metal ions for activity, vary from 30-150 nucleotides while large ones can be a few thousand nucleotides in length. This translates into approximate molecule weights (using this formula for single-stranded RNA: (# nucleotides x 320.5) + 159.0) into 9800 for a 30-mer and 640,000 for a 2000 mer, typical of small and very large proteins/protein complexes, respectively. into two groups depending upon their size – small and large. Large ribozymes, which required metal ions for activity, can vary in size from a few hundred to several thousand nucleotides. Examples of small ones include hammerhead, viroid, hairpin, and riboswitch ribozymes. Examples of large ones include type I and II self-spicing introns, bacterial ribonuclease P, as well as the RNA in spliceosomes and ribosomes. Many also are not really true enzymes since they catalyze their own cleavage, although some can cleave presented RNA substrates. Large ones act as true catalysts.
Since RNAs can carry genetic information and act as enzymes, they probably evolved before proteins, which require nucleic acids for their synthesis. In addition, DNA required a special enzyme (ribonucleotide reductase) encoded by DNA to reduce the 2'OH to a 2'H. Artificial ribozymes have been made to catalyze many reactions that require protein enzymes. We will explore some ribozymes in several classes.
Small self-cleaving RNAs
We'll consider four small self-cleaving RNAs- hammerhead, viroid and hairpin ribozymes as well as the glucosamine-6-phosphate riboswitch (glmS). All catalyze the cleavage of an internal phosphodiester bond (cis catalysis) or in a presented substrate (trans catalysis) by a transesterification reaction. Internal cis catalysis reactions cleave the ribozyme into two fragments, which inactivates their catalytic activity. In that sense, they don't act as true catalysts since they engage in only one cycle of cleavage. Trans catalysis in which a substrate RNA binds to and is cleaved by the ribosome would be considered true catalysis.
The cleavage reaction in an internal cis cleavage is an SN2 trans-esterification reaction as shown in Figure \(\PageIndex{1}\).
In the reaction, an adjacent general base (:B) abstracts a proton of the C'2 OH of nucleotide N1. The resulting 2' O- acts as a nucleophile in a SN2 reaction and attacks the δ+ phosphorous in the phosphodiesterase bond, forming a pentavalent, trigonal pyramidal sp3d hybridized intermediate/transition state, which collapses breaking the phosphodiesterase bond between nucleotide N1 and N2. This is an inline mechanism in that the incoming nucleophilic O in C2' and the exiting one on the O of 5' CH2OH of nucleotide 2 are axial to each other separated by 1800. A general acid, BH, facilitates the departure of the exiting nucleophile by its protonation. Bound metal ions may facilitate the reaction and can be considered cofactors but might be more involved in maintaining a catalytically-active structure. Self-cleaving small RNAs are also found in humans and may be part of long noncoding RNAs.
a. Hammerhead RNA
The hammerhead ribozyme is a small RNA ribozyme with a conserved core with three helical stems. It has a structure similar to the head of a hammerhead shark. One predicted secondary structure of a hammerhead ribozyme is shown below in Figure \(\PageIndex{2}\)
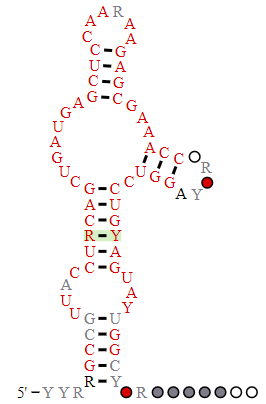.png?revision=1)
_legend.png?revision=1)
A possible trigonal pyramidal intermediate/transition state in the Hammerhead ribosome is shown in Figure \(\PageIndex{3}\).
A deprotonated G-12 in the ribozyme probably acts as a general base that activates the 2'-OH to form the incoming nucleophile that attacks the trans-substrate RNA. The 2′-OH of G-8 in the ribozyme appears to hydrogen bond to the 5′-O of the departing nucleophile in the substrate where bond scission occurs. The ribozyme increases the rate by 1000-fold.
Figure \(\PageIndex{4}\) shows an interactive iCn3D model of the full-length Schistosoma manson catalytically active hammerhead ribozyme (3dz5). This is an example of a ribozyme that acts in trans as the cleaved phosphodiester bond is a bound RNA single-stranded substrate.
Figure \(\PageIndex{4}\): Full-length Schistosoma manson catalytically active hammerhead ribozyme (3dz5). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?qNBw2MY1RZ7xKogH7
The numbering system is a bit different in the iCn3D model above. PDB requires sequential numbering whereas ribozymes sequences are numbered in discontinuous ways. Core residues that are conserved are given common numbers. However, the different projecting double-stranded RNA regions, which vary among ribozymes, are numbered differently. The G8 and G12 in Figure \(\PageIndex{3}\) are numbered G20 and G36, respectively.
b. Viroids
Viroids are small single-stranded circular RNAs that infect plant cells. They are not packaged with viral capsid protein. Some enter cells along with viruses and are called virusoids or viroidlike satellite RNAs. Intrastrand pairing occurs and they are synthesized as tandem repeats containing multiple adjacent copies of the viroid. These repeats are cut and ligated to form the individual mature viroid by internal ribozymes sequences. One example is the hepatitis delta virus (HDV), a satellite of the hepatitis B virus. A possible mechanism of catalysis of the hepatitis delta virus ribozyme involving general acid/base catalysis is shown in Figure \(\PageIndex{5}\).
c. Hairpin ribozyme
Hairpin ribozymes are encoded by the satellite RNA of plant viruses. They are about 50 nucleotides long, and can cleave itself internally, or, in a truncated form, can cleave other RNA strands in a transesterification reaction. The structure consists of two domains, stem A required for binding (self or other RNA molecules) and stem B, required for catalysis. Self-cleavage in the hairpin ribozyme occurs in stem A between an A and G bases (which are splayed apart) when the 2' OH on the A attacks the phosphorous in the phosphodiester bond connecting A and G to form a pentavalent intermediate.
Rupert et al solved the crystal structure of a hairpin ribozyme with a non-cleavable substrate analog containing a 2'-O-CH3 group on the ribose. This acts as a nucleophile in the transesterification cleavage of RNA.
A38 in Stem B appears to be able to interact with the products (the cleaved A now in the form of a cyclic phosphodiester with itself) and the departing G, and with a transition state pentavalent analog of the sessile A-G bond in which the phosphodiester linking A and G in the substrate is replaced with a pentavalent vanadate bridge between A and G. This is Illustrated in Figure \(\PageIndex{6}\).
However, A 38 does not appear to react with the sessile A -G groups in the normal substrate, indicating that the main mechanism used by this ribozyme is transition state binding. Since RNA molecules have fewer groups available for acid/base and electrostatic catalysis (compared to protein enzymes), ribozymes, presumably the earliest type of biological catalyst, probably make more use of transition state binding as their predominant mode of catalytic activity.
Figure \(\PageIndex{7}\) shows an interactive iCn3D model of the hairpin ribozyme in the catalytically-active conformation (1M5K).
V2.png?revision=1)
The hairpin ribozyme is shown in cyan sticks and the inhibitor substrate in brown sticks. The inhibitor contains a 2'-O-methyl adenosine (A2M12), so it can not be cleaved and instead acts as an inhibitor. The A38 shown in the catalytic mechanism is labeled A57 in the iCn3D.
d. Glucosamine-6-phosphate riboswitch (glmS):
A novel use of ribozymes was recently reported by Winkler et. al. They discovered that the 5' end of the mRNA of the gene glmS (from Gram-positive bacteria) is a ribozyme. The GlmS gene encodes glucosamine-6-phosphate synthetase (GlmS), which catalyzes the reaction of fructose-6-phosphate and glutamine to glucosamine-6-phosphate (GlcN6P) and glutamate. This is the first committed step in bacterial cell wall synthesis. Glucosamine-6-phosphate binds to the ribozyme (3' end of the mRNA) and acts as a cofactor leading to self-cleavage of the ribozyme. What an amazing mechanism for pathway inhibition. At high GlcN6P concentrations, it binds to the ribozyme, inhibiting its own synthesis. G40 in the active site appears to act as a general base. Figure \(\PageIndex{8}\) shows an interactive iCn3D model of GlmS Ribozyme Bound to Its Catalytic Cofactor, glucosamine 6 phosphate (GlcN6P) (2NZ4).
.png?revision=1&size=bestfit&width=510&height=222)
Riboswitches are discussed in greater detail in Chapter 28.1: Regulation of Gene Expression in Bacteria.
Group I and Group II Introns
Introns present in RNA molecules must be removed to form mature RNA. In humans, about 80% of introns are less than 200 nucleotides long, but some can be 10,000 nucleotides or longer in length. Before we discuss introns, we'll provide a quick background on RNA splicing. There are two major types of self-splicing introns, Groups I and II. Other introns are removed by a ribonucleoprotein called the spliceosome. Some call these Group III introns. A simple two-step mechanism for the self-splicing Group I and II introns is shown in Figure \(\PageIndex{9}\).
Both required first a scission of the RNA strand followed by ligation of the two exons to form the mature RNA. Note that Group I introns require an external guanosine nucleophile and the removed intron forms a circular RNA when removed. In contrast, in Group II introns, an internal A residue acts as the first nucleophile in the scission reaction and the intron on removal forms a branched lariat structure.
A simple two-step mechanism for Group II introns which are spliced out from pre-mRNA in the nucleus by a ribonucleoprotein complex called the splicesome is shown in Figure \(\PageIndex{10}\).
Figure \(\PageIndex{10}\): Two steps of canonical RNA processing, from pre-mRNA to spliced RNA and the branched lariat intron. https://commons.wikimedia.org/wiki/F...g_reaction.svg. Creative Commons Attribution-Share Alike 3.0 Unported
Note that the mechanism is extremely similar to the auto-removal of Group II introns, suggesting an evolutionary relationship between the two.
Figure \(\PageIndex{11}\) below shows a more detailed view of the catalytic cycle of the spliceosome. Five small ribonucleoproteins (U1, U2, U4/U6 and U5 snRNPs) assemble on the nuclear pre-mRNA and facilitate the removal of the intro,n but the main mechanism involves ribozyme activity.
Group I introns
These are found in bacteria, lower eukaryotes (including mitochondrial and chloroplast RNA) and higher plants and are in ribosomal RNA (rRNA), mRNA and tRNA. They are also found in Gram-positive bacteria bacteriophages (viruses that attack bacteria). As shown in Figure \(\PageIndex{9}\), they require guanosine as a cofactor and have a single active site for both scission and ligation to produce the mature mRNA, tRNA or rRNA. Mg2 is required not for catalysis per se but to maintain the correct tertiary structure of the ribozyme with the correct secondary structure. In Group I introns, the splicing reaction is initiated by a guanosine cofactor. They have one active site that catalyzes the initial cleavage of the phosphodiester bond and the final religation after cleavage.
The Group I catalytic core from Tetrahymena thermophila has two domains. A cleft is formed between them when they pack which can bind the short helix with the 5' splice site. and the guanosine cofactor. This "active" site is preformed without substrates similar to the active sites of protein enzymes. Figure \(\PageIndex{12}\) shows the secondary structure and the reaction of the group I intron ribozyme from Tetrahymena.
The intron runs between the two triangles, which show the 5' (filled triangle) and 3' (open triangle) splice sites. The orange P1 helices contain the 5′-splice site (G:U). The P10 helix (grey), P9.0 helix (green), and the P9.2 helix (blue) are structured to present the appropriate 3′-splice site. During splicing (from (A) to (B)), the P1 helix extension (orange) is opened to expose the 3′-hydroxyl group of the terminal uridine at the 5′-splice site. The P10 helix then facilitates a conformational change, in which the 3′-exon (upper dashed line) is positioned adjacent to the 5′-exon (lower dashed line), allowing the nucleophilic attack of the 3′-uridine the 3′-splice site, joining 5′-exon and 3′-exon.
Figure \(\PageIndex{13}\) shows an interactive iCn3D model from cryoEM of the full-length holo L-16 ScaI Tetrahymena ribozyme (7EZ2).
Figure \(\PageIndex{13}\): Holo L-16 ScaI Tetrahymena ribozyme (7EZ2) (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...PhK3BSbFLDnDt8
Three sets of coplanar bases are found in the active site. These include C262, A263 and G312 (top layer, brown), G264, C311 and the ωG - labeled G3 in the iCn3D model- (cyan layer) and A261, A265 and U310 (bottom layer red). The ωG - labeled G3 - is the nucleophile.
The structure is nearly identical to the apo-form of the ribozyme with just an internal guide RNA sequence undergoing a large change and the guanosine binding site undergoing a small shift on binding RNA substrates.
Group 2 Introns
Group II introns are found in mRNA of bacteria and some Archaea, in rRNA, tRNA, and mRNA of chloroplasts and mitochondria, and in fungi, plants, and protists. No Class 2 introns appear to be found in eukaryotic genomes. Some of these introns are in gene-encoding proteins, but most are in bacterial noncoding sequences. In Group II introns, the splicing reaction is initiated by an adenosine cofactor, as shown in Figure \(\PageIndex{9}\).
What's especially interesting about group II introns is that they can reinsert in DNA so that can be considered to be mobile genetic elements. A maturase/reverse transcriptase enzyme (562 amino acids) is associated with the intron, which helps stabilize the active site of the intron for the reversible session and ligation of the intron. Reintegration of the excised branched lariat intron into DNA is called retrotransposition (copy/paste). Mitochondrial and chloroplast Group 2 intron have lost their mobility and act as classic introns.
The structures of a group II intron from Thermosynechococcus vestitus (a cyanobacteria) before and after integration have been determined. A branch-site domain VI helix swings 90°, enabling DNA integration. The maturase/revere transcriptase protein assists excision of the intron through the interaction of domain VI of that intron that positions the key adenosine for branched lariat formation during forward splicing, as shown in Figure \(\PageIndex{9}\). The changes in the structure of the group II intron retroelement before (6ME0) and after DNA integration (6MEC) are shown in Figure \(\PageIndex{14}\).

The target DNA before integration is shown in cyan spacefill and in orange spacefill after integration. The protein is shown as a colored cartoon. The Group II intron is 867 nucleotides and the sense target DNA is 47 nucleotides in length.
Spliceosomal Introns
A comparison of Figure \(\PageIndex{9}\) through Figure \(\PageIndex{11}\) show the similarities between spliceosomal introns and group II self-splicing introns. Spliceosomal and group II self-splicing introns are structurally and mechanistically homologous, right down to the stereochemistry of the splicing reaction. In eukaryotes, introns in pre-mRNA are removed by splicing and subsequent exon ligation, releasing the intron as a branched lariat molecule. This reaction is performed by the spliceosome, a large nuclear ribonucleoprotein (RNP) complex. Spliceosomes remove introns and splice the exons of most nuclear genes. They are composed of 5 kinds of small nuclear RNA (snRNA) molecules and over 100 different protein molecules. It is the RNA — not the protein — that catalyzes the splicing reactions. The molecular details of the reactions are similar to those of Group II introns, and this has led to speculation that this splicing machinery evolved from them. Figure \(\PageIndex{15}\) shows two views of the cryo-EM structure of the human-activated spliceosome (the Bact complex)
C
Figure \(\PageIndex{15}\): Cryo-EM structure of the human-activated spliceosome (the Bact complex). Zhang et al. https://www.nature.com/articles/cr201814.pdf. Creative Commons Attribution 4.0 Unported License. http:// creativecommons.org/licenses/by/4.0/
There are 52 proteins, 3 small nuclear RNAs (snRNA), and one pre-mRNA. The total molecular mass is 1.8M. U2, U5, and U6 snRNAs are colored marine, orange, and green, respectively. Pre-mRNA is colored red. Figure \(\PageIndex{16}\) shows just the structural changes in RNA and protein components that occur between the early Bact complex (left panel) and the mature Bact complex (right panel).
Ribonuclease P
This enzyme is a ribonucleoprotein that cleaves RNA through the catalytic action of one essential RNA subunit that displays ribozyme activity. It's found in most organisms. As with many ribozymes, the activity is increased 2-3 fold with bound proteins that stabilize the folded ribozyme and help bind the preferred substrate, which is pre-tRNA. Figure \(\PageIndex{17}\) shows bacterial RNase P ribozyme in complex with tRNA (3q1r).
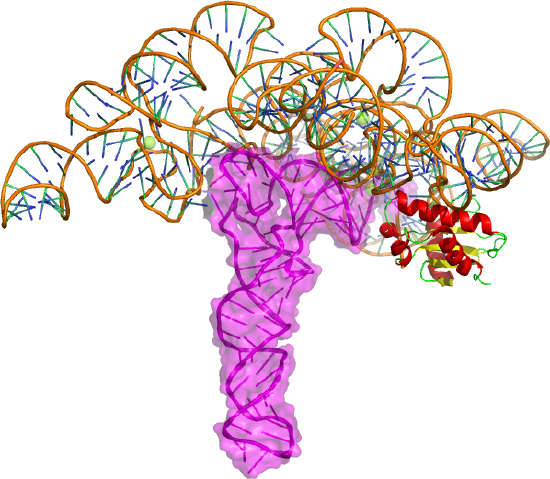
The tRNA is shown as a magenta cartoon/surface and the protein in cartoon form. The enzyme cleaves the 5' head end of the precursors of transfer RNA (tRNA) molecules. In bacteria, the enzyme is a heterodimer with one RNA and protein subunit.
Figure \(\PageIndex{18}\) shows a possible transition state with key residues involve in binding two Mg2+ ions in the active site. These ions are essential for catalysis as they stabilize the pentavalent intermediate/transition state.
Figure \(\PageIndex{19}\) shows an interactive iCn3D model of the active site with key residues labeled of bacterial RNase P ribozyme in complex with tRNA (3q1r).
Figure \(\PageIndex{19}\): Bacterial RNase P holoenzyme in complex with tRNA (3q1r) (Copyright; author via source).
Click the image for a popup or use this external link:https://structure.ncbi.nlm.nih.gov/i...6SjVMa8g6K47x5
RNA Polymerase Ribozyme
If primordial RNA acted as both a metabolic enzyme catalyst as well as the holder of the genetic information, it would need to have RNA polymerase activity. Artificial ribozymes with class I RNA ligase activity have been made. Figure \(\PageIndex{20}\) shows an interactive iCn3D model of the active site region of the Class I ligase ribozyme-substrate preligation complex, C47U mutant, Mg2+ bound ( 3R1L).
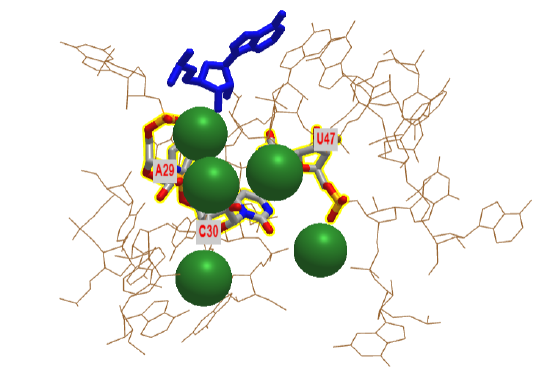
The blue stick represents just the 3' terminal adenosine end of the target substrate (5'UCCAGUA3') to which a new nucleoside would be added. The brown represents the active site region of the ribozyme. Three catalytic residues A29, C30 and C47 have been identified in the actual ribozyme. In iCn3D model is of a mutant, C47U, which has no catalytic activity. The green sphere represents Mg2+ ions.
Figure \(\PageIndex{21}\) show the active site residues and how they might facilitate stabilization of the pentavalent intermediate/transition state and the similarity of the active site to a protein RNA polymerase.
A divalent Mg2+ in the active site of the ribozyme enhances the nucleophilicity of the 3-OH on the primer, which attached the terminal phosphate of the G(1)TP substrate to form a pentavalent intermediate. The Mg cation is stabilized by oxygens on P 29 and 30 of the ribozyme. The Mg ion also stabilizes the developing charge in the transition state and in the charge in the intermediate. Stabilization of analogous divalent cations in the protein polymerase occurs through Asp side changes in the protein.
Ribosome
Protein synthesis from mRNA templates occurs on a ribosome, a nanomachine composed of proteins and ribosomal RNAs (rRNA). The ribosome is composed of two very large structural units. The smaller unit (termed 30S and 40S in bacteria and eukaryotes, respectively) coordinates the correct base pairing of the triplet codon on the mRNA with another small adapter RNA, transfer or tRNA, that brings a covalently connected amino acid to the site. Peptide bond formation occurs when another tRNA-amino acid molecule binds to an adjacent codon on mRNA. The tRNA has a cloverleaf tertiary structure with some intrastrand H-bonded secondary structure. The last three nucleotides at the 3' end of the tRNA are CpCpA. The amino acid is esterified to the terminal 3'OH of the terminal A by a protein enzyme, aminoacyl-tRNA synthetase.
Covalent amide bond formation between the second amino acid to the first, forming a dipeptide, occurs at the peptidyl transferase center, located on the larger ribosomal subunit (50S and 60S in bacteria and eukaryotes, respectively). The ribosome ratchets down the mRNA so the dipeptide-tRNA is now at the P or Peptide site, awaiting a new tRNA-amino acid at the A or Amino site. The figure below shows a schematic of the ribosome with bound mRNA on the 30S subunit and tRNAs covalently attached to amino acid (or the growing peptide) at the A and P site, respectively. Figure \(\PageIndex{22}\) shows a cartoon model of the prokaryotic ribosome with bound mRNA, tRNAs and the P and A sites.
We present another more detailed model of the ribosome complex illustrating protein synthesis in Figure \(\PageIndex{23}\).
A likely mechanism (derived from crystal structures with bound substrates and transition state analogs) for the formation of the amide bond between a growing peptide on the P-site tRNA and the amino acid on the A-site tRNA is shown below. Catalysis does not involve any of the ribosomal proteins (not shown) since none is close enough to the peptidyl transferase center to provide amino acids that could participate in general acid/base catalysis, for example. Hence the rRNA must act as the enzyme (i.e. it is a ribozyme). Initially, it was thought that a proximal adenosine with a perturbed pKa could, at physiological pH, be protonated/deprotonated and hence act as a general acid/base in the reaction. However, none was found. The most likely mechanism to stabilize the oxyanion transition state at the electrophilic carbon attack site is precisely located water, which is positioned at the oxyanion hole by H-bonds to uracil 2584 on the rRNA. The cleavage mechanism involves the concerted proton shuffle shown below. In this mechanism, the substrate (Peptide-tRNA) assists its own cleavage in that the 2'OH is in position to initiate the protein shuttle mechanism. (A similar mechanism might occur to facilitate hydrolysis of the fully elongated protein from the P-site tRNA.) Of course, all of this requires perfect positioning of the substrates and isn't that what enzymes do best? The main mechanisms for catalysis of peptide bond formation by the ribosome (as a ribozyme) are intramolecular catalysis and transition state stabilization by the appropriately positioned water molecule. These processes are illustrated in Figure \(\PageIndex{24}\).
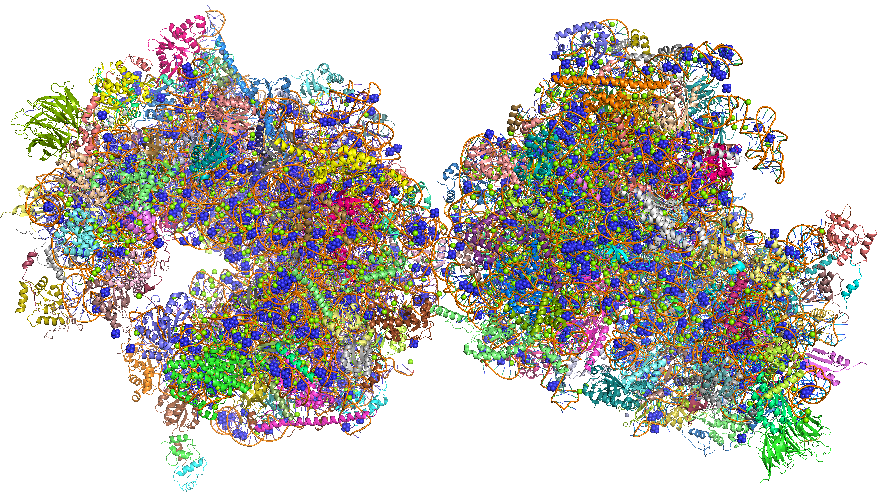
The crystal structure of the eukaryotic ribosome has recently been published (Ben-Shem et al). It is significantly larger (40%) with a mass of around 3x106 Daltons. The 40S subunit has one rRNA chain (18) and 33 associated proteins, while the larger 60S subunit has 3 rRNA chains (25S, 5.8S and 5S) and 46 associated proteins. The larger size of the eukaryotic ribosome facilitates more interactions with cellular proteins and greater regulation of cellular events. Figure \(\PageIndex{25}\) shows the two copies of the 80S yeast ribosome (4v88), presented to humble readers and authors alike.Figure \(\PageIndex{25}\): The 80S yeast ribosome (4v88). Each subunit is given a different color.
Ribozyme methyltransferase
The ribozymes described above and generally found in nature catalyze phosphoryl transfer reactions and with the ribosome, peptide bond formation. In vitro evolution can be used to drive new enzymatic functionalities, which would have been required in a RNA-only world that preceded the use of proteins as catalysts. RNA ribozymes are limited in having only 4 bases that can be employed in binding and catalytic steps, compared to 20 amino acids which can serve the same function in proteins. However, as in the case of protein, small molecule cofactors that bind to a potential ribozyme might facilitate greater catalytic efficiency and an expanded repertoire of reaction types. Indeed, we have seen above how small molecules can bind to riboswitches. Figure \(\PageIndex{26}\) shows the reaction and structure of a methyltransferase 1 ribozyme (MTR1) that acts as a methyltransferase. The small ligand, O6-methylguanine, binds to the ribozymes and acts as a cofactor in the methylation of adenine 63 in the RNA.
Figure \(\PageIndex{26}\): Deng, J., Wilson, T.J., Wang, J. et al. Structure and mechanism of a methyltransferase ribozyme. Nat Chem Biol 18, 556–564 (2022). https://doi.org/10.1038/s41589-022-00982-z. Creative Commons Attribution 4.0 International License. Creative Commons Attribution 4.0 International License, http://creativecommons.org/licenses/by/4.0/
Panel a shows the chemical reaction in which the methyl group of the small ligand O6-methylguanine is transferred to N1 of Adenine 63 in the RNA.
Panel b shows the sequence of the MTR1 ribozyme as crystallized for the experiments. The RNA is a three-way junction composed of three arms P1, P2 and P3. A GNRA tetraloop has been added to the end of the P3 helix so that the entire ribozyme comprises a single RNA strand. Subsections of the strands are named J12 (colored green), J23 (colored blue) and J31 (colored red).
X-ray crystal structures of the ribozyme in the presence of the cofactor O6-methylguanine were determined. The final structure contained guanine and an A63 methylated adenosine (1MA) implying the methyl group of the O6-methylguanine had transferred to A63, leaving guanine bound in the active site. The structure of the guanine bound to the ribozyme is shown in Figure \(\PageIndex{27}\).
Figure \(\PageIndex{27}\): The four planes of nucleobase interactions in the core of the ribozyme. The four planes are composed of (from bottom to top) G12•C38, C11•G41, exogenous guanine hydrogen bonded to C10, U45 and A63 and the triple interaction A9•A46•A40. Deng, J et al, ibid.
A hypothetical reaction mechanism is shown in Figure \(\PageIndex{28}\).
Figure \(\PageIndex{28}\): Proposed catalytic mechanism for the ribozyme methyltransferase. Deng, J et al, ibid.
In step 1, the nucleobase of C10 becomes protonated, and in step 2, the O6-methylguanine becomes bound. However, it is likely that these two steps are coordinated, as the binding will raise the pKa of the cytosine.
The methyl transfer reaction occurs in step 3 by the nucleophilic attack of A63 N1 on the methyl group of O6-methylguanine and the coordinated breakage of the guanine O6–C bond. This involves a train of electron transfers, the movement of the proton from C10 N3 to guanine N1 and a concomitant shift of the positive charge from C10 to the N1-methyladenine at position 63. In principle, the guanine can now be released as product, although there is no evidence that this occurs with the present form of the ribozyme. Regeneration of active ribozyme would also require an exchange of the substrate strand to place unmethylated adenine at position 63.
The proposed mechanism is fully consistent with the structure of the MTR1 riboswitch and the effect of the substitutions at C10 and U45 on activity. The complete loss of methylation activity of the C10U variant is fully consistent with the proposed role as a general acid in addition to ligand binding.
Figure \(\PageIndex{29}\) shows an interactive iCn3D model of a methyltransferase ribozyme (7V9E).
Figure \(\PageIndex{19}\): A methyltransferase ribozyme (7V9E). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...LtyVP4DBnkEgH6
The parts of the ribozyme are named J12 (light green), J23 (cyan) and J31 (magenta). The active site residues, C10, U45 and 1MA63 (1-methyladenosine) are shown in CPK-colored sticks and labeled (disregard small separated spheres). The Ba2+ ion in the crystal structure is not displayed. The guanine is shown in spacefill.




.png?revision=1&size=bestfit&width=412&height=227)
.png?revision=1&size=bestfit&width=322&height=367)
.png?revision=1&size=bestfit&width=347&height=308)
.png?revision=1&size=bestfit&width=418&height=443)