22.5: An Inducible Operon- Lac Operon
- Page ID
- 146168
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Next we will consider gene regulation in prokaryotic cells occurs through inducible operons. The lac operon is a typical inducible operon. While glucose is the preferred energy source, E. coli is able to use other sugars as energy sources when glucose concentrations are low, such as lactose. The lac operon encodes the genes necessary to acquire and break down lactose into glucose and galactose.
We will consider two aspects of regulation for the lac operons. The lac operon is positively regulated in response to glucose levels and negatively regulated in response to lactose levels.
An inducible operon: lac operon
The lac operon is an example of an inducible operon in response to lactose and is also subject to activation in the absence of glucose (Figure \(\PageIndex{1}\)). The lac operon encodes three structural genes necessary to acquire and process the disaccharide lactose from the environment, breaking it down into the simple sugars glucose and galactose. For the lac operon to be expressed, lactose must be present. This makes sense for the cell because it would be energetically wasteful to create the enzymes to process lactose if lactose was not available.
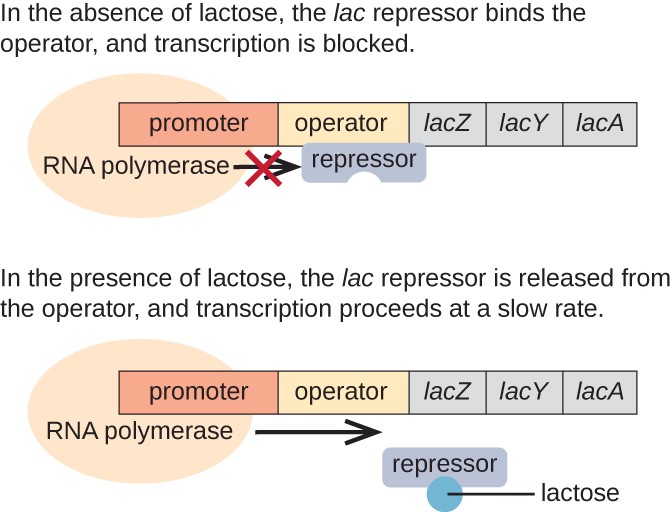
In the absence of lactose: the lac repressor is bound to the operator region of the lac operon, physically preventing RNA polymerase from transcribing the structural genes. Therefore, the lac operon is "OFF" by default.
When lactose is present: the lactose inside the cell is converted to allolactose. Allolactose serves as an inducer molecule, binding to the repressor and changing its shape so that it is no longer able to bind to the operator sequence. Removal of the repressor in the presence of lactose allows RNA polymerase to move through the operator region and begin transcription of the lac structural genes. Thus, the lac operon is "ON" in the presence of lactose.
The lac Operon: Activation by Catabolite Activator Protein
The ability to switch from glucose use to another substrate like lactose is a consequence of the activity of an enzyme called Enzyme IIA (EIIA). When glucose levels drop, cells produce less ATP from catabolism , and EIIA becomes phosphorylated. Phosphorylated EIIA activates adenylyl cyclase, an enzyme that converts some of the remaining ATP to cyclic AMP (cAMP), a cyclic derivative of AMP and an important signaling molecule involved in glucose and energy metabolism in E. coli. As a result, cAMP levels begin to rise in the cell (Figure \(\PageIndex{2}\)). You can think of cAMP as a signal that the cell is hungry and needs an alternate food source.

The lac operon also plays a role in this switch from using glucose to using lactose. When glucose is scarce, the accumulating cAMP caused by increased adenylyl cyclase activity binds to catabolite activator protein (CAP), also known as cAMP receptor protein (CRP). The complex binds to the promoter region of the lac operon (Figure 6). In the regulatory regions of these operons, a CAP binding site is located upstream of the RNA polymerase binding site in the promoter. Binding of the CAP-cAMP complex to this site increases the binding ability of RNA polymerase to the promoter region to initiate the transcription of the structural genes. Thus, in the case of the lac operon, for transcription to occur, lactose must be present (removing the lac repressor protein) and glucose levels must be depleted (allowing binding of an activating protein). When glucose levels are high, there is catabolite repression of operons encoding enzymes for the metabolism of alternative substrates. Because of low cAMP levels under these conditions, there is an insufficient amount of the CAP-cAMP complex to activate transcription of these operons.
Figure \(\PageIndex{3}\): Transcriptional activation by the CAP protein. When glucose levels fall, E. coli may use other sugars for fuel but must transcribe new genes to do so. As glucose supplies become limited, cAMP levels increase. This cAMP binds to the CAP protein, a positive regulator that binds to a promoter region upstream of the genes required to use other sugar sources.
Putting it all together
Watch the video below about the lac operon and the test your knowledge by filling out the table below.
Check out this lac operon interactive. To use the interactive, click "start interactive" and then slowing slow through the content, revealing the information and animations.
Considering lactose and glucose
Make a copy of the table below in your notes and fill in the empty columns. In response to glucose, indicate whether CAP will bind to the promotor or not. In response to lactose, indicate whether the repressor will bind to the operator. For the transcription column, consider your responses to both conditions and determine whether there will be no transcription, some transcription or high transcription of the lac operon. When you have finished, click answer to reveal the solution.
| Table 1. Conditions Affecting Transcription of the lac Operon | ||||
|---|---|---|---|---|
| Glucose | CAP binds | Lactose | Repressor binds | Transcription |
| + | - | |||
| + | + | |||
| - | - | |||
| - | + | |||
- Answer
-
Table 1. Conditions Affecting Transcription of the lac Operon Answer Key Glucose CAP binds Lactose Repressor binds Transcription + – – + None + – + – Some – + – + None – + + – High
Additional Attributions
- OpenStax Microbiology. Provided by: OpenStax CNX. License: CC BY: Attribution. License Terms: Download for free at OpenStax Microbiology.

