19.3: Prokaryotic Transcription
- Page ID
- 139560
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Both prokaryotes and eukaryotes perform fundamentally the same process of transcription, with the important difference of the membrane-bound nucleus in eukaryotes. With the genes bound in the nucleus, transcription occurs in the nucleus of the cell and the mRNA transcript must be transported to the cytoplasm. In prokaryotes, which lack membrane-bound nuclei and other organelles, transcription occurs in the cytoplasm of the cell.
RNA Polymerase
RNA Polymerase is the enzyme that produces the mRNA molecule (just like DNA polymerase produced a new DNA molecule during DNA replication). Prokaryotes use the same RNA polymerase to transcribe all of their genes. In E. coli, the polymerase is composed of five polypeptide subunits. These subunits assemble every time a gene is transcribed, and they disassemble once transcription is complete. Each subunit has a unique role (which you do not need to memorize). The polymerase comprised of all five subunits is called the holoenzyme.
Initiation
Transcription in prokaryotes (and in eukaryotes) requires the DNA double helix to partially unwind in the region of mRNA synthesis. The region of unwinding is called a transcription bubble. The DNA sequence onto which the proteins and enzymes involved in transcription bind to initiate the process is called a promoter. In most cases, promoters exist upstream of the genes they regulate. The specific sequence of a promoter is very important because it determines whether the corresponding gene is transcribed all of the time, some of the time, or hardly at all. The structure and function of a prokaryotic promoter is relatively simple (Figure \(\PageIndex{1}\)). One important sequence in the prokaryotic promoter is located 10 bases before the transcription start site (-10) and is commonly called the TATA box.
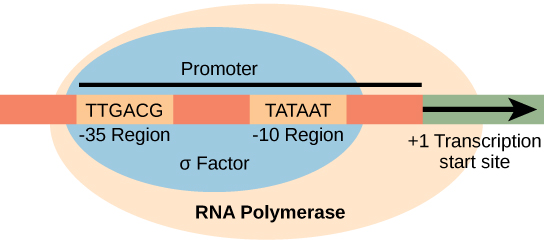
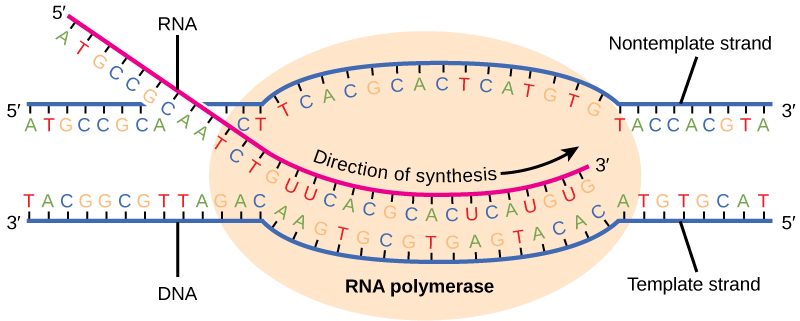
To begin transcription, the RNA polymerase holoenzyme assembles at the promoter. The dissociation of σ allows the core enzyme to proceed along the DNA template, synthesizing mRNA by adding RNA nucleotides according to the base pairing rules, similar to the way a new DNA molecule is produced during DNA replication. Only one of the two DNA strands is transcribed. The transcribed strand of DNA is called the template strand because it is the template for mRNA production. The mRNA product is complementary to the template strand and is almost identical to the other DNA strand, called the non-template strand, with the exception that RNA contains a uracil (U) in place of the thymine (T) found in DNA. Like DNA polymerase, RNA polymerase adds new nucleotides onto the 3′-OH group of the previous nucleotide. This means that the growing mRNA strand is being synthesized in the 5′ to 3′ direction. Because DNA is anti-parallel, this means that the RNA polymerase is moving in the 3′ to 5′ direction down the template strand (Figure \(\PageIndex{2}\)).
Elongation
As elongation proceeds, the DNA is continuously unwound ahead of the core enzyme as the hydrogen bonds that connect the complementary base pairs in the DNA double helix are broken (Figure \(\PageIndex{2}\)). The DNA is rewound behind the core enzyme as the hydrogen bonds are reformed. The base pairing between DNA and RNA is not stable enough to maintain the stability of the mRNA synthesis components. Instead, the RNA polymerase acts as a stable linker between the DNA template and the newly forming RNA strand to ensure that elongation is not interrupted prematurely.
Termination
Once a gene is transcribed, the RNA polymerase needs to be instructed to dissociate from the DNA template and liberate the newly made mRNA. Depending on the gene being transcribed, there are two kinds of termination signals. One is protein-based and the other is RNA-based. Both termination signals rely on specific sequences of DNA near the end of the gene that cause the polymerase to release the mRNA.
Putting it all together
Watch this Transcription Animation and then answer the questions below.
Polyribosome
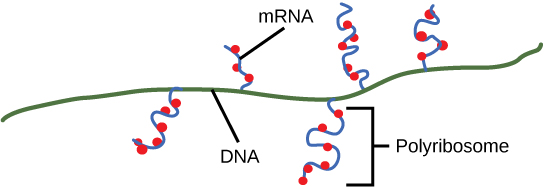
In a prokaryotic cell, by the time transcription ends, the transcript would already have been used to begin making copies of the encoded protein because the processes of transcription and translation can occur at the same time since both occur in the cytoplasm (Figure \(\PageIndex{3}\)). In contrast, transcription and translation cannot occur simultaneously in eukaryotic cells since transcription occurs inside the nucleus and translation occurs outside in the cytoplasm.
References
Unless otherwise noted, images on this page are licensed under CC-BY 4.0 by OpenStax.
OpenStax, Biology. OpenStax CNX. December 21, 2017 https://cnx.org/contents/GFy_h8cu@10...-Transcription

