10.4: Classification of Viruses
- Page ID
- 3235
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- State what criteria are used in viral classification.
- Regarding the naming of enzymes involved in the replication of viral nucleic acid, state what the "dependent" part of the name refers to and what the "polymerase" part of the name refers to.
Viruses can store their genetic information in six different types of nucleic acid which are named based on how that nucleic acid eventually becomes transcribed to the viral mRNA (Figure \(\PageIndex{1}\)) capable of binding to host cell ribosomes and being translated into viral proteins.
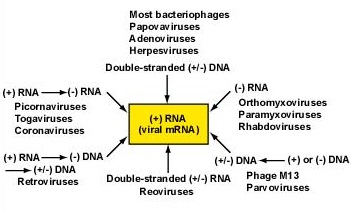
In the diagrams below, (+) and (-) represent complementary strands of nucleic acid. Copying of a (+) strand by complementary base pairing forms a (-) strand. Only a (+) viral mRNA strand can be translated into viral protein. Regarding the enzymes involved in nucleic acid replication, the "dependent" part of the name refers what type of nucleic acid is being copied. The "polymerase" part of the name refers what type of nucleic acid is being synthesized, e.g., DNA-dependent RNA-polymerase would synthesize a strand of RNA complementary to a strand of DNA. These six forms of viral nucleic acid are:
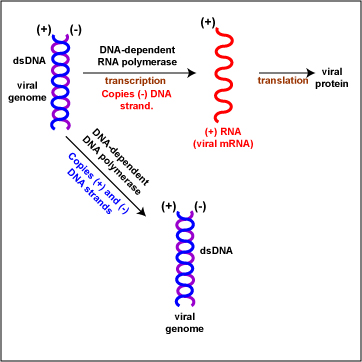
- (+/-) double-stranded DNA (Figure \(\PageIndex{2}\)). To replicate the viral genome, DNA-dependent DNA polymerase enzymes copy both the (+) and (-) DNA strands producing dsDNA viral genomes. To produce viral mRNA molecules, DNA-dependent RNA polymerase enzymes copy the (-) DNA strand into (+) viral mRNA. The (+) viral mRNA can then be translated into viral proteins by host cell ribosomes. Examples include most bacteriophages, Papovaviruses, Adenoviruses, and Herpesviruses.
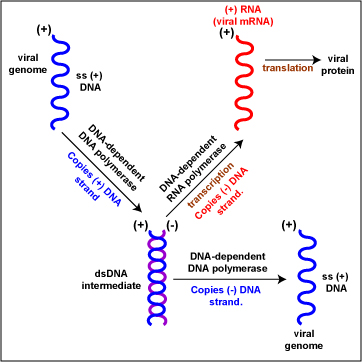
- (+) single-stranded DNA (Figure \(\PageIndex{2}\)). To replicate the viral genome, DNA-dependent DNA polymerase enzymes copy the (+) DNA strand of the genome producing a dsDNA intermediate. DNA-dependent DNA polymerase enzymes then copy the (-) DNA strand into ss (+) DNA genomes. To produce viral mRNA molecules, DNA-dependent RNA polymerase enzymes copy the (-) DNA strand into (+) viral mRNA. The (+) viral mRNA can then be translated into viral proteins by host cell ribosomes. Examples include Phage M13 and Parvoviruses.
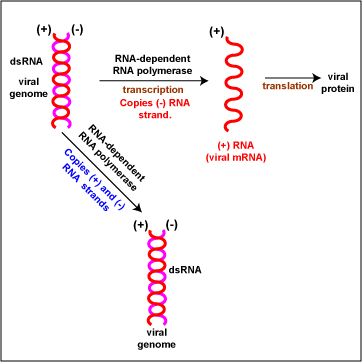
- (+/-) double-stranded RNA (Figure \(\PageIndex{4}\)). To replicate the viral genome, RNA-dependent RNA polymerase enzymes copy both the (+) RNA and (-) RNA strands of the genome producing a dsRNA genomes. To produce viral mRNA molecules, RNA-dependent RNA polymerase enzymes copy the (-) RNA strand into (+) viral mRNA. The (+) viral mRNA can then be translated into viral proteins by host cell ribosomes. Reoviruses are an example.
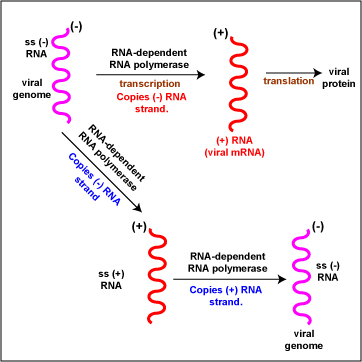
- (-) RNA (Figure \(\PageIndex{5}\)). To replicate the viral genome, RNA-dependent RNA polymerase enzymes copy the (-) RNA genome producing ss (+) RNA. RNA-dependent RNA polymerase enzymes then copy the (+) RNA strands producing ss (-) RNA viral genome. The (+) mRNA strands also function as viral mRNA and can then be translated into viral proteins by host cell ribosomes. Examples include Orthomyxoviruses, Paramyxoviruses, Rhabdoviruses.
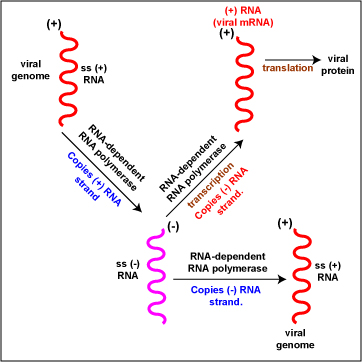
- (+) RNA (Figure \(\PageIndex{6}\)). To replicate the viral genome, RNA-dependent RNA polymerase enzymes copy the (+) RNA genome producing ss (-) RNA. RNA-dependent RNA polymerase enzymes then copy the (-) RNA strands producing ss (+) RNA viral genome. To produce viral mRNA molecules. RNA-dependent RNA polymerase enzymes copy the (-) RNA strand into (+) viral mRNA. The (+) viral mRNA can then be translated into viral proteins by host cell ribosomes. Examples include Picornaviruses, Togaviruses, and Coronaviruses.
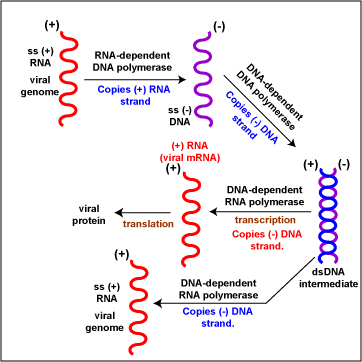
- (+) RNA Retroviruses (Figure \(\PageIndex{7}\)). To replicate the viral genome, reverse transcriptase enzymes (RNA-dependent DNA polymerases) copy the (+) RNA genome producing ss (-) DNA strands. DNA-dependent DNA polymerase enzymes then copy the (-) DNA strands to produce a dsDNA intermediate. DNA-dependent RNA polymerase enzymes then copy the (-) DNA strands to produce ss (+) RNA genomes. To produce viral mRNA molecules, DNA-dependent RNA polymerase enzymes copy the (-) DNA strand into (+) viral mRNA. The (+) viral mRNA can then be translated into viral proteins by host cell ribosomes. Retroviruses, such as HIV-1, HIV-2, and HTLV-1 are examples.
A viral enzyme that synthesizes a complementary RNA copy of an RNA would be called what?
Table \(\PageIndex{1}\) below describes some of the medically important viruses.
| Properties | Viral Family | Size | Example |
|---|---|---|---|
| single-stranded DNA; naked; polyhedral capsid | Parvoviridae | 18-25 nm | parvoviruses (roseola, fetal death, gastroenteritis; some depend on coinfection with adenoviruses) |
| double-stranded, DNA; naked; polyhedral capsid | Papovaviridae; circular dsDNA | 40-57 nm | human papilloma viruses (HPV; benign warts and genital warts; genital and rectal cancers) |
| Adenoviridae; dsDNA | 70-90 nm | adenoviruses (respiratory infections, gastroenteritis, infectious pinkeye, rashes, meningoencephalitis) | |
| double-stranded, circular DNA; enveloped; complex | Poxviridae | 200-350 nm | smallpox virus (smallpox), vaccinia virus (cowpox), molluscipox virus (molluscum contagiosum-wartlike skin lesions) |
| double-stranded DNA; enveloped; polyhedral capsid | Herpesviridae | 150-200 nm | herpes simplex 1 virus (HSV-1; most oral herpes; herpes simplex 2 virus (HSV-2; most genital herpes), herpes simplex 6 virus (HSV-6; roseola), varicella-zoster virus (VZV; chickenpox and shingles), Epstein-Barr virus (EBV; infectious mononucleosis and lymphomas), cytomegalovirus (CMV; birth defects and infections of a variety of body systems in immunosuppressed individuals) |
| Hepadnaviridae | 42 nm | hepatitis B virus (HBV; hepatitis B and liver cancer) | |
| (+)single-stranded RNA; naked; polyhedral capsid | picornaviridae | 28-30 nm | enteroviruses (poliomyelitis), rhinoviruses (most frequent cause of the common cold), Noroviruses (gastroenteritis), echoviruses (meningitis), hepatitis A virus (HAV; hepatitis A) |
| (+)single-stranded RNA; enveloped; usually a polyhedral capsid | Togaviridae | 60-70 nm | arboviruses (eastern equine encephalitis, western equine encephalitis), rubella virus (German measles) |
| Flaviviridae | 40-50 nm | flaviviruses (yellow fever, dengue fever, St. Louis encephalitis), hepatitis C virus (HCV; hepatitis C) | |
| Coronaviridae | 80-160 nm | coronaviruses (upper respiratory infections and the common cold; SARS) | |
| (-)single-stranded RNA; enveloped; pleomorphic | Rhabdoviridae; bullet-shaped | 70-189 nm | rabies virus (rabies) |
| Filoviridae; long and filamentous | 80-14,000 nm | Ebola virus, Marburg virus (hemorrhagic fevers) | |
| Paramyxoviridae; pleomorphic | 150-300 nm | paramyxoviruses (parainfluenza, mumps); measles virus (measles) | |
| (-) strand; multiple strands of RNA; enveloped | Orthomyxoviridae | 80-200 nm | influenza viruses A, B, and C (influenza) |
| Bunyaviridae | 90-120 nm | California encephalitis virus (encephalitis); hantaviruses (Hantavirus pulmonary syndrome, Korean hemorrhagic fever) | |
| Arenaviridae | 50-300 nm | arenaviruses (lymphocytic choriomeningitis, hemorrhagic fevers) | |
| produce DNA from (+) single-stranded RNA using reverse transcriptase; enveloped; bullet-shaped or polyhedral capsid | Retroviridae | 100-120 nm | HIV-1 and HIV-2 (HIV infection/AIDS); HTLV-1 and HTLV-2 (T-cell leukemia) |
| dsRNA; naked; polyhedral capsid | Reoviridae | 60-80 nm | reoviruses (mild respiratory infections, infant gastroenteritis); Colorado tick fever virus (Colorado tick fever) |
Summary
- Viruses can store their genetic information in six different types of nucleic acid which are named based on how that nucleic acid eventually becomes transcribed to the viral mRNA.
- (+) and (-) strands of nucleic acid are complementary. Copying a (+) stand gives a (-) strand; copying a (-) stand gives a (+) strand.
- Only (+) strands of viral RNA can be translated into viral protein.
- Regarding the enzymes involved in nucleic acid replication, the "dependent" part of the name refers what type of nucleic acid is being copied. The "polymerase" part of the name refers what type of nucleic acid is being synthesized.
Questions
Study the material in this section and then write out the answers to these questions. Do not just click on the answers and write them out. This will not test your understanding of this tutorial.
- State what criteria are used in viral classification. (ans)
- What would a DNA-dependent RNA-polymerase enzyme do? (ans)


