5.1: Introduction to Bioinformatics
- Page ID
- 135672
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Bioinformatics and "personalized medicine" has become a model of healthcare that is tailor-made to each person’s unique genetic make-up. Because of its ability to analyze vast amounts of data obtained from procedures, like genome sequencing or microarray analysis, bioinformatics allows scientists to search for mutations or gene variants that could affect a patient’s response to a particular drug or modify the disease prognosis. By knowing a patient’s genetic profile doctors may be able to predict their patient's susceptibility to certain diseases, provide them proper medication and with the proper dose to reduce side-effects.
Introduction
Bioinformatics emerged in the late 20th century as an interdisciplinary field that combines biology, computer science, mathematics, and statistics to analyze and interpret biological data (Figure \(\PageIndex{1}\)). With advancements in genomics, proteomics, and other data-rich technologies, managing the vast amounts of data generated by these fields has become key. In response, bioinformatics develops methods and software to understand this data. Experts working in this field write algorithms, create databases, develop statistical techniques, and utilize powerful tools for analyzing sequences, structures, and functions of DNA, RNA, and proteins. Simply put, researchers working with massive datasets that require computational tools to manage and interpret data turn to bioinformatics. It is through bioinformatics that we can begin to understand complex biological systems and drive discoveries in areas like genetics, molecular biology, medicine, and evolutionary biology.
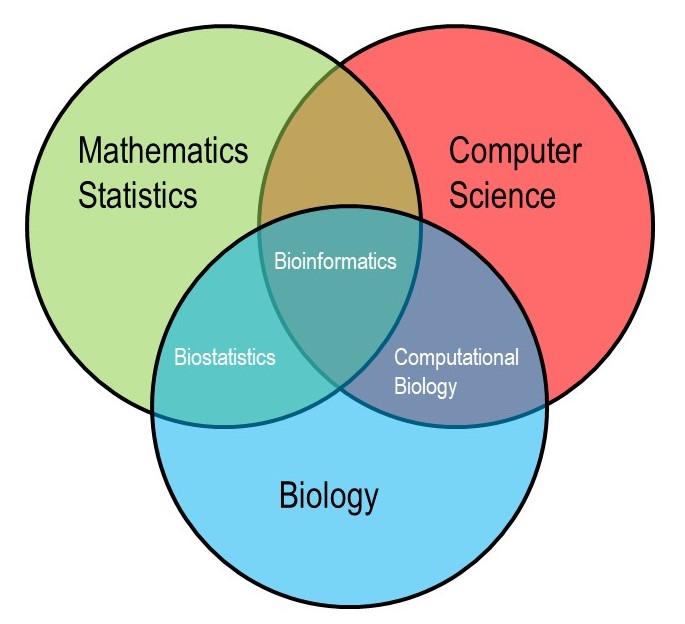
The field of bioinformatics combines biology, mathematics, statistics, and computer science. At the end of this section you will be able to:
- Define and explain bioinformatics
- Explain some of the applications of bioinformatics
- List some tools used in bioinformatics
Applications of Bioinformatics
The primary goal of bioinformatics is to increase our understanding of biological processes through the analysis of raw biological data. While this is not unique, what sets bioinformatics apart from other similar fields is its development and use of computationally-based approaches to achieve this goal. Major research efforts in the field of bioinformatics include sequence alignment, genome assembly and analysis, drug discovery and design, protein structure prediction, gene expression prediction, protein-protein interactions, systems biology, and evolution modeling.
Genomics and Personalized Medicine
Genomics is the study of an organism's entire genome - the complete set of DNA found within a cell. Bioinformatics plays a pivotal role in genomics, especially in the sequencing and analysis of genomes. With the advent of advanced DNA sequencing technologies, like next-generation sequencing (NGS), entire genomes can be sequenced in a day. Because of this, bioinformatics tools are needed in order to assemble the resulting DNA sequences into genomes, in addition to annotating genes, and identifying genetic variations. This genomic information is crucial for understanding genetic diseases and developing personalized medicine approaches. Bioinformatic data can reveal how a patient will respond to certain drugs, such as chemotherapies, enabling clinicians to personalize a patient's treatment regime and prescribe the most effective treatment. Moreover, bioinformatics is integral to genome-wide association studies (GWAS), which identify genetic variants linked to complex diseases like cancer, diabetes, and heart disease. By analyzing large datasets from thousands of individuals, researchers can discover new genetic markers and understand the biological pathways involved in these diseases.
Proteomics and Drug Discovery
Proteomics is the large-scale study of the proteome, the entire set of proteins expressed by a genome (Figure \(\PageIndex{2}\)). Proteomics allows researchers to study the structure and function of proteins, along with their interactions that are central to nearly all biological processes. Unlike the static nature of the genome, the proteome is dynamic and changes in response to different conditions. Bioinformatics is a key tool used in analyzing the proteome. Techniques like mass spectrometry generate vast amounts of protein data, which bioinformatics interprets. Through its study of protein interactions, pathways, and functions, bioinformatics allows us to better study disease mechanisms and identify potential targets for new or existing drugs. Through its analysis of 3D structure, bioinformatics, combined with computational docking simulations, allows for the design of small molecules that can better interact with specific protein targets, leading to the development of more efficacious drugs. Bioinformatics also aids in virtual screening, where large chemical libraries are computationally tested to identify compounds that are likely to bind to a target protein.

Evolutionary Biology and Comparative Genomics
Bioinformatics has revolutionized evolutionary biology by allowing researchers to compare the genomes of different species. Through tools like multiple sequence alignments and phylogenetic tree-building software, scientists can trace the evolutionary history of genes and species, identifying conserved elements and evolutionary innovations. Comparative genomics, which compares genomes across species, helps scientists discover insights into evolutionary relationships, identify conserved genes that have remained unchanged across species, and explore genetic adaptations that have enabled organisms to survive in changing environments. This comparative approach can also provide insights into human evolution and health through its study of genetic similarities between humans and closely related species, such as chimpanzees and other primates. Comparative genomics has been invaluable for evolutionary biology by offering a deeper understanding of how genetic diversity contributes to the complexity of life.
Systems Biology and Big Data
Bioinformatics is central to systems biology, which tries to understand biological systems as whole networks rather than isolated components. In systems biology, bioinformatics tools are used to integrate data from various sources, such as genomics, transcriptomics, proteomics, and metabolomics, in order to build models of cellular processes. These models are critical to helping researchers understand how the components of a biological system interact with one another and lead to insights into disease mechanisms, drug responses, and metabolic pathways. With the rise of "big data" in biology, bioinformatics has become essential for managing, processing, and analyzing the vast datasets that forms the core of systems biology. Data integration, machine learning, and artificial intelligence (AI) are becoming more and more common in systems biology in order to uncover patterns in these datasets. This makes system biology and bioinformatics powerful tools for testing theories about health and disease and may lead to novel discoveries in fields such as cancer research, neurobiology, and microbiome studies.
Tools of Bioinformatics
At its core, bioinformatics relies on biological databases and computational tools to analyze and interpret data. Common activities in bioinformatics include mapping and analyzing DNA and protein sequences, aligning DNA and protein sequences to compare them, and creating and viewing 3-D models of protein structures. Sequence alignment algorithms are used to compare genetic sequences to identify homologous regions. Phylogenetic tree-building algorithms are used to infer evolutionary relationships among species. Machine learning techniques, such as neural networks and support vector machines, are increasingly being applied to predict gene functions, classify proteins, and identify disease-associated genetic variants. Databases like GenBank, EMBL, and UniProt store genetic, protein, and other biological information, allowing researchers to retrieve, compare, and analyze sequences from various organisms. Tools like BLAST (Basic Local Alignment Search Tool) are used for sequence alignment, identifying similarities between genetic or protein sequences, and inferring evolutionary relationships. Finally, bioinformatics tools like PyMOL and Rosetta are used to also model and predict the 3D structures of proteins and nucleic acids.
Animation: What is Bioinformatics?
Bioinformatics is the scientific field that uses powerful computer programs to interpret large amounts of complex biological data. Through bioinformatics, both scientists and clinicians can now utilize massive and complex databases in the hopes of making advances in health and human welfare.
Some important concepts to remember:
- bioinformatics involves the collection, storage, and analysis of biological data generated from genomic and proteomic analysis
- bioinformatics develops computer programs to manage and use this biological data
- bioinformatics creates algorithms and statistical approaches to find patterns and relationships within large databases, thus allowing us to understand complex biological systems as whole
Glossary
Algorithm - a step-by-step computational procedure used to analyze data
Bioinformatics - an interdisciplinary field that combines biology, computer science, mathematics, and statistics to analyze and interpret biological data
Comparative genomics - a field in genomics that compares genomes across species
Database - a structured collection of data
Genome - the totality of genetic information found within the cell
Genome-Wide Association Studies (GWAS) - research methods used to identify genetic variants across the entire genome that are associated with specific traits or diseases
Genomics - the study of the genome
Machine learning techniques - algorithms and methods that allow computers to learn patterns from data and make predictions or decisions without being explicitly programmed for a specific task
Mass spectrometry - an analytical technique used to identify, characterize, and quantify molecules by measuring their mass-to-charge ratio (m/z)
Next-generation sequencing (NGS) - a sequencing technology that enables the sequencing of an entire genome in several hours
Phylogenetic tree building - the process of creating a phylogenetic tree
Phylogenetic tree - a diagram that shows the evolutionary relationships among a group of organisms, genes, or proteins based on their genetic or structural similarities and differences
Proteome - the entire set of proteins expressed by a genome
Proteomics - the study of the proteome
Sequence alignment - the process of arranging two or more DNA, RNA, or protein sequences to identify regions of similarity or difference
Systems biology - a field of biology that seeks understand biological systems as a whole network rather than isolated components

