5.2: Structure and Replication of DNA
- Page ID
- 75835
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the biochemical structure of deoxyribonucleotides
- Identify the base pairs used in the synthesis of deoxyribonucleotides
- Explain why the double helix of DNA is described as antiparallel
- Explain the meaning of semiconservative DNA replication
- Explain why DNA replication is bidirectional and includes both a leading and lagging strand
- Explain why Okazaki fragments are formed
- Describe the process of DNA replication and the functions of the enzymes involved
- Identify the differences between DNA replication in bacteria and eukaryotes
- Explain the process of rolling circle replication
Like other macromolecules, nucleic acids are composed of monomers, called nucleotides, which are polymerized to form large strands. Each nucleic acid strand contains certain nucleotides that appear in a certain order within the strand, called its base sequence. The base sequence of deoxyribonucleic acid (DNA) is responsible for carrying and retaining the hereditary information in a cell. Now, we will discuss in detail the ways in which DNA uses its own base sequence to direct its own synthesis, as well as the synthesis of RNA and proteins, which, in turn, gives rise to products with diverse structure and function. In this section, we will discuss the basic structure and function of DNA.
DNA Nucleotides
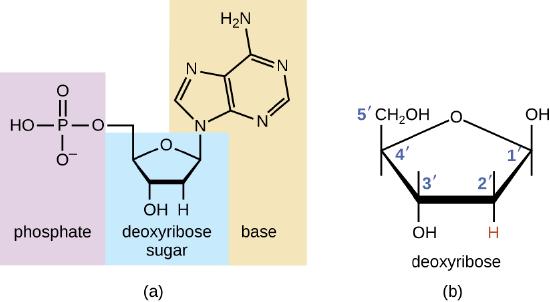
The building blocks of nucleic acids are nucleotides. Nucleotides that compose DNA are called deoxyribonucleotides. The three components of a deoxyribonucleotide are a five-carbon sugar called deoxyribose, a phosphate group, and a nitrogenous base, a nitrogen-containing ring structure that is responsible for complementary base pairing between nucleic acid strands (Figure \(\PageIndex{1}\)). The carbon atoms of the five-carbon deoxyribose are numbered 1ʹ, 2ʹ, 3ʹ, 4ʹ, and 5ʹ (1ʹ is read as “one prime”).
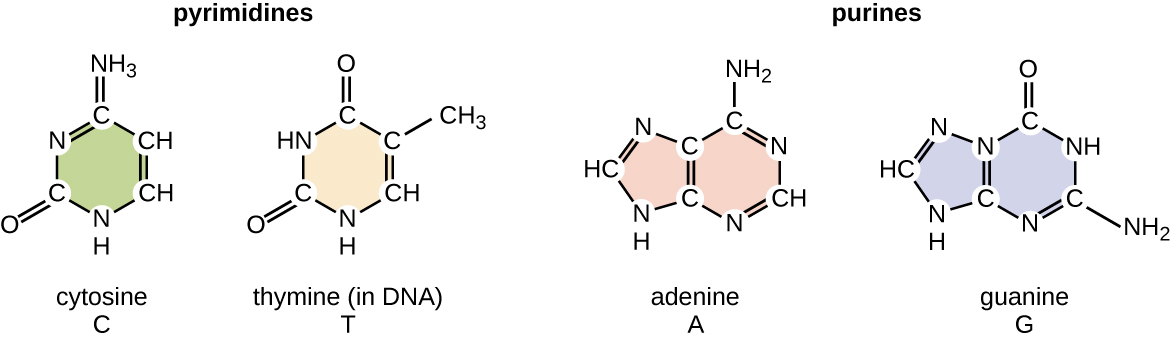
The deoxyribonucleotide is named according to the nitrogenous bases (Figure \(\PageIndex{2}\)). The nitrogenous bases adenine (A) and guanine (G) are the purines; they have a double-ring structure with a six-carbon ring fused to a five-carbon ring. The pyrimidines, cytosine (C) and thymine (T), are smaller nitrogenous bases that have only a six-carbon ring structure.
Individual nucleotides with three phosphates combine with each other by covalent bonds known as 5ʹ-3ʹ phosphodiester bonds, or linkages whereby the phosphate group attached to the 5ʹ carbon of the sugar of one nucleotide bonds to the hydroxyl group of the 3ʹ carbon of the sugar of the next nucleotide. Phosphodiester bonding between nucleotides forms the sugar-phosphate backbone, the alternating sugar-phosphate structure composing the framework of a nucleic acid strand (Figure \(\PageIndex{3}\)). During the polymerization process, deoxynucleotide triphosphates (dNTP) are used. To construct the sugar-phosphate backbone, the two terminal phosphates are released from the dNTP as a pyrophosphate. The resulting strand of nucleic acid has a free phosphate group at the 5ʹ carbon end and a free hydroxyl group at the 3ʹ carbon end. The two unused phosphate groups from the nucleotide triphosphate are released as pyrophosphate during phosphodiester bond formation. Pyrophosphate is subsequently hydrolyzed, releasing the energy used to drive nucleotide polymerization.
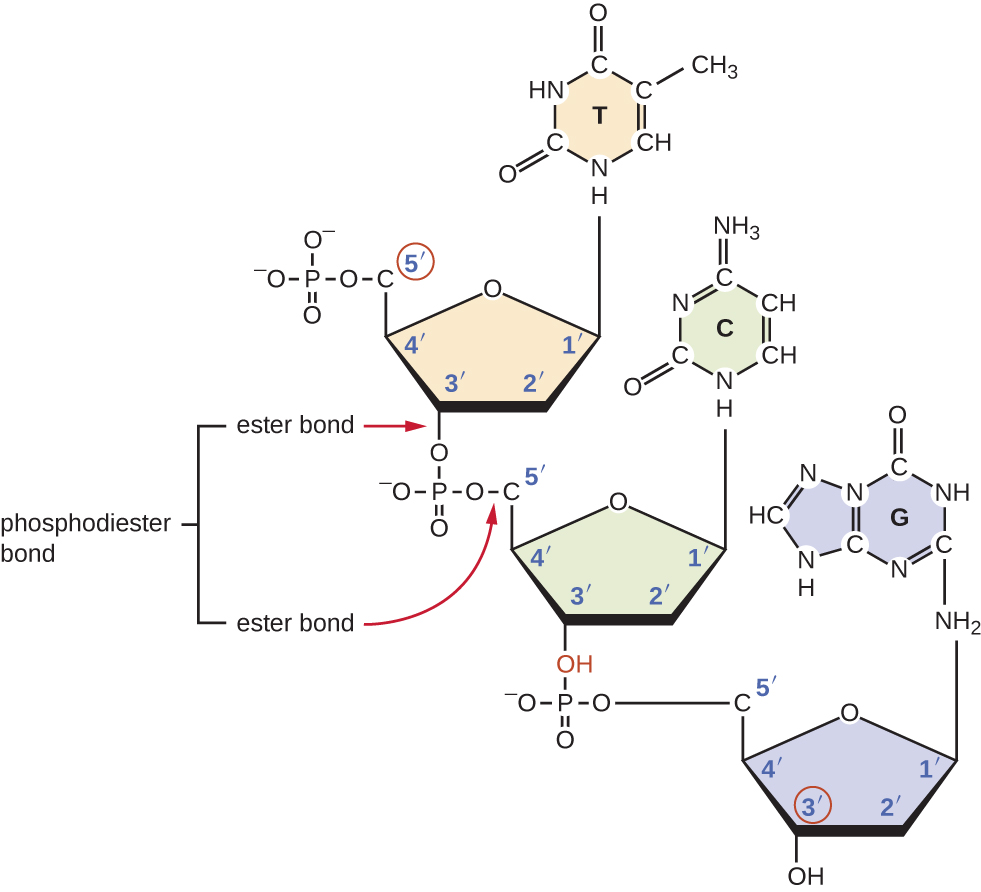
What is meant by the 5ʹ and 3ʹ ends of a nucleic acid strand?
Discovering the Double Helix
By the early 1950s, considerable evidence had accumulated indicating that DNA was the genetic material of cells, and now the race was on to discover its three-dimensional structure. Around this time, Austrian biochemist Erwin Chargaff1(1905–2002) examined the content of DNA in different species and discovered that adenine, thymine, guanine, and cytosine were not found in equal quantities, and that it varied from species to species, but not between individuals of the same species. He found that the amount of adenine was very close to equaling the amount of thymine, and the amount of cytosine was very close to equaling the amount of guanine, or A = T and G = C. These relationships are also known as Chargaff’s rules.
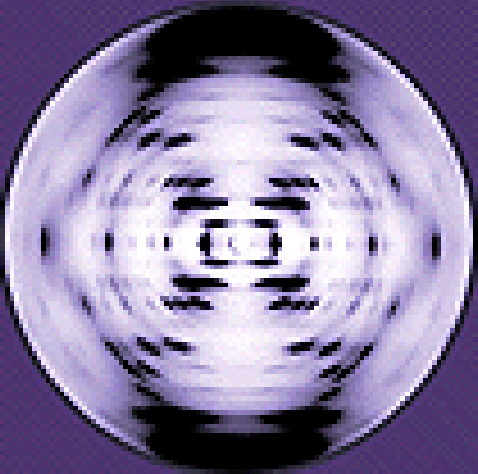
Other scientists were also actively exploring this field during the mid-20th century. In 1952, American scientist Linus Pauling (1901–1994) was the world’s leading structural chemist and odds-on favorite to solve the structure of DNA. Pauling had earlier discovered the structure of protein α helices, using X-ray diffraction, and, based upon X-ray diffraction images of DNA made in his laboratory, he proposed a triple-stranded model of DNA.2 At the same time, British researchers Rosalind Franklin (1920–1958) and her graduate student R.G. Gosling were also using X-ray diffraction to understand the structure of DNA (Figure \(\PageIndex{4}\)). It was Franklin’s scientific expertise that resulted in the production of more well-defined X-ray diffraction images of DNA that would clearly show the overall double-helix structure of DNA.

James Watson (1928–), an American scientist, and Francis Crick (1916–2004), a British scientist, were working together in the 1950s to discover DNA’s structure. They used Chargaff’s rules and Franklin and Wilkins’ X-ray diffraction images of DNA fibers to piece together the purine-pyrimidine pairing of the double helical DNA molecule (Figure \(\PageIndex{5}\)). In April 1953, Watson and Crick published their model of the DNA double helix in Nature.3 The same issue additionally included papers by Wilkins and colleagues,4 as well as by Franklin and Gosling,5 each describing different aspects of the molecular structure of DNA. In 1962, James Watson, Francis Crick, and Maurice Wilkins were awarded the Nobel Prize in Physiology and Medicine. Unfortunately, by then Franklin had died, and Nobel prizes at the time were not awarded posthumously. Work continued, however, on learning about the structure of DNA. In 1973, Alexander Rich (1924–2015) and colleagues were able to analyze DNA crystals to confirm and further elucidate DNA structure.6

Which scientists are given most of the credit for describing the molecular structure of DNA?
DNA Structure
Watson and Crick proposed that DNA is made up of two strands that are twisted around each other to form a right-handed helix. The two DNA strands are antiparallel, such that the 3ʹ end of one strand faces the 5ʹ end of the other (Figure \(\PageIndex{6}\)). The 3ʹ end of each strand has a free hydroxyl group, while the 5ʹ end of each strand has a free phosphate group. The sugar and phosphate of the polymerized nucleotides form the backbone of the structure, whereas the nitrogenous bases are stacked inside. These nitrogenous bases on the interior of the molecule interact with each other, base pairing.
Analysis of the diffraction patterns of DNA has determined that there are approximately 10 bases per turn in DNA. The asymmetrical spacing of the sugar-phosphate backbones generates major grooves (where the backbone is far apart) and minor grooves (where the backbone is close together) (Figure \(\PageIndex{6}\)). These grooves are locations where proteins can bind to DNA. The binding of these proteins can alter the structure of DNA, regulate replication, or regulate transcription of DNA into RNA.
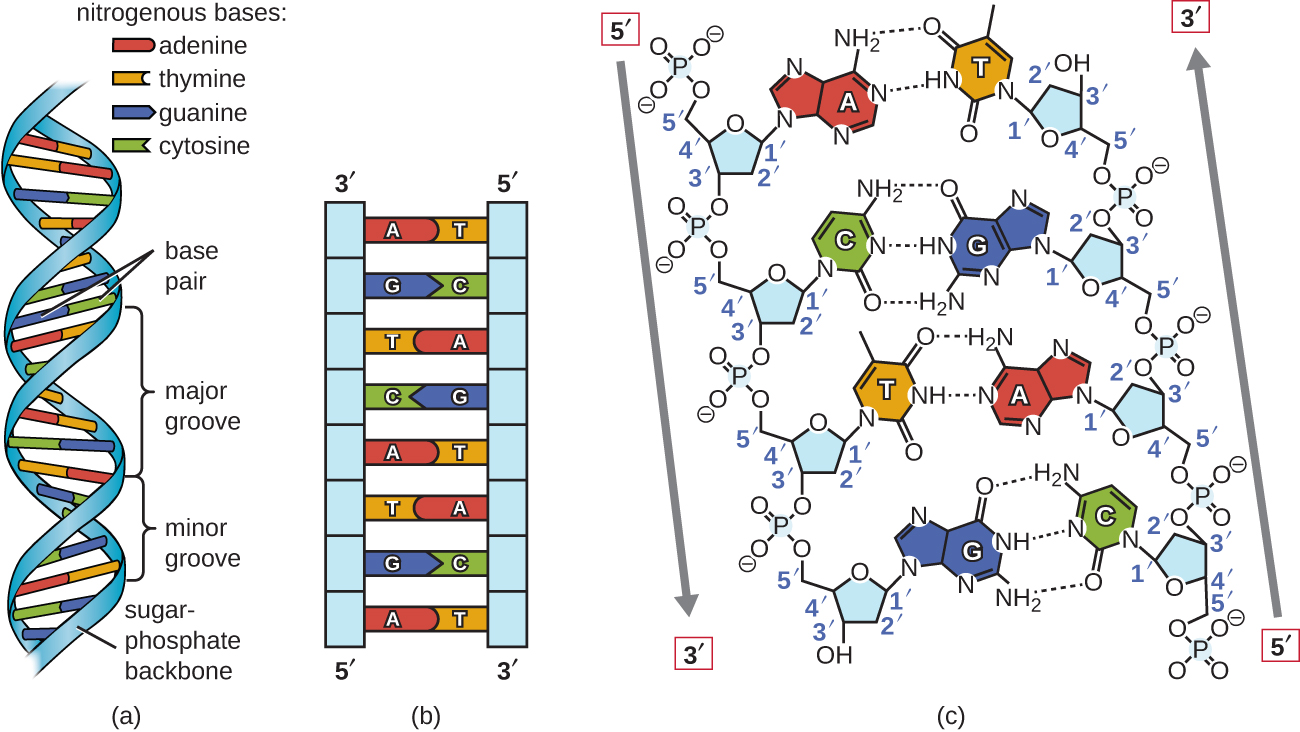
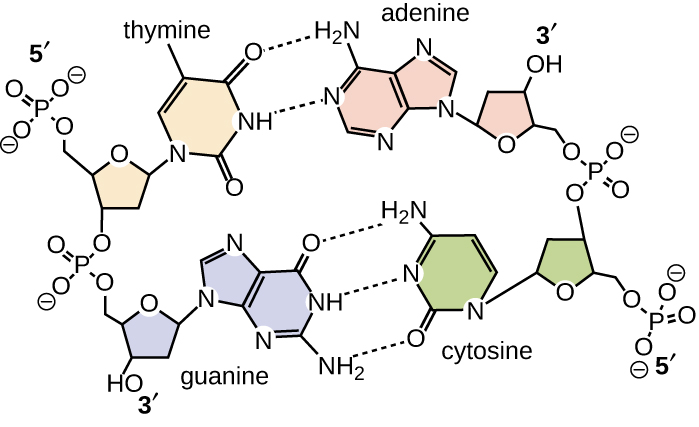
Base pairing takes place between a purine and pyrimidine. In DNA, adenine (A) and thymine (T) are complementary base pairs, and cytosine (C) and guanine (G) are also complementary base pairs, explaining Chargaff’s rules (Figure \(\PageIndex{7}\)). The base pairs are stabilized by hydrogen bonds; adenine and thymine form two hydrogen bonds between them, whereas cytosine and guanine form three hydrogen bonds between them.
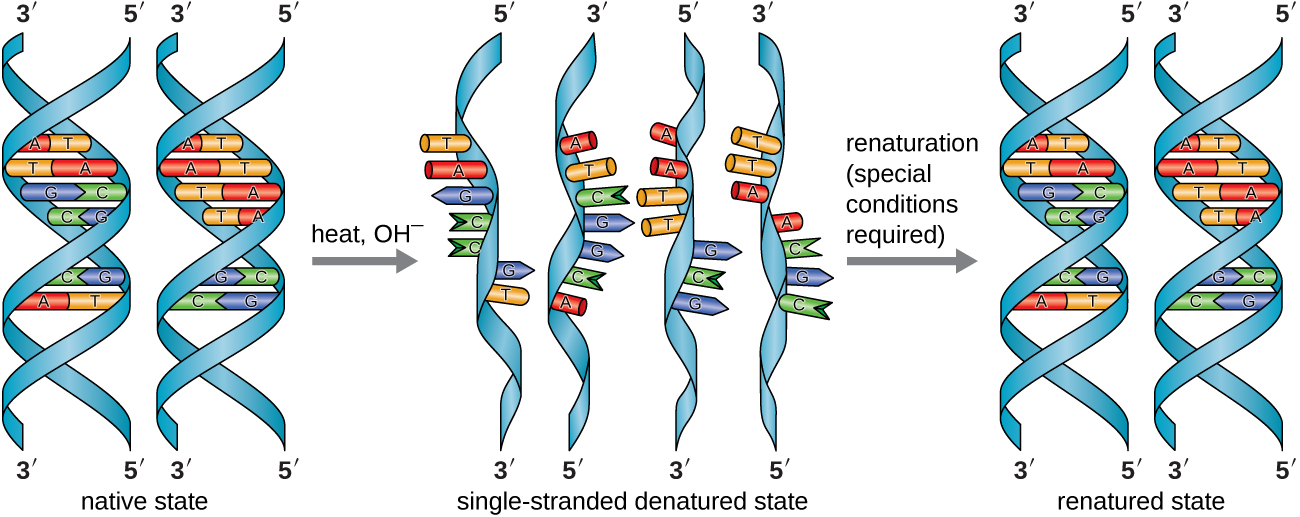
In the laboratory, exposing the two DNA strands of the double helix to high temperatures or to certain chemicals can break the hydrogen bonds between complementary bases, thus separating the strands into two separate single strands of DNA (single-stranded DNA [ssDNA]). This process is called DNA denaturation and is analogous to protein denaturation. The ssDNA strands can also be put back together as double-stranded DNA (dsDNA), through reannealing or renaturing by cooling or removing the chemical denaturants, allowing these hydrogen bonds to reform. The ability to artificially manipulate DNA in this way is the basis for several important techniques in biotechnology (Figure \(\PageIndex{8}\)). Because of the additional hydrogen bonding between the C = G base pair, DNA with a high GC content is more difficult to denature than DNA with a lower GC content.
View an animation on DNA structure from the DNA Learning Center to learn more.
What are the two complementary base pairs of DNA and how are they bonded together?
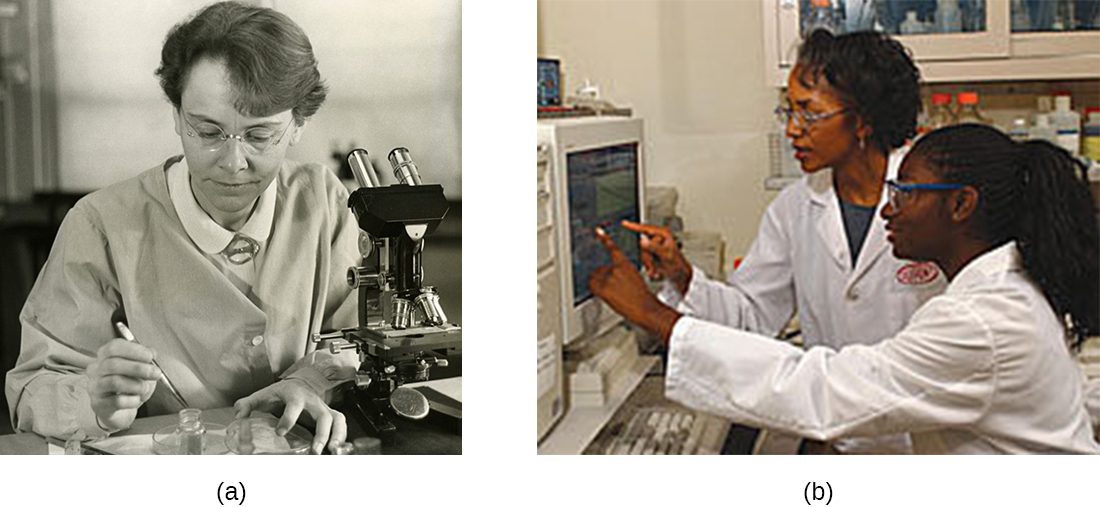
Historically, women have been underrepresented in the sciences and in medicine, and often their pioneering contributions have gone relatively unnoticed. For example, although Rosalind Franklin performed the X-ray diffraction studies demonstrating the double helical structure of DNA, it is Watson and Crick who became famous for this discovery, building on her data. There still remains great controversy over whether their acquisition of her data was appropriate and whether personality conflicts and gender bias contributed to the delayed recognition of her significant contributions. Similarly, Barbara McClintock did pioneering work in maize (corn) genetics from the 1930s through 1950s, discovering transposons (jumping genes), but she was not recognized until much later, receiving a Nobel Prize in Physiology or Medicine in 1983 (Figure \(\PageIndex{9}\)).
Today, women still remain underrepresented in many fields of science and medicine. While more than half of the undergraduate degrees in science are awarded to women, only 46% of doctoral degrees in science are awarded to women. In academia, the number of women at each level of career advancement continues to decrease, with women holding less than one-third of the positions of Ph.D.-level scientists in tenure-track positions, and less than one-quarter of the full professorships at 4-year colleges and universities.7 Even in the health professions, like nearly all other fields, women are often underrepresented in many medical careers and earn significantly less than their male counterparts, as shown in a 2013 study published by the Journal of the American Medical Association.8
Why do such disparities continue to exist and how do we break these cycles? The situation is complex and likely results from the combination of various factors, including how society conditions the behaviors of girls from a young age and supports their interests, both professionally and personally. Some have suggested that women do not belong in the laboratory, including Nobel Prize winner Tim Hunt, whose 2015 public comments suggesting that women are too emotional for science9 were met with widespread condemnation.
Perhaps girls should be supported more from a young age in the areas of science and math (Figure \(\PageIndex{9}\)). Science, technology, engineering, and mathematics (STEM) programs sponsored by the American Association of University Women (AAUW)10 and National Aeronautics and Space Administration (NASA)11 are excellent examples of programs that offer such support. Contributions by women in science should be made known more widely to the public, and marketing targeted to young girls should include more images of historically and professionally successful female scientists and medical professionals, encouraging all bright young minds, including girls and women, to pursue careers in science and medicine.

Based upon his symptoms, Alex’s physician suspects that he is suffering from a foodborne illness that he acquired during his travels. Possibilities include bacterial infection (e.g., enterotoxigenic E. coli, Vibrio cholerae, Campylobacter jejuni, Salmonella), viral infection (rotavirus or norovirus), or protozoan infection (Giardia lamblia, Cryptosporidium parvum, or Entamoeba histolytica).
His physician orders a stool sample to identify possible causative agents (e.g., bacteria, cysts) and to look for the presence of blood because certain types of infectious agents (like C. jejuni, Salmonella, and E. histolytica) are associated with the production of bloody stools.
Alex’s stool sample showed neither blood nor cysts. Following analysis of his stool sample and based upon his recent travel history, the hospital physician suspected that Alex was suffering from traveler’s diarrhea caused by enterotoxigenic E. coli (ETEC), the causative agent of most traveler’s diarrhea. To verify the diagnosis and rule out other possibilities, Alex’s physician ordered a diagnostic lab test of his stool sample to look for DNA sequences encoding specific virulence factors of ETEC. The physician instructed Alex to drink lots of fluids to replace what he was losing and discharged him from the hospital.
ETEC produces several plasmid-encoded virulence factors that make it pathogenic compared with typical E. coli. These include the secreted toxins heat-labile enterotoxin (LT) and heat-stabile enterotoxin (ST), as well as colonization factor (CF). Both LT and ST cause the excretion of chloride ions from intestinal cells to the intestinal lumen, causing a consequent loss of water from intestinal cells, resulting in diarrhea. CF encodes a bacterial protein that aids in allowing the bacterium to adhere to the lining of the small intestine.
Why did Alex’s physician use genetic analysis instead of either isolation of bacteria from the stool sample or direct Gram stain of the stool sample alone?
DNA Function
DNA stores the information needed to build and control the cell. The transmission of this information from mother to daughter cells is called vertical gene transfer and it occurs through the process of DNA replication. DNA is replicated when a cell makes a duplicate copy of its DNA, then the cell divides, resulting in the correct distribution of one DNA copy to each resulting cell. DNA can also be enzymatically degraded and used as a source of nucleotides for the cell. Unlike other macromolecules, DNA does not serve a structural role in cells.
The elucidation of the structure of the double helix by James Watson and Francis Crick in 1953 provided a hint as to how DNA is copied during the process of replication. Separating the strands of the double helix would provide two templates for the synthesis of new complementary strands, but exactly how new DNA molecules were constructed was still unclear. In one model, semiconservative replication, the two strands of the double helix separate during DNA replication, and each strand serves as a template from which the new complementary strand is copied; after replication, each double-stranded DNA includes one parental or “old” strand and one “new” strand. There were two competing models also suggested: conservative and dispersive, which are shown in Figure \(\PageIndex{10}\).
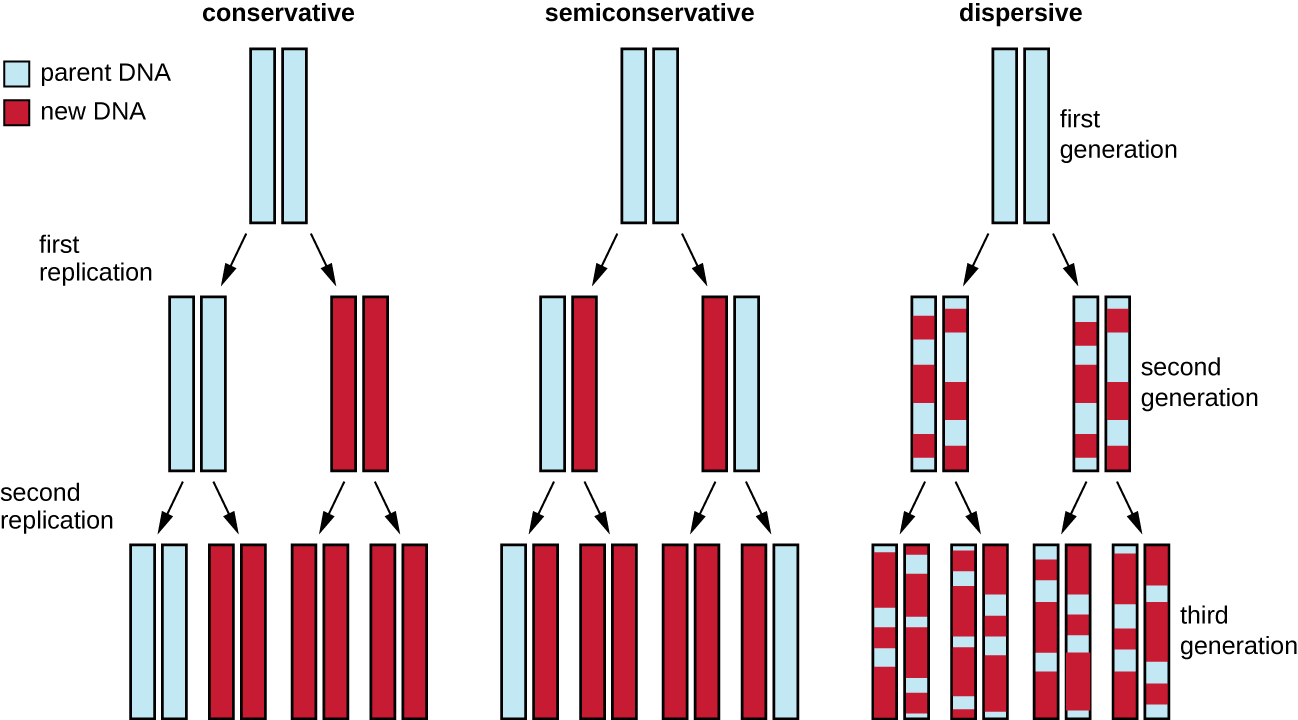
Matthew Meselson (1930–) and Franklin Stahl (1929–) devised an experiment in 1958 to test which of these models correctly represents DNA replication. They grew E. coli for several generations in a medium containing a “heavy” isotope of nitrogen (15N) that was incorporated into nitrogenous bases and, eventually, into the DNA. This labeled the parental DNA. The E. coli culture was then shifted into a medium containing 14N and allowed to grow for one generation. The cells were harvested and the DNA was isolated. The DNA was separated by ultracentrifugation, during which the DNA formed bands according to its density. DNA grown in 15N would be expected to form a band at a higher density position than that grown in 14N. Meselson and Stahl noted that after one generation of growth in 14N, the single band observed was intermediate in position in between DNA of cells grown exclusively in 15N or 14N. This suggested either a semiconservative or dispersive mode of replication. Some cells were allowed to grow for one more generation in 14N and spun again. The DNA harvested from cells grown for two generations in 14N formed two bands: one DNA band was at the intermediate position between 15N and 14N, and the other corresponded to the band of 14N DNA. These results could only be explained if DNA replicates in a semiconservative manner. Therefore, the other two models were ruled out. As a result of this experiment, we now know that during DNA replication, each of the two strands that make up the double helix serves as a template from which new strands are copied. The new strand will be complementary to the parental or “old” strand. The resulting DNA molecules have the same sequence and are divided equally into the two daughter cells. This means the middle explanation from Figure \(\PageIndex{10}\) is correct.
DNA Replication in Bacteria
DNA replication has been well studied in bacteria primarily because of the small size of the genome and the mutants that are available. E. coli has 4.6 million base pairs (Mbp) in a single circular chromosome and all of it is replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the circle bidirectionally (i.e., in both directions). This means that approximately 1000 nucleotides are added per second. The process is quite rapid and occurs with few errors.
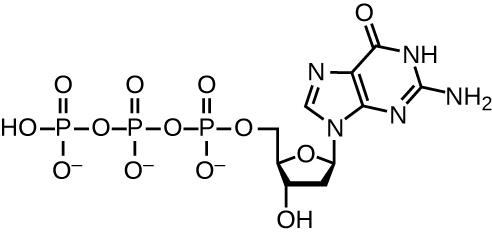
DNA replication uses a large number of proteins and enzymes (Table \(\PageIndex{1}\)). One of the key players is the enzyme DNA polymerase, also known as DNA pol. In bacteria, three main types of DNA polymerases are known: DNA pol I, DNA pol II, and DNA pol III. It is now known that DNA pol III is the enzyme required for DNA synthesis; DNA pol I and DNA pol II are primarily required for repair. DNA pol III adds deoxyribonucleotides each complementary to a nucleotide on the template strand, one by one to the 3’-OH group of the growing DNA chain. The addition of these nucleotides requires energy. This energy is present in the bonds of three phosphate groups attached to each nucleotide (a triphosphate nucleotide), similar to how energy is stored in the phosphate bonds of adenosine triphosphate (ATP) (Figure \(\PageIndex{11}\)). When the bond between the phosphates is broken and diphosphate is released, the energy released allows for the formation of a covalent phosphodiester bond by dehydration synthesis between the incoming nucleotide and the free 3’-OH group on the growing DNA strand.
Initiation
The initiation of replication occurs at specific nucleotide sequence called the origin of replication, where various proteins bind to begin the replication process. E. coli has a single origin of replication (as do most prokaryotes), called oriC, on its one chromosome. The origin of replication is approximately 245 base pairs long and is rich in adenine-thymine (AT) sequences.
Some of the proteins that bind to the origin of replication are important in making single-stranded regions of DNA accessible for replication. Chromosomal DNA is typically wrapped around histones (in eukaryotes and archaea) or histone-like proteins (in bacteria), and is supercoiled, or extensively wrapped and twisted on itself. This packaging makes the information in the DNA molecule inaccessible. However, enzymes can change the shape and supercoiling of the chromosome. For bacterial DNA replication to begin, the supercoiled chromosome is relaxed. An enzyme called helicase then separates the DNA strands by breaking the hydrogen bonds between the nitrogenous base pairs. Recall that AT sequences have fewer hydrogen bonds and, hence, have weaker interactions than guanine-cytosine (GC) sequences. These enzymes require ATP hydrolysis. As the DNA opens up, Y-shaped structures called replication forks are formed. Two replication forks are formed at the origin of replication, allowing for bidirectional replication and formation of a structure that looks like a bubble when viewed with a transmission electron microscope; as a result, this structure is called a replication bubble. The DNA near each replication fork is coated with single-stranded binding proteins to prevent the single-stranded DNA from rewinding into a double helix.
Once single-stranded DNA is accessible at the origin of replication, DNA replication can begin. However, DNA pol III is able to add nucleotides only in the 5’ to 3’ direction (a new DNA strand can be only extended in this direction). This is because DNA polymerase requires a free 3’-OH group to which it can add nucleotides by forming a covalent phosphodiester bond between the 3’-OH end and the 5’ phosphate of the next nucleotide. This also means that it cannot add nucleotides if a free 3’-OH group is not available, which is the case for a single strand of DNA. The problem is solved with the help of an RNA sequence that provides the free 3’-OH end. Because this sequence allows the start of DNA synthesis, it is appropriately called the primer. The primer is five to 10 nucleotides long and complementary to the parental or template DNA. It is synthesized by RNA primase, which is an RNA polymerase. Unlike DNA polymerases, RNA polymerases do not need a free 3’-OH group to synthesize an RNA molecule. Now that the primer provides the free 3’-OH group, DNA polymerase III can now extend this RNA primer, adding DNA nucleotides one by one that are complementary to the template strand.
Elongation
During elongation in DNA replication, the addition of nucleotides occurs at its maximal rate of about 1000 nucleotides per second. DNA polymerase III can only extend in the 5’ to 3’ direction, which poses a problem at the replication fork. The DNA double helix is antiparallel; that is, one strand is oriented in the 5’ to 3’ direction and the other is oriented in the 3’ to 5’ direction. During replication, one strand, which is complementary to the 3’ to 5’ parental DNA strand, is synthesized continuously toward the replication fork because polymerase can add nucleotides in this direction. This continuously synthesized strand is known as the leading strand. The other strand, complementary to the 5’ to 3’ parental DNA, grows away from the replication fork, so the polymerase must move back toward the replication fork to begin adding bases to a new primer, again in the direction away from the replication fork. It does so until it bumps into the previously synthesized strand and then it moves back again (Figure \(\PageIndex{12}\)). These steps produce small DNA sequence fragments known as Okazaki fragments, each separated by RNA primer. Okazaki fragments are named after the Japanese research team and married couple Reiji and Tsuneko Okazaki, who first discovered them in 1966. The strand with the Okazaki fragments is known as the lagging strand, and its synthesis is said to be discontinuous.
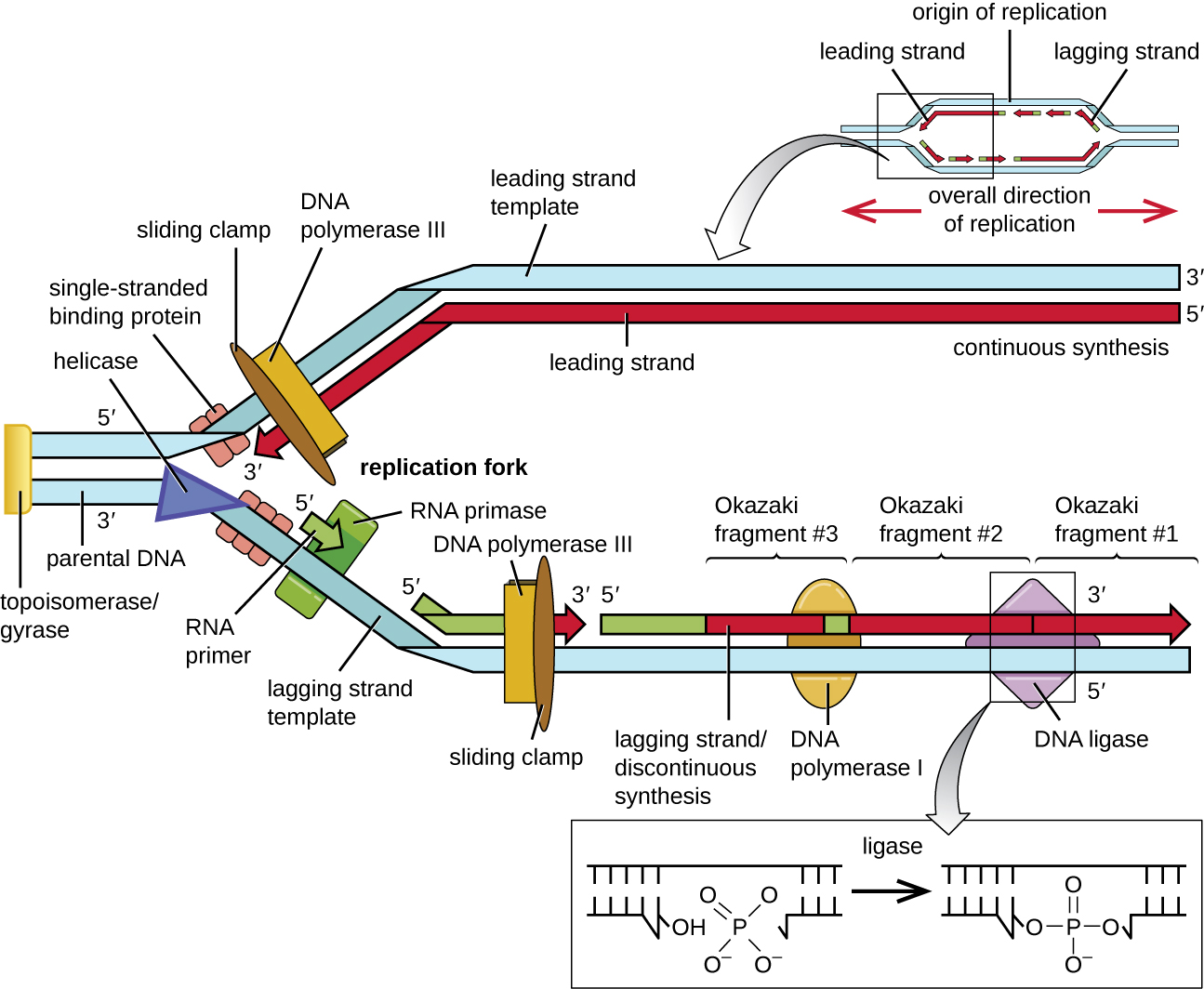
The leading strand can be extended from one primer alone, whereas the lagging strand needs a new primer for each of the short Okazaki fragments. The overall direction of the lagging strand will be 3’ to 5’, and that of the leading strand 5’ to 3’. A protein called the sliding clamp holds the DNA polymerase in place as it continues to add nucleotides. The sliding clamp is a ring-shaped protein that binds to the DNA and holds the polymerase in place. As synthesis proceeds, the RNA primers are replaced by DNA by the exonuclease activity of DNA polymerase I, and the gaps are filled in. Any nicks that remain between the newly synthesized DNA (that replaced the RNA primer) and the previously synthesized DNA are sealed by the enzyme DNA ligase, stabilizing the sugar-phosphate backbone of the DNA molecule.
Termination
Once the complete chromosome has been replicated, termination of DNA replication must occur. Although much is known about initiation of replication, less is known about the termination process. Following replication, the resulting complete circular genomes of prokaryotes are concatenated, meaning that the circular DNA chromosomes are interlocked and must be separated from each other. This is accomplished through the activity of bacterial topoisomerase IV, which introduces double-stranded breaks into DNA molecules, allowing them to separate from each other; the enzyme then reseals the circular chromosomes. The resolution of concatemers is an issue unique to prokaryotic DNA replication because of their circular chromosomes. Because both bacterial DNA gyrase and topoisomerase IV are distinct from their eukaryotic counterparts, these enzymes serve as targets for a class of antimicrobial drugs called quinolones.
- Which enzyme breaks the hydrogen bonds holding the two strands of DNA together so that replication can occur?
- Is it the lagging strand or the leading strand that is synthesized in the direction toward the opening of the replication fork?
- Which enzyme is responsible for removing the RNA primers in newly replicated bacterial DNA?
DNA Replication in Eukaryotes
Eukaryotic genomes are much more complex and larger than prokaryotic genomes and are typically composed of multiple linear chromosomes (Table \(\PageIndex{2}\)). The human genome, for example, has 3 billion base pairs per haploid set of chromosomes, and 6 billion base pairs are inserted during replication. There are multiple origins of replication on each eukaryotic chromosome (Figure \(\PageIndex{13}\)); the human genome has 30,000 to 50,000 origins of replication. The rate of replication is approximately 100 nucleotides per second—10 times slower than prokaryotic replication.
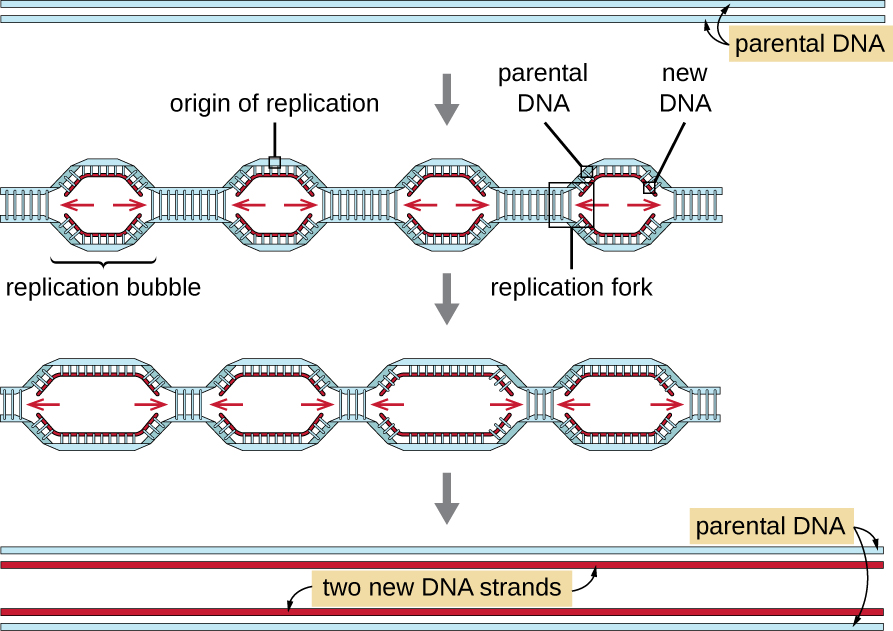
The essential steps of replication in eukaryotes are the same as in prokaryotes. Before replication can start, the DNA has to be made available as a template. Eukaryotic DNA is highly supercoiled and packaged, which is facilitated by many proteins, including histones. At the origin of replication, a prereplication complex composed of several proteins, including helicase, forms and recruits other enzymes involved in the initiation of replication, including topoisomerase to relax supercoiling, single-stranded binding protein, RNA primase, and DNA polymerase. Following initiation of replication, in a process similar to that found in prokaryotes, elongation is facilitated by eukaryotic DNA polymerases. The leading strand is continuously synthesized by the eukaryotic polymerase enzyme pol δ, while the lagging strand is synthesized by pol ε. A sliding clamp protein holds the DNA polymerase in place so that it does not fall off the DNA. The enzyme ribonuclease H (RNase H), instead of a DNA polymerase as in bacteria, removes the RNA primer, which is then replaced with DNA nucleotides. The gaps that remain are sealed by DNA ligase.
Because eukaryotic chromosomes are linear, one might expect that their replication would be more straightforward. As in prokaryotes, the eukaryotic DNA polymerase can add nucleotides only in the 5’ to 3’ direction. In the leading strand, synthesis continues until it reaches either the end of the chromosome or another replication fork progressing in the opposite direction. On the lagging strand, DNA is synthesized in short stretches, each of which is initiated by a separate primer. When the replication fork reaches the end of the linear chromosome, there is no place to make a primer for the DNA fragment to be copied at the end of the chromosome. These ends thus remain unpaired and, over time, they may get progressively shorter as cells continue to divide.
The ends of the linear chromosomes are known as telomeres and consist of noncoding repetitive sequences. The telomeres protect coding sequences from being lost as cells continue to divide. In humans, a six base-pair sequence, TTAGGG, is repeated 100 to 1000 times to form the telomere. The discovery of the enzyme telomerase clarified our understanding of how chromosome ends are maintained. Telomerase contains a catalytic part and a built-in RNA template. It attaches to the end of the chromosome, and complementary bases to the RNA template are added on the 3’ end of the DNA strand. Once the 3’ end of the lagging strand template is sufficiently elongated, DNA polymerase can add the nucleotides complementary to the ends of the chromosomes. In this way, the ends of the chromosomes are replicated. In humans, telomerase is typically active in germ cells and adult stem cells; it is not active in adult somatic cells and may be associated with the aging of these cells. Eukaryotic microbes including fungi and protozoans also produce telomerase to maintain chromosomal integrity. For her discovery of telomerase and its action, Elizabeth Blackburn (1948–) received the Nobel Prize for Medicine or Physiology in 2009.
- How does the origin of replication differ between eukaryotes and prokaryotes?
- What polymerase enzymes are responsible for DNA synthesis during eukaryotic replication?
- What is found at the ends of the chromosomes in eukaryotes and why?
Key Concepts and Summary
- Nucleic acids are composed of nucleotides, each of which contains a pentose sugar, a phosphate group, and a nitrogenous base. Deoxyribonucleotides within DNA contain deoxyribose as the pentose sugar.
- DNA contains the pyrimidines cytosine and thymine, and the purines adenine and guanine.
- Nucleotides are linked together by phosphodiester bonds between the 5ʹ phosphate group of one nucleotide and the 3ʹ hydroxyl group of another. A nucleic acid strand has a free phosphate group at the 5ʹ end and a free hydroxyl group at the 3ʹ end.
- Chargaff discovered that the amount of adenine is approximately equal to the amount of thymine in DNA, and that the amount of the guanine is approximately equal to cytosine. These relationships were later determined to be due to complementary base pairing.
- Watson and Crick, building on the work of Chargaff, Franklin and Gosling, and Wilkins, proposed the double helix model and base pairing for DNA structure.
- DNA is composed of two complementary strands oriented antiparallel to each other with the phosphodiester backbones on the exterior of the molecule. The nitrogenous bases of each strand face each other and complementary bases hydrogen bond to each other, stabilizing the double helix.
- Heat or chemicals can break the hydrogen bonds between complementary bases, denaturing DNA. Cooling or removing chemicals can lead to renaturation or reannealing of DNA by allowing hydrogen bonds to reform between complementary bases.
- DNA stores the instructions needed to build and control the cell. This information is transmitted from parent to offspring through vertical gene transfer.
- The DNA replication process is semiconservative, which results in two DNA molecules, each having one parental strand of DNA and one newly synthesized strand.
- In bacteria, the initiation of replication occurs at the origin of replication, where supercoiled DNA is unwound by DNA gyrase, made single-stranded by helicase, and bound by single-stranded binding protein to maintain its single-stranded state. Primase synthesizes a short RNA primer, providing a free 3’-OH group to which DNA polymerase III can add DNA nucleotides.
- During elongation, the leading strand of DNA is synthesized continuously from a single primer. The lagging strand is synthesized discontinuously in short Okazaki fragments, each requiring its own primer. The RNA primers are removed and replaced with DNA nucleotides by bacterial DNA polymerase I, and DNA ligase seals the gaps between these fragments.
- Termination of replication in bacteria involves the resolution of circular DNA concatemers by topoisomerase IV to release the two copies of the circular chromosome.
- Eukaryotes typically have multiple linear chromosomes, each with multiple origins of replication. Overall, replication in eukaryotes is similar to that in prokaryotes.
- The linear nature of eukaryotic chromosomes necessitates telomeres to protect genes near the end of the chromosomes. Telomerase extends telomeres, preventing their degradation, in some cell types.
Footnotes
- N. Kresge et al. “Chargaff's Rules: The Work of Erwin Chargaff.” Journal of Biological Chemistry 280 (2005):e21.
- L. Pauling, “A Proposed Structure for the Nucleic Acids.” Proceedings of the National Academy of Science of the United States of America 39 no. 2 (1953):84–97.
- J.D. Watson, F.H.C. Crick. “A Structure for Deoxyribose Nucleic Acid.” Nature 171 no. 4356 (1953):737–738.
- M.H.F. Wilkins et al. “Molecular Structure of Deoxypentose Nucleic Acids.” Nature 171 no. 4356 (1953):738–740.
- R. Franklin, R.G. Gosling. “Molecular Configuration in Sodium Thymonucleate.” Nature 171 no. 4356 (1953):740–741.
- R.O. Day et al. “A Crystalline Fragment of the Double Helix: The Structure of the Dinucleoside Phosphate Guanylyl-3',5'-Cytidine.” Proceedings of the National Academy of Sciences of the United States of America 70 no. 3 (1973):849–853.
- N.H. Wolfinger “For Female Scientists, There's No Good Time to Have Children.” The Atlantic July 29, 2013. www.theatlantic.com/sexes/arc...ildren/278165/.
- S.A. Seabury et al. “Trends in the Earnings of Male and Female Health Care Professionals in the United States, 1987 to 2010.” Journal of the American Medical Association Internal Medicine 173 no. 18 (2013):1748–1750.
- E. Chung. “Tim Hunt, Sexism and Science: The Real 'Trouble With Girls' in Labs.” CBC News Technology and Science, June 12, 2015. http://www.cbc.ca/news/technology/ti...labs-1.3110133. Accessed 8/4/2016.
- American Association of University Women. “Building a STEM Pipeline for Girls and Women.” www.aauw.org/what-we-do/stem-education/. Accessed June 10, 2016.
- National Aeronautics and Space Administration. “Outreach Programs: Women and Girls Initiative.” http://women.nasa.gov/outreach-programs/. Accessed June 10, 2016.
Contributors and Attributions
Nina Parker, (Shenandoah University), Mark Schneegurt (Wichita State University), Anh-Hue Thi Tu (Georgia Southwestern State University), Philip Lister (Central New Mexico Community College), and Brian M. Forster (Saint Joseph’s University) with many contributing authors. Original content via Openstax (CC BY 4.0; Access for free at https://openstax.org/books/microbiology/pages/1-introduction)


