17.4: Immune Evasion and Viral Virulence Factors
- Page ID
- 158889
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the mechanisms that bacteria and viruses use for immune evasion
- Evaluate the mechanisms viruses use for adhesion and antigenic variation
- Analyze the roles of antigenic drift and antigenic shift in viral evolution
Virulence Factors for Survival in the Host and Immune Evasion
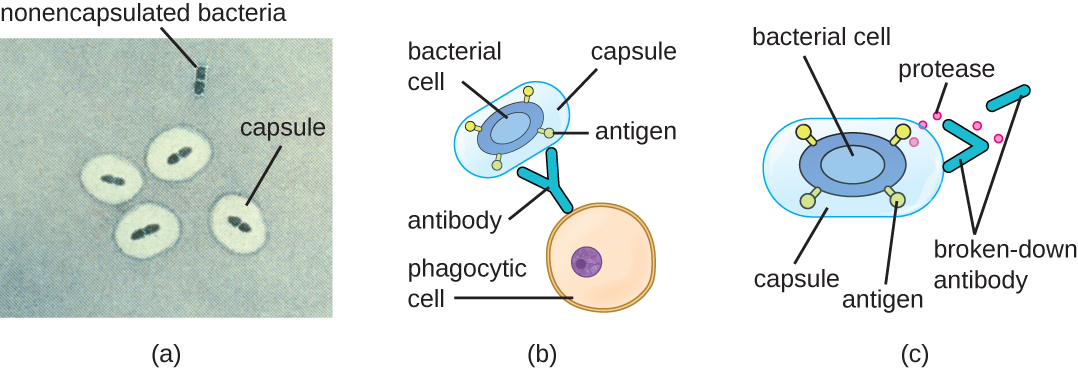
Evading the immune system is also important to invasiveness. Bacteria use a variety of virulence factors to evade phagocytosis by cells of the immune system. For example, many bacteria produce capsules, which are used in adhesion but also aid in immune evasion by preventing ingestion by phagocytes. The composition of the capsule prevents immune cells from being able to adhere and then phagocytose the cell. In addition, the capsule makes the bacterial cell much larger, making it harder for immune cells to engulf the pathogen (Figure \(\PageIndex{8}\)). A notable capsule-producing bacterium is the gram-positive pathogen Streptococcus pneumoniae, which causes pneumococcal pneumonia, meningitis, septicemia, and other respiratory tract infections. Encapsulated strains of S. pneumoniae are more virulent than nonencapsulated strains and are more likely to invade the bloodstream and cause septicemia and meningitis.
Some pathogens can also produce proteases to protect themselves against phagocytosis. As described in Adaptive Specific Host Defenses, the human immune system produces antibodies that bind to surface molecules found on specific bacteria (e.g., capsules, fimbriae, flagella, LPS). This binding initiates phagocytosis and other mechanisms of antibacterial killing and clearance. Proteases combat antibody-mediated killing and clearance by attacking and digesting the antibody molecules (Figure \(\PageIndex{8}\)).
In addition to capsules and proteases, some bacterial pathogens produce other virulence factors that allow them to evade the immune system. The fimbriae of certain species of Streptococcus contain M protein, which alters the surface of Streptococcus and inhibits phagocytosis by blocking the binding of the complement molecules that assist phagocytes in ingesting bacterial pathogens. The acid-fast bacterium Mycobacterium tuberculosis (the causative agent of tuberculosis) produces a waxy substance known as mycolic acid in its cell envelope. When it is engulfed by phagocytes in the lung, the protective mycolic acid coat enables the bacterium to resist some of the killing mechanisms within the phagolysosome.
Some bacteria produce virulence factors that promote infection by exploiting molecules naturally produced by the host. For example, most strains of Staphylococcus aureus produce the exoenzyme coagulase, which exploits the natural mechanism of blood clotting to evade the immune system. Normally, blood clotting is triggered in response to blood vessel damage; platelets begin to plug the clot, and a cascade of reactions occurs in which fibrinogen, a soluble protein made by the liver, is cleaved into fibrin. Fibrin is an insoluble, thread-like protein that binds to blood platelets, cross-links, and contracts to form a mesh of clumped platelets and red blood cells. The resulting clot prevents further loss of blood from the damaged blood vessels. However, if bacteria release coagulase into the bloodstream, the fibrinogen-to-fibrin cascade is triggered in the absence of blood vessel damage. The resulting clot coats the bacteria in fibrin, protecting the bacteria from exposure to phagocytic immune cells circulating in the bloodstream.
Whereas coagulase causes blood to clot, kinases have the opposite effect by triggering the conversion of plasminogen to plasmin, which is involved in the digestion of fibrin clots. By digesting a clot, kinases allow pathogens trapped in the clot to escape and spread, similar to the way that collagenase, hyaluronidase, and DNAse facilitate the spread of infection. Examples of kinases include staphylokinases and streptokinases, produced by Staphylococcus aureusand Streptococcus pyogenes, respectively. It is intriguing that S. aureus can produce both coagulase to promote clotting and staphylokinase to stimulate the digestion of clots. The action of the coagulase provides an important protective barrier from the immune system, but when nutrient supplies are diminished or other conditions signal a need for the pathogen to escape and spread, the production of staphylokinase can initiate this process.
A final mechanism that pathogens can use to protect themselves against the immune system is called antigenic variation, which is the alteration of surface proteins so that a pathogen is no longer recognized by the host’s immune system. For example, the bacterium Borrelia burgdorferi, the causative agent of Lyme disease, contains a surface lipoprotein known as VlsE. Because of genetic recombination during DNA replication and repair, this bacterial protein undergoes antigenic variation. Each time fever occurs, the VlsE protein in B. burgdorferi can differ so much that antibodies against previous VlsE sequences are not effective. It is believed that this variation in the VlsE contributes to the ability B. burgdorferi to cause chronic disease. Another important human bacterial pathogen that uses antigenic variation to avoid the immune system is Neisseria gonorrhoeae, which causes the sexually transmitted disease gonorrhea. This bacterium is well known for its ability to undergo antigenic variation of its type IV pili to avoid immune defenses.
Query \(\PageIndex{1}\)
Viral Virulence
Although viral pathogens are not similar to bacterial pathogens in terms of structure, some of the properties that contribute to their virulence are similar. Viruses use adhesins to facilitate adhesion to host cells, and certain enveloped viruses rely on antigenic variation to avoid the host immune defenses. These virulence factors are discussed in more detail in the following sections.
Viral Adhesins
One of the first steps in any viral infection is adhesion of the virus to specific receptors on the surface of cells. This process is mediated by adhesins that are part of the viral capsid or membrane envelope. The interaction of viral adhesins with specific cell receptors defines the tropism (preferential targeting) of viruses for specific cells, tissues, and organs in the body. The spike protein hemagglutinin found on Influenza virus is an example of a viral adhesin; it allows the virus to bind to the sialic acid on the membrane of host respiratory and intestinal cells. Another viral adhesin is the glycoprotein gp120, found on HIV. For HIV to infect cells of the immune system, it must interact with two receptors on the surface of cells. The first interaction involves binding between gp120 and the CD4 cellular marker that is found on some essential immune system cells. However, before viral entry into the cell can occur, a second interaction between gp120 and one of two chemokine receptors (CCR5 and CXCR4) must occur. Table \(\PageIndex{5}\) lists the adhesins for some common viral pathogens and the specific sites to which these adhesins allow viruses to attach.
| Pathogen | Disease | Adhesin | Attachment Site |
|---|---|---|---|
| Influenzavirus | Influenza | Hemagglutinin | Sialic acid of respiratory and intestinal cells |
| Herpes simplex virus I or II | Oral herpes, genital herpes | Glycoproteins gB, gC, gD | Heparan sulfate on mucosal surfaces of the mouth and genitals |
| Human immunodeficiency virus | HIV/AIDS | Glycoprotein gp120 | CD4 and CCR5 or CXCR4 of immune system cells |
Query \(\PageIndex{1}\)
Antigenic Variation in Viruses
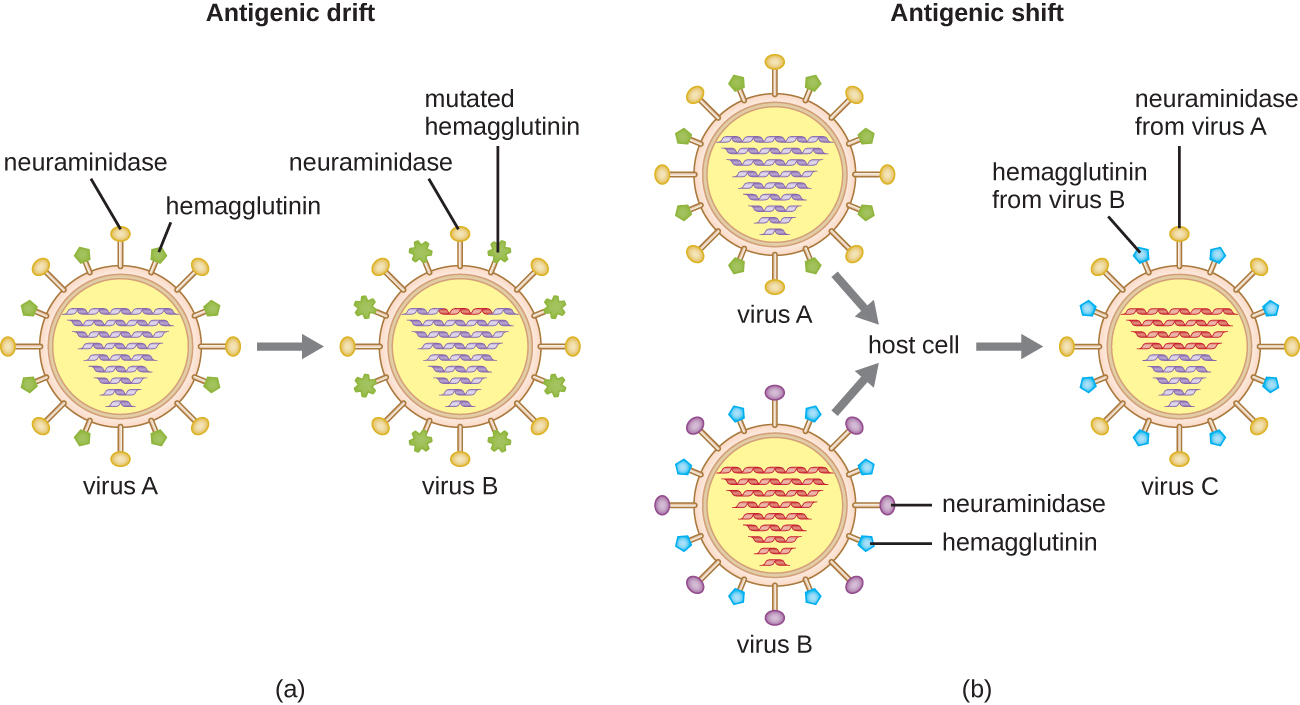
Antigenic variation also occurs in certain types of enveloped viruses, including influenza viruses, which exhibit two forms of antigenic variation: antigenic drift and antigenic shift (Figure \(\PageIndex{9}\)). Antigenic drift is the result of point mutations causing slight changes in the spike proteins hemagglutinin (H) and neuraminidase (N). On the other hand, antigenic shift is a major change in spike proteins due to gene reassortment. This reassortment for antigenic shift occurs typically when two different influenza viruses infect the same host.
The rate of antigenic variation in influenza viruses is very high, making it difficult for the immune system to recognize the many different strains of Influenza virus. Although the body may develop immunity to one strain through natural exposure or vaccination, antigenic variation results in the continual emergence of new strains that the immune system will not recognize. This is the main reason that vaccines against Influenza virus must be given annually. Each year’s influenza vaccine provides protection against the most prevalent strains for that year, but new or different strains may be more prevalent the following year.
For another explanation of how antigenic shift and drift occur, watch this video.
Query \(\PageIndex{1}\)
Key Concepts and Summary
Virulence factors contribute to a pathogen’s ability to cause disease. Exoenzymes and toxins allow pathogens to invade host tissue and cause tissue damage. Exoenzymes are classified according to the macromolecule they target and exotoxins are classified based on their mechanism of action. Bacterial toxins include endotoxin and exotoxins. Endotoxin is the lipid A component of the LPS of the gram-negative cell envelope. Exotoxins are proteins secreted mainly by gram-positive bacteria, but also are secreted by gram-negative bacteria. Bacterial pathogens may evade the host immune response by producing capsules to avoid phagocytosis, surviving the intracellular environment of phagocytes, degrading antibodies, or through antigenic variation. Viral pathogens use adhesins for initiating infections and antigenic variation to avoid immune defenses. Influenza viruses use both antigenic drift and antigenic shift to avoid being recognized by the immune system.
Footnotes
- 1 V. Meka. “Panton-Valentine Leukocidin.” http://www.antimicrobe.org/h04c.file...L-S-aureus.asp


