10: Transcription: RNA polymerases
- Page ID
- 378
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Recall the Central Dogma of molecular biology: DNA is transcribed into RNA, which is translated into protein. We will cover the material in that order, since that is the direction that information flows.
Introduction
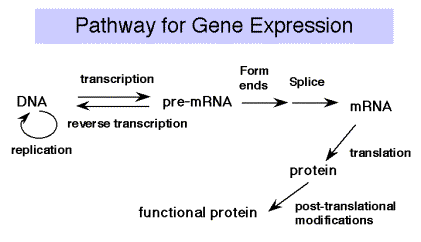
The liberated pyrophosphate is cleaved in the cell to 2 Pi, an energetically favorable reaction that drives the reaction in the direction of synthesis. In the presence of excess PPi, the reverse reaction of pyrophosphorolysis can occur. Synthesis always proceeds in a 5' to 3' direction (with respect to the growing RNA chain). The template is read in a 3' to 5' direction.
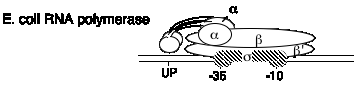
B. E. coli RNA polymerase structure
1. This one RNA polymerase synthesizes all classes of RNA
mRNA, rRNA, tRNA
2. It is composed of four subunits.
a. Core and holoenzyme
a2bb's a2bb' + s
Holoenzyme = a2bb's = core + s = can initiatetranscription accurately as the proper site, as determined by the promoter
Core = a2bb' = can elongate a growing RNA chain
A promoter can be defined in two ways.
- The sequence of DNA required for accurate, specific intiation of transcription
- The sequence of DNA to which RNA polymerase binds to accurately initiate transcription.
b. Subunits
| Subunit | Size | Gene | Function |
|---|---|---|---|
| b' | 160 kDa | rpoC | b' + b form the catalytic center. |
| b | 155 kDa | rpoB | b' + b form the catalytic center. |
| a | 40 kDa | rpoA | enzyme assembly; also binds UP sequence in the promoter |
| s | 70 kDa (general) | rpoD | confers specificity for promoter; binds to -10 and -35 sites in the promoter |
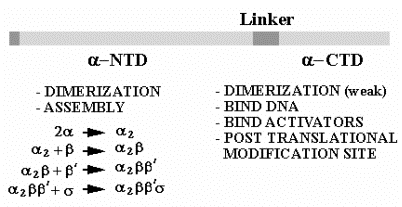
C. E. coli RNA polymerase mechanism
Mode of action of sfactors
The presence of the s factor causes the RNA polymerase holoenzyme to be selective in choosing the site of initiation. This is accomplished primarily through effects on the dissociation rate of RNA polymerase from DNA.
- a. Core has strong affinity for general DNA sequences. The t1/2 for dissociation of the complex of core‑DNA is about 60 min. This is useful during the elongation phase, but not during initiation.
- b. Holoenzyme has a reduced affinity for general DNA; it is decreased about 104 fold. The t1/2 for dissociation of holoenzyme from general DNA is reduced to about 1 sec.
- c. Holoenzyme has a greatly increased affinity for promoter sequences.The t1/2 for dissociation of holoenzyme from promoter sequences is of the order of hours.
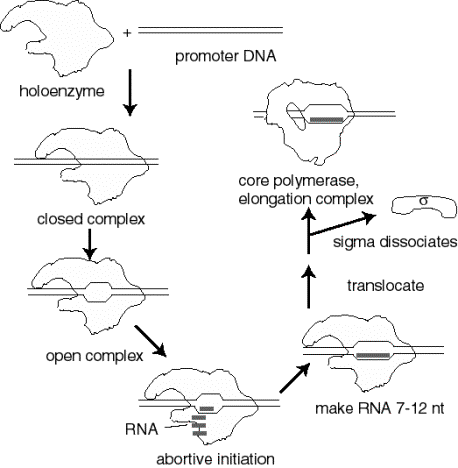
Events at initiation of transcription
- a. RNA polymerase holoenzyme binds to the promoter to form a closed complex; at this stage there is no unwinding of DNA.
- b. The polymerase‑promoter complex undergoes the closed to open transition, which is a melting or unwinding of about 12 bp.
- c. The initiating nucleotides can bind to the enzyme, as directed by their complementary nucleotides in the DNA template strand, and the enzyme will catalyze formation of a phosphodiester bond between them. This polymerase‑DNA‑RNA complex is referred to as the ternary complex.
- d. During abortive initiation, the polymerase catalyzes synthesis of short transcripts about 6 or so nucleotides long and then releases them.
- e. This phase ends when the nascent RNA of ~6 nucleotides binds to a second RNA binding site on the enzyme; this second site is distinct from the catalytic center. This binding is associated with "resetting" the catalytic center so that the enzyme will now catalyze the synthesis of oligonucleotides 7-12 long.
- f. The enzyme now translocates to an new position on the template. During this process sigma leaves the complex. A conformational change in the enzyme associated with sigma leaving the complex lets the "thumb" wrap around the DNA template, locking in processivity. Thus the core enzyme catalyzes RNA synthesis during elongation, which continues until "signals" are encountered which indicate termination.
Figure 3.1.8. Events at initiation
3. Transcription cycle
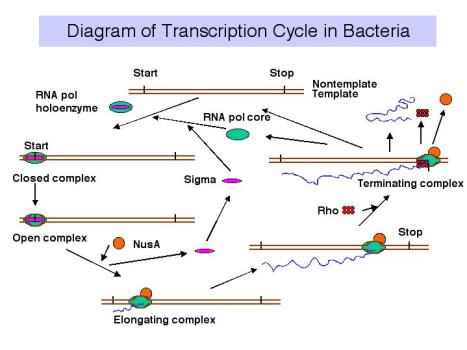
4. Sites on RNA Polymerase core
a. The enzyme covers about 60 bp of DNA, with a transcription bubble of about 17 bp unwound.
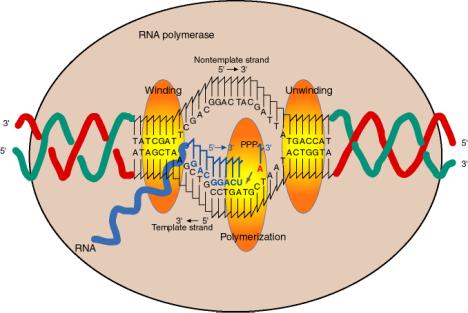
d The incoming nucleotide (NTP) that will be added to the growing RNA chain binds adjacent to the 3' end of the growing RNA chain, as directed by the template, at the active site for polymerization.
e. The incoming nucleotide is linked to the growing RNA chain by nucleophilic attack of the 3' OH on the a phosphoryl of the NTP, with liberation of pyrophosphate.
f. The reaction progresses (the enzyme moves) about 50 nts per sec. This is much slower than the rate of replication (about 1000 nts per sec).
g. If the template is topologically constrained, the DNA ahead of the RNA polymerase becomes overwound (positive superhelical turns) and the DNA behind the RNA polymerase becomes underwound (negative superhelical turns).
The effect of the unwinding of the DNA template by RNA polymerase is to decrease T by 1 for every 10 bp unwound. Thus DT = -1, and since DL = 0, then DW = +1 for every 10 bp unwound. This effect of the increase in W will be exerted in the DNA ahead of the polymerase.
The effect of rewinding the DNA template by RNA polymerase is just the opposite, of course. T will increase by 1 for every 10 bp rewound. Thus DT= +1, and since DL = 0, then DW = -1 for every 10 bp rewound. This effect of the decrease in W will be exerted in the DNA behind the polymerase, since that is where the rewinding is occurring.
5. Inhibitors: useful reagents and clues to function
a. Rifamycins, e.g. rifampicin: bind the b subunit to block initiation. The drug prevents addition of the 3rd or 4th nucleotide, hence the initiation process cannot be completed.
How do we know the site of rifampicin action is the b subunit? Mutations that confer resistance to rifampicin map to the rpoBgene.
b. Streptolydigins: bind to the b subunit to inhibit chain elongation.
These effects of rifamycins and streptolydigins, and the fact that they act on the b subunit, argue that the b subunit is required for nucleotide addition to the growing chain.
c. Heparin, a polyanion, binds to the b' subunit to prevent binding to DNA in vitro
D. Eukaryotic RNA polymerases
1. Eukaryotes have 3 different RNA polymerases in their nuclei.
- a. Each nuclear RNA polymerase is a large protein with about 8 to 14 subunits. MW is approximately 500,000 for each.
- b. Each polymerase has a different function:
| RNA polymerase | localization | synthesizes | effect of a‑amanitin |
|---|---|---|---|
| RNA polymerase I | nucleolus | pre‑rRNA | none |
| RNA polymerase II | nucleoplasm | pre‑mRNA some snRNAs | inhibited by low concentrations (0.03 mg/ml) |
| RNA polymerase III | nucleoplasm | pre‑tRNA, other small RNAs some snRNAs | inhibited by high concentrations (100 mg/ml) |
2. Subunit structures
a. The genes and encoded proteins for the subunits of the yeast RNA polymerases have been isolated and the sequences determined, and some functional analysis has been done.
b. Some of the subunits are homologous to bacterial RNA polymerases: The largest two subunits are homologs of b and b'. The roughly 40 kDa subunit is the homolog of a.
c. Some subunits are common to all three RNA polymerases.
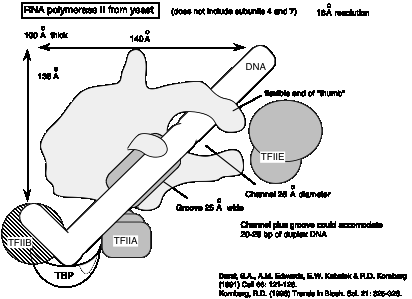
d. Example of yeast RNA polymerase II:
| Approximate size (kDa) | subunits per polymerase | role / comment | |
|---|---|---|---|
| 220 | 1 | related to b' | catalytic? |
| 130 | 1 | related to b | catalytic? |
| 40 | 2 | related to a | assembly? |
| 35 | < 1 | ||
| 30 | 2 | common to all 3 | |
| 27 | 1 | common to all 3 | |
| 24 | < 1 | ||
| 20 | 1 | common to all 3 | |
| 14 | 2 | ||
| 10 | 1 |
e. The largest subunit has a carboxy‑terminal domain (CTD)with an unusual structure: tandem repeats of the sequence Tyr‑Ser‑Pro‑Thr‑Ser‑Pro‑Thr. The yeast enzyme has 26 tandem repeats and the mammalian enzyme has about 50. These can be phophorylated on Ser and Thr to give a highly charged CTD.
- RNA Pol IIa is not phosphorylated in the CTD.
- RNA Pol IIo is phosphorylated in the CTD.

4. RNA polymerases in chloroplasts (plastids) and mitochondria
- a. The RNA polymerase found in plastids is encoded on the plastid chromosome. In some species the mitochondrial RNA polymerase is encoded by the mitochondrial DNA.
- b. These organellar RNA polymerases are much more related to the bacterial RNA polymerases than to the nuclear RNA polymerases. This is a strong argument in favor of the origins of these organelles being bacterial, supporting the endo-symbiotnt model for acquisition of these organelles in eukaryotes.
- c. These RNA polymerases catalyze specific transcription of organellar genes.
E. General transcription factors for eukaryotic RNA polymerase II
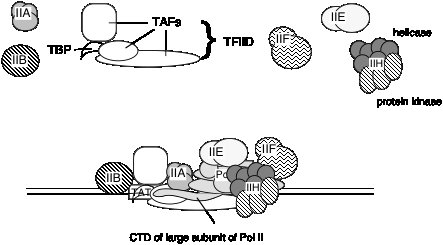
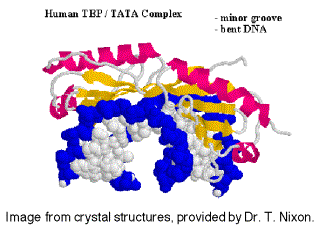
It is not known if the same set of TAFs are in the TFIID for all promoters transcribed by RNA polymerase II, or if some are used only for certain types of promoters. TFIID is the only sequence‑specific general transcription factor so far characterized, and it binds in the minor groove of the DNA. It is also used at TATA‑less promoters, so the role of the sequence ‑specific binding is still under investigation.
3. Summary of general transcription factors for RNA polymerase II.
Factors for RNA polymerase II (human cells)
| Factor | No. of subunits | Molecular mass (kDa) | Functions | Functions to Recruit: |
|---|---|---|---|---|
| TFIID: TBP | 1 | 38 | Recognize core promoter (TATA) | TFIIB |
| TFIID: TAFs | 12 | 15-250 | Recognize core promoter (non-TATA); Positive and negative regulation | RNA Pol II? |
| TFIIA | 2 | 12, 19, 35 | Stabilize TBP-DNA binding; Anti-repression | |
| TFIIB | 1 | 35 | Select start site for RNA Pol II | RNA PolII-TFIIF |
| RNA Pol II | 12 | 10-220 | Catalyze RNA synthesis | TFIIE |
| TFIIF | 2 | 30, 74 | Target RNA PolII to promoter; destabilize non-specific interactions between PolII and DNA | |
| TFIIE | 2 | 34, 57 | Modulate TFIIH helicase, ATPase and kinase activities; Directly enhance promoter melting? | TFIIH |
| TFIIH | 9 | 35-89 | Helicase to melt promoter; CTD kinase; promoter clearance? |
Roeder, R.G. (1996) TIBS 21: 327-335.
4. TFIIH is a multisubunit transcription factor also involved in DNA repair.
Subunits of the human factor
| Gene | Molec. mass of protein (kDa) | Function/ Structure | Proposed Role |
|---|---|---|---|
| XPB | 89 | helicase, tracks 3' to 5' | Unwind duplex for transcription/ Repair |
| XPD | 80 | helicase, tracks 5' to 3' | Unwind duplex, Repair |
| P62 | 62 | unknown | |
| P52 | 52 | unknown | |
| P44 | 44 | Zn-finger | Binds DNA |
| P34 | 34 | Zn-finger | |
| MAT1 | 32 | CDK assembly factor | |
| Cyclin H | 38 | Cyclin partner for CDK7/MO15 | |
| CDK7/MO15 | 32 | Protein kinase | Kinase for CTD |

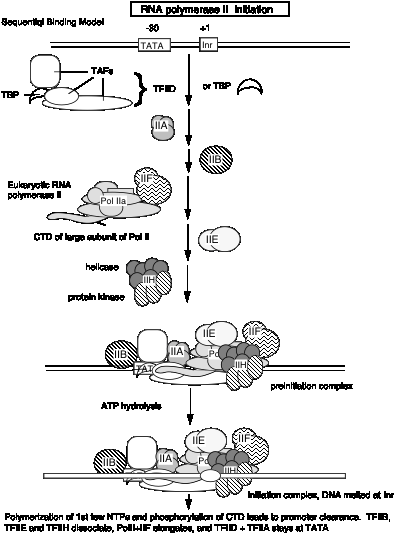
Table 3.1.6. RNA polymerase II holoenzyme and mediator
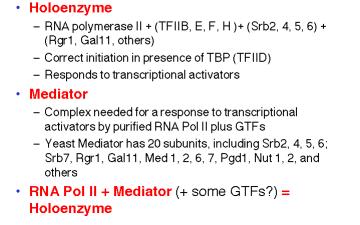

These studies show that RNA polymerase II can exist in several different states or complexes. One is in a very large holocomplex containing the mediator. In this state, it will accurately initiate transcription when directed by TFIID, and respond to activators (Table 3.1.6). The mediator subcomplex appears to be able to dissociate and reassociate with RNA polymerase II and GTFs. Indeed, this reassociation could be the step that was assayed in the identification of mediator. Without mediator, RNA polymerase II plus GTFs can initiate transcription at the correct place (as directed by TFIID), but they do not respond to activators. In the absence of GTFs, RNA polymerase II is capable of transcribing DNA templates, but it will not begin transcription at the correct site. Hence it is competent for elongation but not initiation.
Table 3.1.7. Expanding the functions of RNA polymerase II
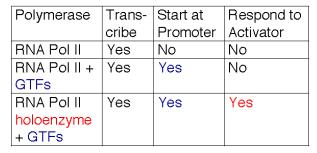
Fig.3.1.17
![]()
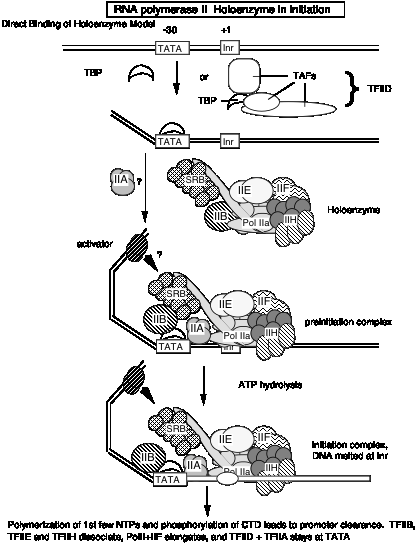
If the holoenzyme is the primary enzyme involved in transcription initiation in eukaryotic cells, then the progressive assembly pathway observed in vitro(see section d above) may be of little relevance in vivo. Perhaps the holoenzyme will bind to promoters simply marked by binding of TBP (or TFIID) to the TATA box, in contrast to the progressive assembly model that has a more extensive, ordered assembly mechanism. In both models, TBP or TFIID binding is the initial step in assembly of the preinitiation complex. However, at this point one cannot rule out the possibility that the holoenzyme is used at some promoters, and progressive assembly occurs at others.
7. Targets for the activator proteins
The targets for transcriptional activator proteins may be some component of the initiation complex. One line of investigation is pointing to the TAFs in TFIID as well as TFIIB as targets for the activators. Thus the activators may facilitate the ordered assembly of the intiation complex by recruiting GTFs. However, the holoenzyme contains the "mediator" or SRB complex that can mediate response to activators. Thus the activators may serve to recruit the holoenzyme to the promoter. Further studies are required to establish whether one or the other is correct, or if these are separate paths to activation.
F. General transcription factors for eukaryotic RNA polymerases I and III
1. General transcription factors for RNA polymerase I
a. Core promoter covers the start site of transcription, plus an upstream control element located about 70 bp further 5'.
b. The factor UBF1 binds to a G+C rich sequence in both the upstream control element and in the core promoter.
c. A multisubunit complex called SL1 binds to the UBF1‑DNA complex, again at both the upstream and core elements.
d. One of the subuntis of SL1 is TBP ‑ the TATA‑binding protein from TFIID!
e. RNA polymerase I then binds to this complex of DNA+UBF1+SL1 to initiate transcription at the correct nucleotide and the elongate to make pre‑rRNA.
2. General transcription factors for RNA Pol III
- a. Internal control sequences are characteristic of genes transcribed by RNA Pol III (see below).
- b. TFIIIA: binds to the internal control region of genes that encode 5S RNA (type 1 internal promoter)
- c. TFIIIC: binds to internal control regions of genes for 5S RNA (alongside TFIIIA) and for tRNAs (type 2 internal promoters)
- d. TFIIIB: The binding of TFIIIC directs TFIIIB to bind to sequences (-40 to +11) that overlap the start site for transcription. One subunit of TFIIIB is TBP, even though no TATA box is required for transcription. TFIIIA and TFIIIC can now be removed without affecting the ability of RNA polymerase III to initiate transcription. Thus TFIIIA and TFIIIC are assembly factors, and TFIIIB is the initiation factor.
Figure 3.1.18.
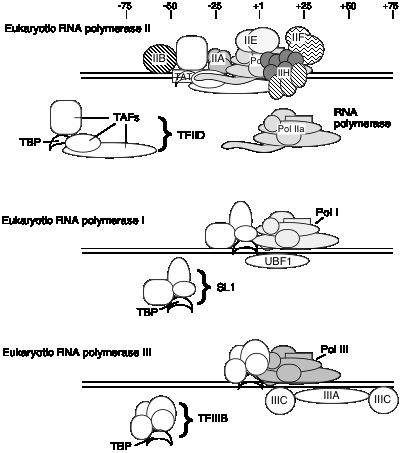
e. RNA polymerase III binds to the complex of TFIIIB+DNA to accurately and efficiently initiated transcription.
3 Transcription factor used by all 3 RNA Pol'ases: TBP
TBP seems to play a common role in directing RNA polymerase (I, II and III) to initiate at the correct place. The multisubunit factors that contain TBP (TFIID, SL1 and TFIIIB) may serve as positioning factors for their respective polymerases.


