9.6: Classes of Transposable Elements
- Page ID
- 366
The two major classes of transposable elements are defined by the intermediates in the transposition process. One class moves by DNA intermediates, using transposases and DNA polymerases to catalyze transposition. The other class moves by RNA intermediates, using RNA polymerase, endonucleases and reverse transcriptase to catalyze the process. Both classes are abundant in many species, but some groups of organisms have a preponderance of one or the other. For instance, bacteria have mainly the DNA intermediate class of transposable elements, whereas the predominant transposable elements in mammalian genomes move by RNA intermediates.
- Transposable elements that move via DNA intermediates
- Transposable elements that move via RNA intermediates
Transposable elements that move via DNA intermediates
Among the most thoroughly characterized transposable elements are those that move by DNA intermediates. In bacteria, these are either short insertion sequences or longer transposons.
An insertion sequences, or IS, is a short DNA sequence that moves from one location to another. They were first recognized by the mutations they cause by inserting into bacterial genes. Different insertion sequences range in size from about 800 bp to 2000 bp. The DNA sequence of an IS has inverted repeats (about 10 to 40 bp) at its termini (Figure 9.10A.). Note that this is different from the FDRs, which are duplications of the target site. The inverted repeats are part of the IS element itself. The sequences of the inverted repeats at each end of the IS are very similar but not necessarily identical. Each family of insertion sequence in a species is named IS followed by a number, e.g. IS1, IS10, etc.
An insertion sequence encodes a transposase enzyme that catalyzes the transposition. The amount of transposase is well regulated and is the primary determinant of the rate of transposition. Transposons are larger transposable elements, ranging in size from 2500 to 21,000 bp. They usually encode a drug resistance gene or other marker besides the functions required for transposition (Figure 9.10.B.). One type of transposon, called a composite transposon, has an IS element at each end (Figure 9.10.C.). One or both IS elements may be functional; these encode the transposition function for this class of transposons. The IS elements flank the drug resistance gene (or other selectable marker). It is likely that the composite transposon evolved when two IS elements inserted on both sides of a gene. The IS elements at the end could either move by themselves or they can recognize the ends of the closely spaced IS elements and move them together with the DNA between them. If the DNA between the IS elements confers a selective advantage when transposed, then it will become fixed in a population.
Exercise 9.3
What are the predictions of this model for formation of a composite transposon for the situation in which a transposon in a small circular replicon, such as a plasmid?
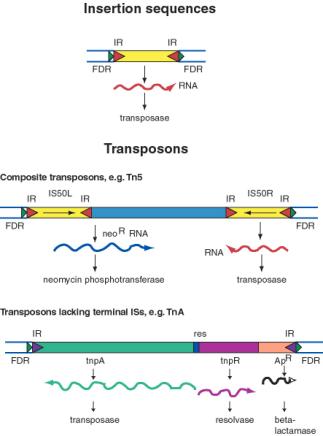
The TnA family of transposons has been intensively studied for the mechanism of transposition. Members of the TnA family have terminal inverted repeats, but lack terminal IS elements (Figure 9.10). The tnpAgene of the TnA transposon encodes a transposase, and the tnpRgene encodes a resolvase. TnA also has a selectable marker, ApR, which encodes a beta-lactamase and makes the bacteria resistance to ampicillin.
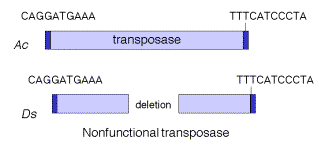
Transposable elements that move via DNA intermediates are not limited to bacteria, but rather they are found in many species. The P elements and copiafamily of repeats are examples of such transposable elements in Drosophila, as are marinerelements in mammals and the controlling elements in plants. Indeed, the general structure of controlling elements in maize is similar to that of bacterial transposons. In particular, they end in inverted repeats and encode a transposase. As illustrated in Figure 9.11, the DNA sequences at the ends of an Ac element are very similar to those of a Dselement. However, internal regions, which normally encode the transposase, have been deleted. This is why Dselements cannot transpose by themselves, but rather they require the presence of the intact transposon, Ac, in the cell to provide the transposase. Since transposase works in trans, the Acelement can be anywhere in the genome, but it can act on Dselements at a variety of sites. Note that Ac is an autonomous transposon because it provides its own transposase and it has the inverted repeats needed to act as the substrate for transposase.
Mechanism of DNA-mediated transposition
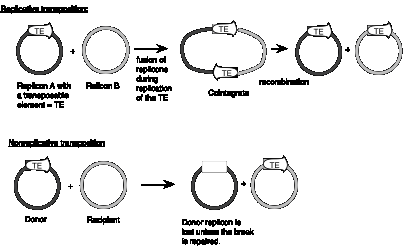
Some families of transposable elements that move via a DNA intermediate do so in a replicative manner. In this case, transposition generates a new copy of the transposable element at the target site, while leaving a copy behind at the original site. A cointegrate structure is formed by fusion of the donor and recipient replicons, which is then resolved (Figure 9.12). Other families use a nonreplicative mechanism. In this case, the original copy excises from the original site and move to a new target site, leaving the original site vacant.
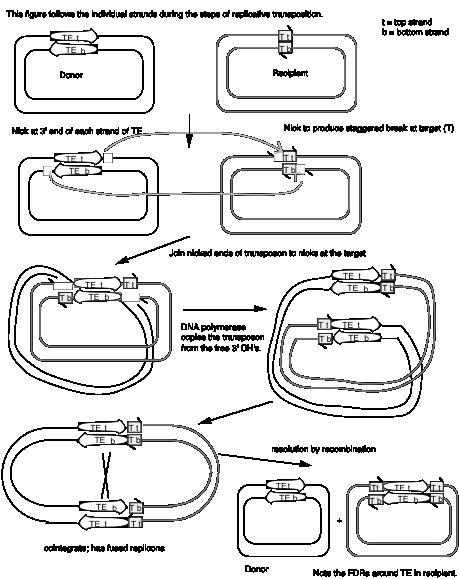
Studies of bacterial transposons have shown that replicative transposition and some types of nonreplicative transposition proceed through a strand-transfer intermediate (also known as a crossover structure), in which both the donor and recipient replicons are attached to the transposable element (Figure 9.13). For replicative transposition, DNA synthesis through the strand-transfer intermediate produces a transposable element at both the donor and target sites, forming the cointegrate intermediate. This is subsequently resolved to separate the replicons. DNA synthesis does not occur at the crossover structure in nonreplicative transposition, thus leaving a copy only at the new target site. In an alternative pathway for nonreplicative transposition, the transposon is excised by two double strand breaks, and is joined to the recipient at a staggered break (illustrated at the bottom of Figure 9.12).
In more detail, there are two steps in common for replicative and nonreplicative transposition, generating the strand-transfer intermediate (Figure 9.13).
- The transposase encoded by a transposable element makes four nicks initially. Two nicks are made at the target site, one in each strand, to generate a staggered break with 5' extensions (3' recessed). The other two nicks flank the transposon; one nick is made in one DNA strand at one end of the transposon, and the other nick is made in the other DNA strand at the other end. Since the transposon has inverted repeats at each end, these two nicks that flank the transposon are cleavages in the same sequence. Thus the transposase has a sequence-specific nicking activity. For instance, the transposase from TnA binds to a sequence of about 25 bp located within the 38 bp of inverted terminal repeat (Figure 9.10). It nicks a single strand at each end of the transposon, as well as the target site (Figure 9.13). Note that although the target and transposon are shown apart in the two-dimensional drawing in Figure 9.13, they are juxtaposed during transposition.
- At each end of the transposon, the 3' end of one strand of the transposon is joined to the 5' extension of one strand at the target site. This ligation is also catalyzed by transposase. ATP stimulates the reaction but it can occur in the absence of ATP if the substrate is supercoiled. Ligation of the ends of the transposon to the target site generates a strand-transfer intermediate, in which the donor and recipient replicons are now joined by the transposon.
After formation of the strand-transfer intermediate, two different pathways can be followed. For replicative transposition, the 3' ends of each strand of the staggered break (originally at the target site) serve as primers for repair synthesis (Figure 9.13). Replication followed by ligation leads to the formation of the cointegrate structure, which can then be resolved into the separate replicons, each with a copy of the transposon. The resolvase encoded by transposon TnA catalyzes the resolution of the cointegrate structure. The site for resolution (res) is located between the divergently transcribed genes for tnpA and tnpR (Figure 9.10). TnA resolvase also negatively regulates expression of both tnpA and tnpR (itself).
For nonreplicative transposition, the strand-transfer intermediate is released by nicking at the ends of the transposon not initially nicked. Repair synthesis is limited to the gap at the flanking direct repeats, and hence only one copy of the transposon is left. This copy is ligated to the new target site, leaving a vacant site in the donor molecule.
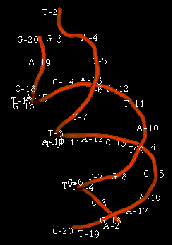
The enzyme transposase can recognize specific DNA sequences, cleave two duplex DNA molecules in four places, and ligate strands from the donor to the recipient. This enzyme has a remarkable ability to generate and manipulate the ends of DNA. A three-dimensional structure for the Tn5 transposase in complex with the ends of the Tn5 DNA has been solved by Rayment and colleagues. One static view of this protein DNA complex is in Figure 9.14.A. The transposase is a dimer, and each double-stranded DNA molecule (donor and target) is bound by both protein subunits. This orients the transposon ends into the active sites, as shown in the figure. Also, an image with just the DNA (Figure 9.14.B.) shows considerable distortion of the DNA helix at the ends. This recently determined structure is a good starting point to better understand the mechanism for strand cleavage and transfer.

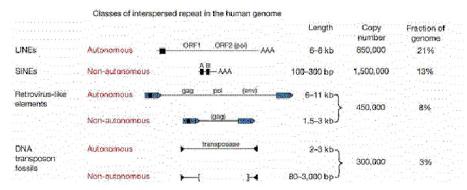
Transposable elements that move via RNA intermediates
Transposable DNA sequences that move by an RNA intermediate are called retrotransposons. They are very common in eukaryotic organisms, but some examples have also been found in bacteria. Some retrotransposons have long terminal repeats (LTRs) that regulate expression (Figure 9.15). The LTRs were initially discovered in retroviruses. They have now been seen in some but not all retrotransposons. They have a strong promoter and enhancer, as well as signals for forming the 3’ end of mRNAs after transcription. The presence of the LTR is distinctive for this family, and members are referred to as LTR-containing retrotransposons. Examples include the yeast Ty-1family and retroviral proviruses in vertebrates. Retroviral proviruses encode a reverse transcriptase and an endonuclease, as well as other proteins, some of which are needed for viral assembly and structure.
Others retrotransposons are in the large and diverse class of non-LTR retrotransposons (Figure 9.15). One of the most prevalent examples is the family of long interspersed repetitive elements, or LINEs. It was initially found in mammals but has now been found in a broad range of phyla, including fungi. The first and most common LINE family in mammals is the LINE1 family, also called L1. An older family, but discovered later, is called LINE2. Full-length LINEs are about 7000 bp long, and there are about 10,000 copies in humans. Many other copies are truncated from the 5’ ends. Like retroviral proviruses, the full-length L1 encodes a reverse transcriptase and an endonuclease, as well as other proteins. However, the promoter is not an LTR. Other abundant non-LTR retrotransposons, initially discovered in mammals, are short interspersed repetitive elements, or SINEs. These are about 300 bp long. Alurepeats, with over a million copies, comprise the predominant class of SINEs in humans. Non-LTR retrotransposons besides LINEs are found in many other species, such as jockey repeats in Drosophila.
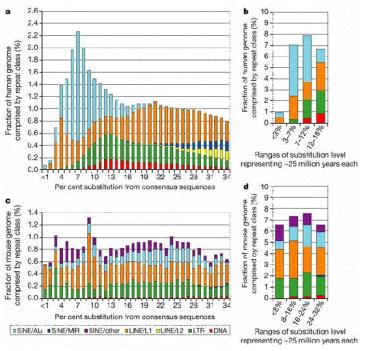
Extensive studies in of genomic DNA sequences have allowed the reconstruction of the history of transposable elements in humans and other mammals. The major approach has been to classify the various types of repeats (themselves transposable elements), align the sequences and determine how different the members of a family are from each other. Since the vast majority of the repeats are no longer active in transposition, and have no other obvious function, they will accumulate mutations rapidly, at the neutral rate. Thus the sequence of more recently transposing members are more similar to the source sequence than are the members that transposed earlier. The results of this analysis show that the different families of repeats have propagated in distinct waves through evolution (Figure 9.16). The LINE2 elements were abundant prior to the mammalian divergence, roughly 100 million years ago. Both LINE1 and Alu repeats have propagated more recently in humans. It is likely that the LINE1 elements, which encode a nuclease and a reverse transcriptase, provide functions needed for the transposition and expansion of Alu repeats. LINE1 elements have expanded in all orders of mammals, but each order has a distinctive SINE, all of which are derived from a gene transcribed by RNA polymerase III. This has led to the idea that LINE1 elements provide functions that other different transcription units use for transposition.
Mechanism of retrotransposition
Although the mechanism of retrotransposition is not completely understood, it is clear that at least two enzymatic activities are utilized. One is an integrase, which is an endonuclease that cleaves at the site of integration to generate a staggered break (Figure 9.17). The other is RNA-dependent DNA polymerase, also called reverse transcriptase. These activities are encoded in some autonomous retrotransposons, including both LTR-retrotransposons such as retroviral proviruses and non-LTR-retrotransposons such as LINE1 elements.
The RNA transcript of the transposable element interacts with the site of cleavage at the DNA target site. One strand of DNA at the cleaved integration site serves as the primer for reverse transcriptase. This DNA polymerase then copies the RNA into DNA. That cDNA copy of the retrotransposon must be converted to a double stranded product and inserted at a staggered break at the target site. The enzymes required for joining the reverse transcript (first strand of the new copy) to the other end of the staggered break and for second strand synthesis have not yet been established. Perhaps some cellular DNA repair functions are used.
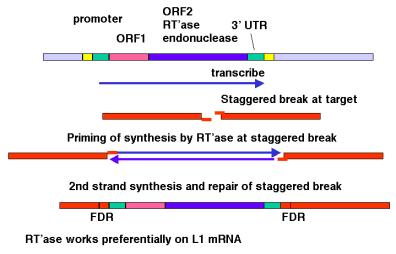
The model shown in Figure 9.17 is consistent with any RNA serving as the template for synthesis of the cDNA from the staggered break. However, LINE1 mRNA is clearly used much more often than other RNAs. The basis for the preference of the retrotransposition machinery for LINE1 mRNA is still being studied. Perhaps the endonuclease and reverse transcriptase stay associated with the mRNA that encodes them after translation has been completed, so that they act in cis with respect to the LINE1 mRNA. Other repeats that have expanded recently, such as Alu repeats in humans, may share sequence determinants with LINE1 mRNA for this cis preference.
Clear evidence that retrotransposons can move via an RNA intermediate came from studies of the yeast Ty-1 elements by Gerald Fink and his colleagues. They placed a particular Ty-1 element, called TyH3 under control of a GAL promoter, so that its transcription (and transposition) could be induced by adding galactose to the media. They also marked TyH3 with an intron. After inducing transcription of TyH3, additional copies were found at new locations in the yeast strain. When these were examined structurally, it was discovered that the intron had been removed. If the RNA transcript is the intermediate in moving the Ty-1 element, it is subject to splicing and the intron can be removed. Hence, these results fit the prediction of an RNA-mediated transposition. They demonstrate that during transposition, the flow of Ty-1 sequence information is from DNA to RNA to DNA.
Exercise 9.4
If yeast Ty-1 moved by the mechanism illustrated for DNA-mediated replicative transposition in Figure 9.13, what would be predicted in the experiment just outlined? Also, would you expect an increase in transposition when transcription is induced?
Additional Consequences of Transposition
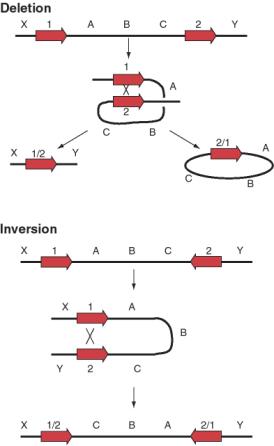
Not only can transposable elements interrupt genes or disrupt their regulation, but they can cause additional rearrangements in the genome. Homologous recombination can occur between any two nearly identical sequences. Thus when transposition makes a new copy of a transposable element, the two copies are now potential substrates for recombination. The outcome of recombination depends on the orientation of the two transposable elements relative to each other. Recombination between two transposable elements in the same orientation on the same chromosome leads to a deletion, whereas it results in an inversion if they are in opposite orientations (Figure 9.18).
The preference of the retrotransposition machinery for LINE1 mRNA does not appear to be absolute. Many processed genes have been found in eukaryotic genomes; these are genes that have no introns. In many cases, a homologous gene with introns is seen in the genome, so it appears that these processed genes have lost their introns. It is likely that these were formed when processed mRNA derived from the homologous gene with introns was copied into cDNA and reinserted into the genome. Many, but not all, of these processed genes are pseudogenes, i.e. they have been mutated such that they no longer encode proteins. Other examples of active processed genes have inserted next to promoters and encode functional proteins.


