4.6: Large Scale Genome Organization
- Page ID
- 314
How to get by with the smallest possible genome
The Mycoplasma species have the smallest genomes of any free-living species. They are most related to the Bacillaceae family, but have lost their cell walls and many other functions in a process of reductive evolution. They are obligate parasites, e.g. living in the lungs of humans. Their genomes encode many transport proteins, so that amino acids, sugars, etc. can be taken up from their hosts. They have very little metabolic capacity, utilizing only glycolysis in the case of M. genitalium. There is very little biosynthetic capacity, depending largely on uptake from the host for these nutrients.
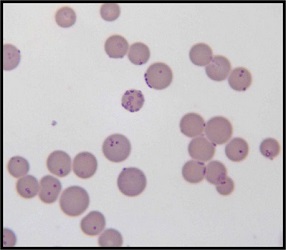
Mycoplasma Haemofelis, Wright-Geiemsa Staining 100X. (CC BY-SA 3.0; Nr387241).
One might have thought that the Mycoplasmal species would retain only the most highly conserved genes in bacteria, under the premise that these are the most critical genes. However, they have retained a proportion of conserved and variable genes that is quite similar to the proportion seen in E. coli. This indicates that these bacteria are maintaining a balance between conserved and variable genes that perhaps reflects an equilibrium between the stability of major physiological processes and the need for environmental adaptability.
More information from E. coli
The complete sequence of the E. coli genome provides an overview of genome structure within a well-understood context. For more information, see Blattner et al. (1997) Science, vol. 277, pp. 1453- 1462.
(1) Organization with respect to direction of replication
Since replication proceeds bidirectionally from the origin (oriC) and ends at the terminus, one can divide the genome into two "replicores." The replication fork proceeds clockwise in Replicore 1 and counter-clockwise in Replicore 2 (Figure 4.19).

Several features of the genome are oriented with respect to replication. All the rRNA genes, 53 of 86 tRNA genes, and 55% of the protein coding genes are transcribed in the same direction as the replication fork moves. In other species, such as the Mycoplasma, the transcriptional polarity is even more pronounced, and it also corresponds to the direction of replication.
These replicores show a pronounced skew in base composition, such that an excess of G over C is seen on the top strand (i.e. the one presented in the sequence file) in Replicore 1 and the opposite in Replicore 2. This nucleotide bias is striking and unexpected. As will be appreciated more after we study DNA synthesis in Part Two, this means that the leading strand for both replication forks is richer in G than C. Such an nucleotide bias may reflect differential mutation in the leading and lagging strands as a result of the asymmetry inherent in the DNA replication mechanism.
The recombination hotspot chi (GCTGGTGG) also shows a prominent strand preference, being more abundant on the leading strand of each replicore. The role of chi sites in recombination is covered in Chapter 8.
(2) Repeats, prophage and transposable elements
The E. coli chromosome contains several prophages and remnants of prophage, including lambda and three lambdoid prophages. The genome is peppered with at least 18 families of repeated DNA. The longest are the 5 Rhselements, which are 5.7 to 9.6 kb in length. Others are as short as the 581 copies of the 40 bp palindromic REP repeat. Several families of insertion sequences, which are transposable elements, are found. Note that repetitive elements are common in bacteria as well as in eukaryotes.
(3) General categories of genes
Many of the genes are similar to other genes in E. coli. Homologous genes that have diverged because of gene duplications are paralogous. The genes that encode proteins of similar but not necessarily identical function are referred to as a paralogous family. About 1/3 of the E. coli genes (1345) have at least one paralogous sequence in the genome. Some paralogous groups are quite large, the largest being the ABC transporters with 80 members. The larger number of genes in E. coli could reflect some redundancy in function as well as greater diversification of function compared to other bacteria with fewer genes.


Based on current understanding of the function of the gene products, about 1/4 are involved in small-molecule metabolism, about 1/8 are used in large-molecule metabolism, and at least 1/5 are associated with cell structure and processes. A specific function has not been assigned to the products of about 40% of the E. coli genes. Segmental duplications are common, as illustrated in Figure 4.21 for chromosomes 22.



