22.3: Molecules Derived from Amino Acids
- Page ID
- 15180
Introduction
Once made or ingested, amino acids have many metabolic fates. Of course, they are used for the synthesis of proteins. Aspartate and glutamate (and indirectly glutamine) can be converted to oxaloacetate and α-ketoglutarate, respectively, and used in the citric acid cycle for energy production. They can also be used for gluconeogenesis using mitochondrial and cytoplasmic enzymes. Branched-chain amino acids can be converted to acetyl-CoA and used in energy production or fat synthesis. A review summary of the use of amino acids in energy and biosynthetic metabolic pathways is shown in Figure \(\PageIndex{1}\).
Figure \(\PageIndex{1}\): Review summary of the use of amino acid in energy and biosynthetic metabolic pathways. Lieu, E.L., Nguyen, T., Rhyne, S., et al. Amino acids in cancer. Exp Mol Med 52, 15–30 (2020). https://doi.org/10.1038/s12276-020-0375-3. Creative Commons Attribution 4.0 International License, http://creativecommons.org/licenses/by/4.0/.
Amino acids are shown in green and other metabolites are in red. Orange represents transporters. Yellow boxes signify enzymes. Lesser known abbreviations for species include SHMT1 serine hydroxymethyltransferase, cytosolic, BCAT branched-chain amino acid transaminase, mitochondrial, BCAA branched-chain amino acid (valine, leucine, isoleucine), BCKA branched-chain ketoacid, GOT1 aspartate transaminase, cytosolic (AST), GLS glutaminase, GS glutamine synthetase (cytosolic and mitochondrial), ASNS asparagine synthetase, PRODH pyrroline-5-carboxylate dehydrogenase, PYCR pyrroline-5-carboxylate reductase, P5C pyrroline-5-carboxylate, GSH glutathione, PRPP phosphoribosyl pyrophosphate, LAT1 large-neutral amino acid transporter 1, SLC25A44 solute carrier family 25 member 44, GLUT glucose transporter,
Cancer cells from an increased need for fuels and biosynthetic intermediates. Both can come from amino acids as described previously. Glutamine is a key amino acid, especially if glucose is depleted as α-ketoglutarate (α-KG) and subsequently oxaloacetate (OAA generated from it powers the TCA cycle as fumarate, malate, and citrate are significantly increased. Hence it is both anaplerotic and a source of fuel. Similar increases in citrate occur in hypoxia. Aerobic glycolysis (Warburg effect) occurs in cancer cells, which show enhanced glucose uptake and conversion to lactate even in the presence of oxygen. This process can go so quickly that the amount of ATP produced in cancer cells from aerobic glycolysis can be similar to the from oxidative metabolism in the mitochondria, even though it is far less efficient. More information on cancer cell metabolism is found in Chapter 23.
In this chapter, we will discuss the conversion of amino acids to other molecules not directly involved in those metabolic pathways. We will focus on their use for the synthesis of polyamines, heme, and neurotransmitters in this chapter section. We won't discuss detailed mechanisms or structures for the proteins and enzymes involved in these pathways. In the next chapter section (22.4), we will present amino acids as substrates in the synthesis of pyrimidine and purine bases for nucleotides and nucleic acids.
Polyamine synthesis
If a non-quaternary amine has a single positive charge when protonated, a polyamine can have multiple positive charges. Hence they would be expected to bind to almost any negatively charged biomolecule, but especially those with multiple negative charges. These would include the polyanions RNA and DNA as well as proteins and lipid bilayers. They would then have the potential ability to regulate many features of cell life, including DNA replication and transcription, RNA translation, and a multitude of binding interactions. The question arises if these interactions are nonspecific, or specific, in which case they can be considered key regulators of cellular activity. Polyamine response elements have been found that regulate the transcription of genes including c-Myc and c-Jun. Polyamines have been shown tumor growth and aggressiveness.
The main biological polyamines include putrescine, spermine, and spermidine. Another is cadaverine. Given their names, you can surmise that they smell horrible. The synthesis of three polyamines from arginine and SAM is shown in Figure \(\PageIndex{2}\).
Figure \(\PageIndex{2}\): Polyamine synthesis form arginine and SAM
Glutathione synthesis and redox balance
Glutathione, γ-glutamylcysteinylglycine (GSH), is a chief regulator of the oxidation state of a cell. As a disulfide bond can be cleaved and hence reduced by the excess concentration of a thiol (sulfhydryl) like b-mercaptoethanol (which gets oxidized in the process), the free thiol on glutathione can act as a reducing agent in the cell. The production of reactive oxygen species (ROS) in normal but especially tumor cells, which have increased O2 demand and use, is countered by the generation of an antioxidant defense state. This is characterized in part by increased levels of reductants such as NADPH but especially glutathione. It can react with H2O2 through the enzyme GSH peroxidase to form water and the oxidized disulfide form of GSH, GSSG. The GSSG is oxide back to GSH by glutathione reductase (GR) and NADPH. Figure \(\PageIndex{3}\) shows the synthesis of glutathione from glutamate, cysteine, and glycine.
Figure \(\PageIndex{3}\): Synthesis of glutathione
NADPH is generated in the cell by the phosphopentose pathway metabolism of glucose and by malic enzyme. It can also be generated from the Ser-Glycine One Carbon Cycle (SGOT) that we saw in Chapter 18.4, which is shown again in Figure \(\PageIndex{4}\). Under appropriate conditions, this cycle can produce NADPH.
Figure \(\PageIndex{4}\): The Ser-Gly One Carbon (SGOC) Cycle
Serine Hydroxymethyltransferases (SHMTs) 2 is upregulated by HIF1α and helps maintain the NADPH/NADP+ ratio. Given the connection between the SGOC and the methionine cycle through folate, a decrease in serine concentration leads to a decrease in GSH.
Heme Biosynthesis
This section is derived from Aminat S. Ogun; Neena V. Joy; Menogh Valentine. https://www.ncbi.nlm.nih.gov/books/NBK537329/. Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/)
Heme is a macrocyclic tetrapyrrole ring structure containing two nonpolar vinyl groups on one edge and two charge propionates on the other. It is extensively conjugated with 26 π electrons (4n+2 = 4(6)+2) so it is aromatic. The molecule without Fe2+ is called protoporphyrin IX and with a centrally-coordinate Fe2+, it is called heme. The structures of both are shown in Figure \(\PageIndex{5}\).
Figure \(\PageIndex{5}\): Structure of protoporphyrin IX and heme
It is found in oxygen-binding proteins and as substrates and cofactors for enzymes involved in electron transport. It is synthesized in the bone marrow and liver. Alternative forms of heme include heme b (in hemoglobin), heme a (cytochrome a), and heme c (cytochrome c).
Its synthesis, as expected given its macrocyclic structure, is complicated. The key enzyme in the pathway for regulation is 5'-Aminolevulinic acid synthase (ALA-S). Liver and bone express ALAS2 while ALAS1 is expressed in all tissues. The synthesis starts in the mitochondria and ends in the cytosol. The overall pathway for heme synthesis is shown in Figure \(\PageIndex{6}\).
_pdftoPhotoshop.png?revision=1)
Figure \(\PageIndex{6}\): Heme biosynthetic pathway. Wikimedia Commonsile: Heme-Synthesis-Chemical-Details-Mirror.svg
5'-Aminolevulinic acid synthase (ALA-S), a pyridoxal phosphate-dependent enzyme, catalyzes the rate-limiting step in heme synthesis in the liver and erythroid cells. It is highly regulated There are two forms of ALA Synthase, ALAS1, and ALAS2. All cells express ALAS1 while only the liver and bone marrow expresses ALAS2. The gene for ALAS2 is on the X-chromosome. After the synthesis of ALA in the mitochondria, it moves into the cytoplasm for the remaining steps.
Figure \(\PageIndex{7}\) shows a likely mechanism for the first committed step, the production of ALA. This enzyme is used in the synthesis of all tetrapyrroles, including heme, chlorophyll, and cobalamin.
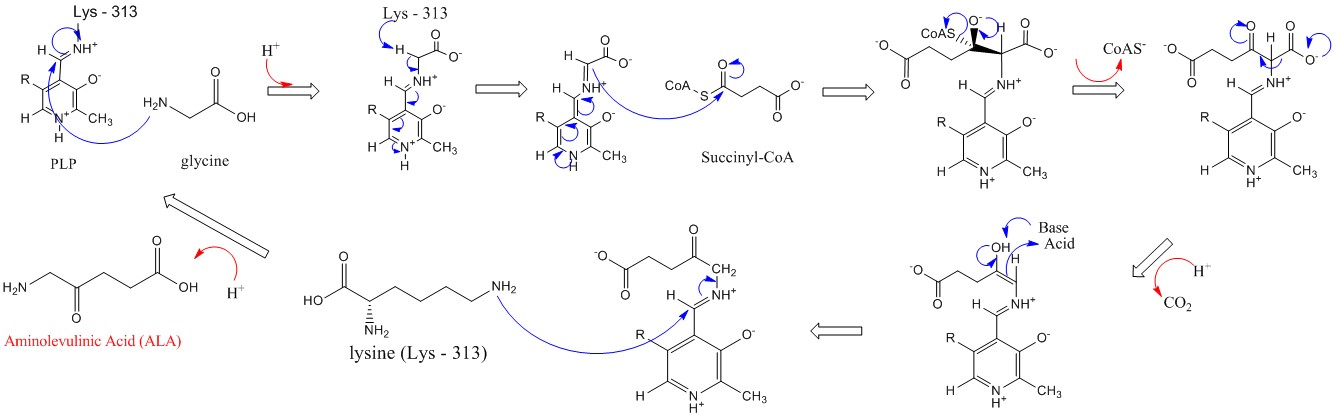 Figure \(\PageIndex{7}\): Mechanism for 5'-Aminolevulinic acid synthesis by ALAS (Wikipedia. https://en.Wikipedia.org/wiki/Aminol..._acid_synthase)
Figure \(\PageIndex{7}\): Mechanism for 5'-Aminolevulinic acid synthesis by ALAS (Wikipedia. https://en.Wikipedia.org/wiki/Aminol..._acid_synthase)
The pathway shown in Figure 6 above is called the C4 pathway and is found in mammals, fungi, and purple nonsulfur bacteria. A C5 pathway is found in most bacteria, all archaea, and plants. The biosynthetic pathway for heme synthesis in E. Coli is shown in Figure \(\PageIndex{8}\).
Figure \(\PageIndex{8}\): Heme pathway in E. coli. Zhang, J., Kang, Z., Chen, J. et al. Optimization of the heme biosynthesis pathway for the production of 5-aminolevulinic acid in Escherichia coli. Sci Rep 5, 8584 (2015). https://doi.org/10.1038/srep08584. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/
The pathway is divided into three modules (module I, module II, and module III in the dotted box). The arrows in green and red represent the enzymes that are positive and negative to ALA accumulation, respectively. Dotted red arrows represent the feedback inhibition. α-KG: α-ketoglutarate, GSA: glutamate-1-semialdehyde, ALA: 5-aminolevulinic acid, PBG: porphobilinogen, HMB: hydroxymethylbilane, GltX: glutamyl-tRNA synthetase, HemA: glutamyl-tRNA reductase, HemL: glutamate-1-semialdehyde aminotransferase, HemB: 5-aminolevulinic acid dehydratase, HemC: porphobilinogen deaminase, HemD: uroporphyrinogen III synthase, HemE: uroporphyrinogen decarboxylase, HemF: coproporphyrinogen III oxidase, HemG: protoporphyrin oxidase, HemH: ferrochelatase.
In immature red blood cells (reticulocytes), heme increase globin protein synthesis. The hormone erythropoietin increases heme synthesis. In the liver, heme is part of cytochrome P450s. Increased concentration of drugs causes increases in ALAS1 to produce the cytochrome P450s to metabolize them. Also, low heme concentration increases ALAS1 transcription. Mutations in ALAS2 can lead to X-linked sideroblastic anemia from decreased heme production even as Fe2+ continues to enter the cell.
Yeast ALAS is a homodimer with PLP covalently attached through a Schiff base link to lysine 337 of just one of the monomers. The structures of a noncovalent complex of PLP with ALAS (pdb 5TXR) and the covalently bound one (5TXT) show large changes in the protein conformation. PLP when covalently attached reorders the active. A C-terminal extension not found in bacteria wraps around the dimer and binds near the active site and is important for activity. Mutations in the tail can result in human diseases.
Figure \(\PageIndex{9}\) shows an interactive iCn3D model of the 5-aminolevulinic acid synthase with covalently attached PLP (5TXT).
Figure \(\PageIndex{9}\): 5-aminolevulinic acid synthase with covalently attached PLP (5TXT). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?fTBbWuS3HPTP8uRm9
Lysine 337 (spacefill, CPK colors, labeled) in the A chain (magenta, no bound PLP) is shown. Lys 337 in the B chain (cyan) is covalently linked to PLP. The side chain of lysine 337 covalently attached to PLP is shown in spacefill, CPK colors, and labeled. The C-terminal extension (493–548) is shown as a red backbone chain. The very distal end of the extension is disordered and missing in the B chain (cyan).
Figure \(\PageIndex{10}\) shows an interactive iCn3D model of aligned 5-aminolevulinic acid synthase with free PLP (not covalently attached, 5TXR) and with covalently attached PLP (5TXT ).
Figure \(\PageIndex{11}\): Alignment of 5-aminolevulinic acid synthase with free PLP (5TXR) and with and with covalently attached PLP (5TXT ). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...aqN4Qpi2pSLkD7
The 5TXT structure contains two molecules of a stabilizing molecule shown in stick form, which you can ignore. The A chain is shown in magenta and the B chain is in cyan. Press "a" to toggle back and forth between the structures. The C-terminal extension is missing from the figure.
Figure \(\PageIndex{11}\) shows another view of heme synthesis which emphasizes the role of mitochondrial and cytoplasmic enzymes.
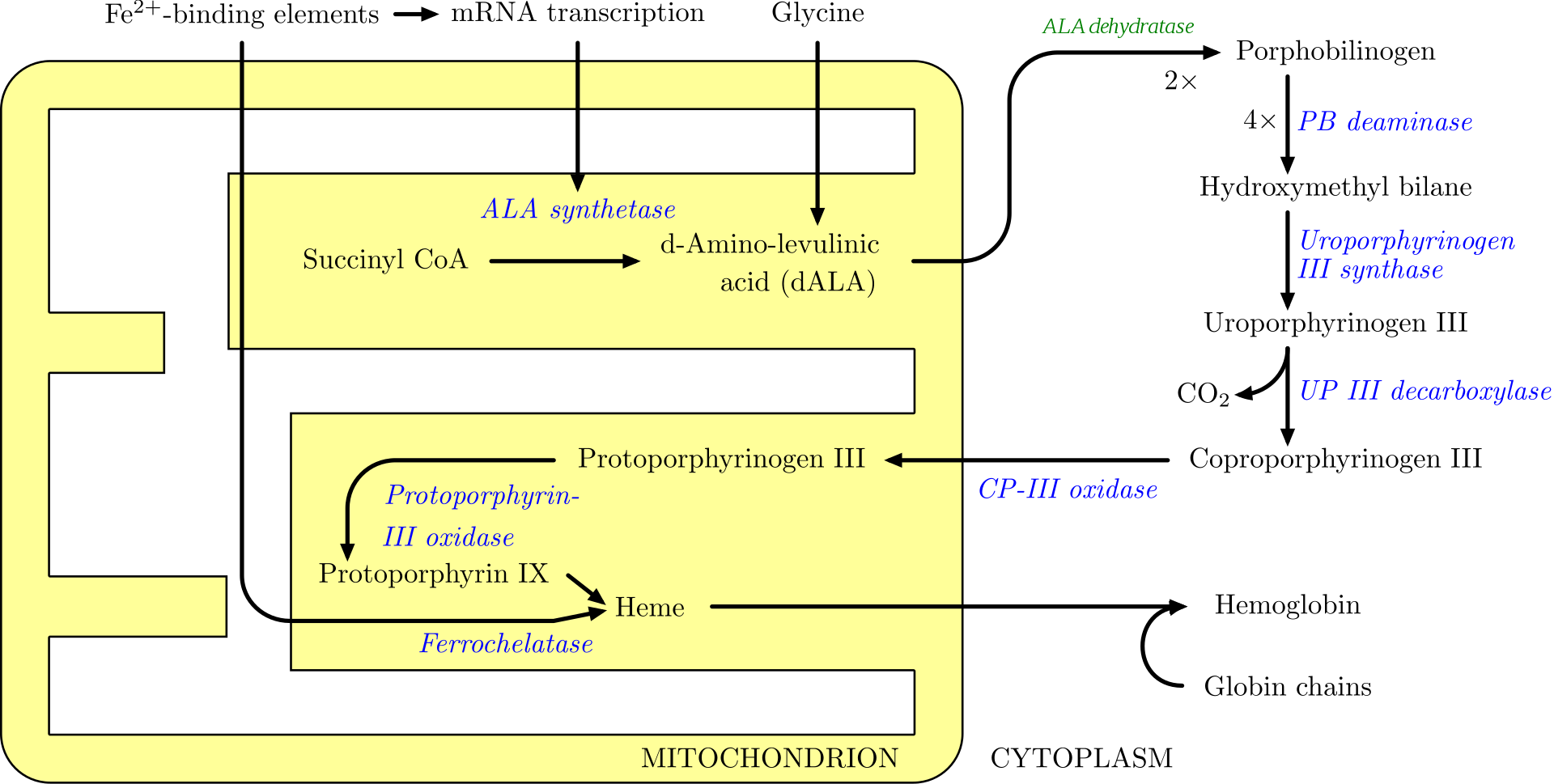
Figure \(\PageIndex{11}\): Mitochondrial and cytosolic contributions to heme synthesis. https://commons.wikimedia.org/wiki/F..._synthesis.png
Neurotransmitters
The section below is modified from Manorama Patri. Synaptic Transmission and Amino Acid Neurotransmitters. DOI: 10.5772/intechopen.82121. https://www.intechopen.com/books/neu...rotransmitters. Creative Commons Attribution 3.0 License,
There are three major categories of amino acids and their derivatives act as neurotransmitters are:
- Amino acids: The neurotransmitters of this group are involved in fast synaptic transmission and are inhibitory and excitatory in action (primarily glutamic acid, GABA, aspartic acid, and glycine).
- Amines: Amines are modified amino acids such as biogenic amines, e.g., catecholamines. The neurotransmitters of this group involve in slow synaptic transmission and are inhibitory and excitatory in action (noradrenaline, adrenaline, dopamine, serotonin, and histamine).
- Others: The ones which do not fit in any of these categories (acetylcholine and nitric oxide). Amino acids are among the most abundant of all neurotransmitters present within the central nervous system (CNS).
Amino acid transmitters provide the majority of excitatory and inhibitory neurotransmission in the nervous system. Amino acids used for synaptic transmission are compartmentalized (e.g., glutamate, compartmentalized from metabolic glutamate used for protein synthesis by packaging the transmitter into synaptic vesicles for subsequent Ca2+-dependent release). Amino acid neurotransmitters are all products of intermediary metabolism except GABA. Unlike all the other amino acid neurotransmitters, GABA is not used in protein synthesis and is produced by an enzyme (glutamic acid decarboxylase; GAD) uniquely located in neurons.
Here is some more specific information:
- Glutamate: Glutamate is used at the great majority of fast excitatory synapses in the brain and spinal cord. Glutamate binds to glutamate receptors of which there are many subtypes based on other molecules (some amino acid derivatives) that can bind to them. These other molecules include N-methyl-D-aspartate (NMDA), α-amino-3-hydroxy-5-methyl-4-isoxazolepropionate (AMPA) kainate, and quisqualate.
- Aspartate: Aspartate is the most abundant excitatory neurotransmitter in the CNS. Like glycine, aspartate is primarily localized to the ventral spinal cord. Note that the two major excitatory neurotransmitters both have carboxylic acid side chains.
- Gamma-aminobutyric acid (GABA): GABA, which is not one of the canonical amino acids used in protein biosynthesis, is the most ubiquitous inhibitory neurotransmitter in the brain.
- Glycine: lycine receptors are ligand-gated ion channels that increase Cl− influx and hence are generally inhibitory. Hydroxymethyl transferase converts the amino acid serine to glycine. Glycine has been found to play a role in the functional modulation of NMDA receptors
The pathways for the synthesis of amino acid-derived bioactive amines and neurotransmitters are shown in Figure \(\PageIndex{12}\).
Figure \(\PageIndex{12}\): Pathways for the synthesis of amino acid-derived bioactive amines and neurotransmitters
Note the structural similarity of the psychotropic and hallucinogenic drug LSD to serotonin (5HT), amphetamines to norepinephrine and epinephrine, and melatonin (a substance some take as a sleeping aid and which forms in the dark at night in brains. The name catecholamines derive from the common name of the 1,2-dihydroxybenzene group (catechol).
The first and rate-limiting step in catecholamine synthesis is catalyzed by tyrosine hydroxylase (TH). It has no heme but it has an Fe2+ and tetrahydrobiopterin as a cofactor used in the synthesis of dihydroxyphenylalanine (DOPA). Tyrosine hydroxylase is rate-limiting for the synthesis of all three transmitters.
The enzyme is inhibited by catecholamines including dopamine, a downstream product, and is activated by phosphorylation on serine 40. The structures of TH in the absence of dopamine and the pSer40 state are known. The protein is a tetramer with a regulatory domain (dimer) and catalytic domain (also a dimer) separated by 15 Å.
The mammalian TH is a member of the aromatic amino acid hydroxylases (AAAHs) which are mainly found as homotetramers. Each subunit has 3 domains:
- The N-terminal regulatory domain (RD) that has an unstructured variable-length section followed by an ACT (aspartate kinase-chorismate mutase-TyrA) domain. The N-terminal tail contains serine 40 which on phosphorylation relieves the inhibition when dopamine is bound to the catalytic domain.
- a central catalytic domain (CD) that binds Fe2+, aromatic amino acid substrates, and the tetrahydrobiopterin cofactor
- C-terminal oligomerization domain (OD) which leads to dimer and tetramer formation.
Figure \(\PageIndex{13}\) shows a potential model that illustrates dopamine (DA)-mediated feedback inhibition and its regulation by serine 40 phosphorylation through the interaction of the N-terminal tail of the regulatory domain (RD) with the catalytic domain. All forms containing bound dopamine (yellow star) are inactive.
Figure \(\PageIndex{13}\): Cartoon model of DA-mediated feedback inhibition and its regulation by S40 phosphorylation. Bueno-Carrasco, M.T., Cuéllar, J., Flydal, M.I. et al. Structural mechanism for tyrosine hydroxylase inhibition by dopamine and reactivation by Ser40 phosphorylation. Nat Commun 13, 74 (2022). https://doi.org/10.1038/s41467-021-27657-y. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/.
In the active, apo, and non-phospho states, the 39−58 α-helix of the N-terminal regulatory domain of TH is detached from the main structure (I, apo-TH). The feedback inhibitor DA binds to the TH active site, most likely in the open conformation (I′, TH(DA)). DA-binding favors the interaction of the N-terminal α-helix with the same binding site, which blocks DA exit and contributes to the high-affinity binding and strong inhibition of TH activity (II, TH(DA)). Protein Kinase (PK) phosphorylation of S40 in TH(DA), leads to state III (THS40p(DA)), prompting the detachment of the α-helix from the TH active site (IV′), which opens up for DA-dissociation and activation (IV, THS40p). PKs and protein phosphatase(s) (PP) control the transition between THS40p and unphosphorylated TH for both DA bound (I′ ↔ IV′ and II ↔ III) and apo-TH (I ↔ IV). States I′ and III are expected to be only transiently populated during DA binding as states II and IV' will be more stable. Hence states I′ and III states are faded. S40 is also expected to be less accessible in state II than in state I, which is indicated by stippled lines for phosphorylation of TH in state II. The case for dephosphorylation is not known, but it could be expected that state III is a poorer substrate for PP than the open states IV′ and IV. The dephosphorylation reaction III → II is therefore also stippled. The states where we provide structural details in this work (I, II, and IV) are marked with circles.
Figure \(\PageIndex{14}\) shows a model of the TH active site changes on phosphorylation of serine 40 using structural and molecular dynamics approaches that led to the cartoon model above.
Figure \(\PageIndex{14}\): Modeling of the TH active site.
Panel (a) shows models demonstrating the effect of serine 40 phosphorylation on the interaction of the N-terminal α-helix with bound dopamine (DA). Representative conformations from the last 50 ns of a 500 ns MD simulations for TH(DA) (grey ribbon) and pS40-TH(DA) (light blue ribbon) are shown. The resulting structures show a slight shift of the N-terminal α-helix upon phosphorylation, most probably due to electrostatic repulsion between the phosphate and E325, E375, and D424.
Panel (b) shows a detailed view of the atomic model of the TH(DA) active site. (left) The N-terminal α-helix (orange), establishes connections with the adjacent helix D360-E375 and with residues of the 290–297 and 420–429 loops (blue, right).
Pane (c) shows a cartoon depicting the interactions established between residues of the N-terminal α-helix that enters the active site, and residues of adjacent regions.
Figure \(\PageIndex{15\) below shows an interactive iCn3D model of the full-length tyrosine hydroxylase in complex with dopamine (residues 40-497) in which the regulatory domain (residues 40-165) has been included only with the backbone atoms (6zvp)
Figure \(\PageIndex{15}\): Tyrosine hydroxylase dopamine complex (6zvp). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...hu6vJn22eDkWw7
Finally, Figure \(\PageIndex{16}\) shows an interactive iCn3D model of the active site of tyrosine hydroxylase in complex with dopamine (6zvp)
Figure \(\PageIndex{16}\): Active site of tyrosine hydroxylase dopamine complex (6zvp). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?sx5obQnzojmKXuLS9
The side chains binding the active site Fe2+ and the interaction of Fe2+ with dopamine (LDP) are shown in sticks and labeled. The oxygen of serine 40 is shown as a red sphere. Dopamine is the molecule containing the 1,2-dihydroxybenzene.
Figure \(\PageIndex{17}\) shows a final summary presentation of the conversion of phenylalanine, tyrosine, and tryptophan to neurotransmitters.
Figure \(\PageIndex{17}\): Comparison of monoamine synthesis pathways. Adapted from Hochman, Shawn. (2015). Neural Regeneration Research. 10. 10.4103/1673-5374.169625. Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International




.png?revision=1&size=bestfit&width=470&height=342)
_and_with_and_with_covalently_attached_PLP_(5TXT_).png?revision=1&size=bestfit&width=285&height=228)
.png?revision=1&size=bestfit&width=357&height=337)
.png?revision=1&size=bestfit&width=307&height=319)