13.4: Pentose Phosphate Pathway of Glucose Oxidation
- Page ID
- 15008
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Introduction
The pentose phosphate pathway (PPP), also known as the pentose phosphate shunt, is an important part of glucose metabolism. The PPP branches after the first step of glycolysis and consumes the intermediate glucose 6-phosphate (G6P) to generate fructose 6-phosphate (F6P) and glyceraldehyde 3-phosphate (G3P) through the oxidative and non-oxidative branches of the PPP. Unlike glycolysis and glucose aerobic oxidation, the PPP does not provide adenosine 5′-triphosphate (ATP) to meet the energy demands of cells. Instead, it supplies NADPH and ribose 5-phosphate (R5P). These two metabolites are vital for the survival and proliferation of cells. R5P is a building block for nucleic acid synthesis. NADPH is the reducing power required for the synthesis of fatty acids, sterols, nucleotides, and non-essential amino acids. Moreover, NADPH-derived conversion of oxidized glutathione (GSSG) to reduced glutathione (GSH) via glutathione reductase is important for cellular antioxidant defenses. Interestingly, NADPH also serves as the substrate of NADPH oxidases (NOXs) which produce reactive oxygen species (ROS).
Both the oxidative branch and non-oxidative branch of the PPP take place in the cytosol (Figure \(\PageIndex{1}\)). Glucose 6-phosphate dehydrogenase (G6PD) is the rate-limiting enzyme of the oxidative PPP, determining the flux of G6P directed into the pathway. G6PD catalyzes the conversion of G6P to 6-phosphogluconolactone, accompanied by NADPH production. 6-phosphogluconolactonase (6PGL) is the enzyme that hydrolyzes 6-phosphogluconolactone to produce 6-phosphogluconate (6PG). 6-phosphogluconate dehydrogenase (6PGD) converts 6-PG to ribulose 5-phosphate (Ru5P) and generates NAPDH (Figure \(\PageIndex{1}\)). The largest contributor to cytosolic NADPH is the oxidative PPP in mammalian cells.
The non-oxidative branch is composed of a series of reversible transfer reactions of chemical groups. Ribose 5-phosphate isomerase (RPI) and ribulose 5-phosphate epimerase (RPE) catalyze reversible reactions converting Ru5P to R5P and xylulose 5-phosphate (Xu5P), respectively. TKT catalyzes two reversible reactions. One is the conversion of Xu5P and R5P to G3P and sedoheptulose 7-phosphate (S7P). The other is the conversion of Xu5P and erythrose 4-phosphate (E4P) to G3P and F6P. Therefore, TKT can bi-directionally regulate the carbon flux between the non-oxidative PPP and glycolysis or gluconeogenesis. Transaldolase (TALDO) reversibly converts G3P and S7P to E4P and F6P. The non-oxidative branch not only replenishes metabolites of the oxidative branch (by their reversal), but also regulates the flux of glycolysis or gluconeogenesis by providing F6P and G3P (Figure \(\PageIndex{1}\)).
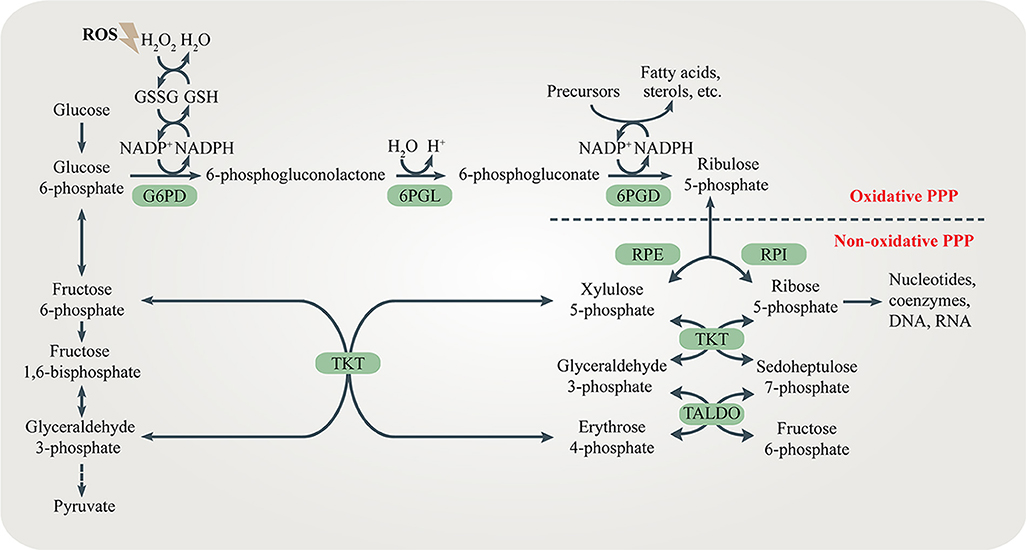
Figure \(\PageIndex{1}\): The pentose phosphate pathway (PPP). The PPP branches after the first step of glycolysis and goes back to fructose 6-phosphate and glyceraldehyde 3-phosphate in the glycolytic and gluconeogenic pathway. The PPP produces R5P and NADPH for biosynthesis and redox regulation. Enzymes in the oxidative and non-oxidative PPP are shaded in green. Figure from: Ge, T., et al. (2020) Cellular Endocrinology DOI:10.3389
Oxidative branch
The oxidative branch of PPP (ox-PPP) is a non-reversible metabolic pathway where glucose-6-phosphate (G6P) is transformed into 6-phosphoglucono-δ-lactone by glucose-6-phosphate dehydrogenase (G6PD) and, subsequently, to ribulose-5-phosphate by 6-phosphogluconate dehydrogenase (6PGD) with the concomitant production of nicotinamide adenine dinucleotide phosphate (NADPH). The resulting ribulose-5-phosphate is then converted to ribose-5-phosphate and used for the biosynthesis of nucleotides (Figure\(\PageIndex{2}\)).
Figure \(\PageIndex{2}\): Summary of the oxidative branch of phosphopentose pathway
The first enzyme in the oxidative branch is the glucose-6-phosphate dehydrogenase enzyme. It is also the first committed enzyme in the pathway and is involved in the regulation. PPP metabolizes from 5 to 30% of glucose depending on the tissue type. The enzymatic reaction is:
1. Glucose-6-phosphate dehydrogenase: Glc6P + NADP+ → 6-phosphogluconolactone + NADPH
The oxidative branch produces NADPH for reductive biosynthesis and it also maintains the reducing condition of the cell to protect it against oxidative stress, which is especially important in erythrocytes (Figure\(\PageIndex{3}\)). It also starts the pathway to produce 5-carbon sugars for nucleotide biosynthesis.
G6PDH is very important for the protection of free radicals in red blood cells since they don't have mitochondria that could provide another source of NADPH for protection. NADPH is involved in protection against ROS through the three cycles shown in Figure \(\PageIndex{3}\).
Figure \(\PageIndex{3}\): Function of G6PD enzyme in the PPP from red blood cells. In G6PD-normal red cells, the NADPH is produced by the action of glucose 6-phosphate dehydrogenase (G6PD) and 6-phosphogluconate dehydrogenase (6PGD) enzymes. The NADPH serves as a proton donor to regenerate the GSSG oxidized. Cat = Catalase; GPx = Glutathione peroxidase; GR = Glutathione reductase; G6PD = glucose 6-phosphate dehydrogenase; 6PGL = 6-phosphogluconolactonase; 6GPD = 6-phosphogluconate dehydrogenase; SOD = Superoxide dismutase; GSH = Reduced glutathione; GSSG = Oxidized glutathione; H2O2 = Peroxide; O2− = Superoxide. Gomez-Manzo et al. Int J Mol Sci. 2016 Dec; 17(12): 2069. doi: 10.3390/ijms17122069. Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
NADPH can keep glutathione in its reduced form, which as a substrate for catalase can help rid the cell of hydrogen peroxide and indirectly other ROS.
The enzyme is active as a dimer or tetramer. Each monomer in the complex has a substrate binding site that binds to G6P, and a catalytic coenzyme binding site that binds to NADP+/NADPH. For some higher organisms, such as humans, G6PD contains an additional NADP+ binding site, called the NADP+ structural site, that does not seem to participate directly in the reaction catalyzed by G6PD. The evolutionary purpose of the NADP+ structural site is unknown, however, it does play a role in the overall stability of the enzyme.
Mutations that cause dysfunction or deficiencies are very common, especially in males and in Africa, Asia, the Mediterranean, and the Middle East, in a geographic distribution that parallels the incidence of malaria. Interestingly, many of the mutations occur near the NADP+ structural site. Glucose-6-phosphate dehydrogenase deficiency is very common worldwide and causes acute hemolytic anemia in the presence of simple infection, ingestion of fava beans, or reaction with certain medicines.
An abbreviate mechanism for glucose-6-phosphate dehydrogenase from Leuconostoc mesenteroides is shown in Figure \(\PageIndex{4}\).
Figure \(\PageIndex{4}\): Mechanism of Leuconostoc mesenteroides glucose-6-phosphate dehydrogenase. Ribeiro AJM et al. (2017), Nucleic Acids Res, 46, D618-D623. Mechanism and Catalytic Site Atlas (M-CSA): a database of enzyme reaction mechanisms and active sites. DOI:10.1093/nar/gkx1012. PMID:29106569 https://www.ebi.ac.uk/thornton-srv/m-csa/entry/843/. Creative Commons Attribution 4.0 International (CC BY 4.0) License.
Figure \(\PageIndex{5}\) shows an interactive iCn3D model of the glucose 6-phosphate dehydrogenase from Leuconostoc mesenteroides (1DPG).
Figure \(\PageIndex{5}\): Glucose 6-phosphate dehydrogenase from Leuconostoc mesenteroides (1DPG). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...m8NCXiBuQbTQb7
Interestingly, this bacteria has an incomplete glycolytic pathway and can use this enzyme with either NADP+ for anabolism or NAD+ for catabolism.
Each monomer can bind two NADP+, one at a site that promotes structure integrity and stability and the other at the catalytic site where NADP+ serves as a substrate (or cofactor). The other substrate, glucose-6-phosphate bind between the two.
Figure \(\PageIndex{6}\) shows an interactive iCn3D model of the human glucose 6-phosphate dehydrogenase with bound structural and substrate NADP+ (2BH9).
Figure \(\PageIndex{6}\): Human glucose 6-phosphate dehydrogenase with bound structural and substrate NADP (2BH9).(Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?NFJnS7UtUt3Uwu2D7
The biological unit shown is a dimer. The structural NADP+ is shown in spacefill bound in the N-terminal end of each monomer, while the substrate (cofactor) NADP+ is shown in sticks. Arg 459 is shown in spacefill.
A common disease-causing mutation of G6PDH is the Canton R459L mutation. It's common in China and Southeast Asia. The activity of the enzyme is significantly decreased and the enzyme is less able to form tetramers. It unfolds at a lower temperature. This suggests that the mutation causes a significant conformation change. The Arg459 is shown in the above model. Its contribution to interhelical noncovalent attractions between it and D181 and N185. D181 is shown in proximity to Arg459 in the above model.
Figure \(\PageIndex{7}\)s shows a static image of glucose 6-phosphate dehydrogenase with bound structural and substrate NADP+ as well as glucose-6-phosphate.
Figure \(\PageIndex{7}\): Glucose 6-phosphate dehydrogenase with bound structural and substrate NADP+ as well as glucose-6-phosphate (2BHL and 2BH9). Structural NADP+ (blue molecular surface), catalytic NADP+ (dark purple molecular surface), and G6P substrate (yellow molecular surface) in the dimer are shown. The two monomers are shown in cyan and green. Right inset, close-up of the dimer interface and both structural NADP+ molecules. Gomez-Manzo et al., ibid.
The second reaction of the oxidative branch is mediated by the phosphogluconolactonase (6PGL, PGLS) enzyme. The overall reaction is shown here:
2. 6-phosphogluconolactamase: 6-phosphogluconolactone + H2O → 6-phosphogluconate
6PGL is a cytosolic enzyme found in all organisms that catalyzes the hydrolysis of 6-phosphogluconolactone to 6-phosphogluconic acid in the oxidative phase of the pentose phosphate pathway. 6PGL hydrolysis of 6-phosphogluconolactone to 6-phosphogluconic acid has been proposed to proceed via proton transfer to the O5 ring oxygen atom (Figure \(\PageIndex{8}\)). The reaction initiates via the attack of a hydroxide ion at the C5 ester. A tetrahedral intermediate forms and the elimination of the ester linkage follows, aided by the donation of a proton from an active site histidine residue.
Figure \(\PageIndex{8}\): Mechanism for6-phosphogluconolactamaseHis 165 and Asp 163 appear to be involved in a proton relay scheme. His 163 is conserved in lactonase. Arg 77 and 200 are involved in binding the substrate.
Figure \(\PageIndex{9}\) shows an interactive iCn3D model of the 6-phosphogluconolactonase from Trypanosoma brucei complexed with 6-phosphogluconic acid (3E7F). This enzyme is a target for developing drug treatment strategies for African sleeping sickness.
Figure \(\PageIndex{9}\): 6-phosphogluconolactonase from Trypanosoma brucei complexed with 6-phosphogluconic acid (3E7F). The key residues involved in binding and the charge relay system are shown in sticks and labeled. The product, 6-phosphogluconic acid, is shown in spacefill. The single sphere is a Zn2+ which does not appear to be involved in catalysis. (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?jRgi1nfrSGGgTYWo8
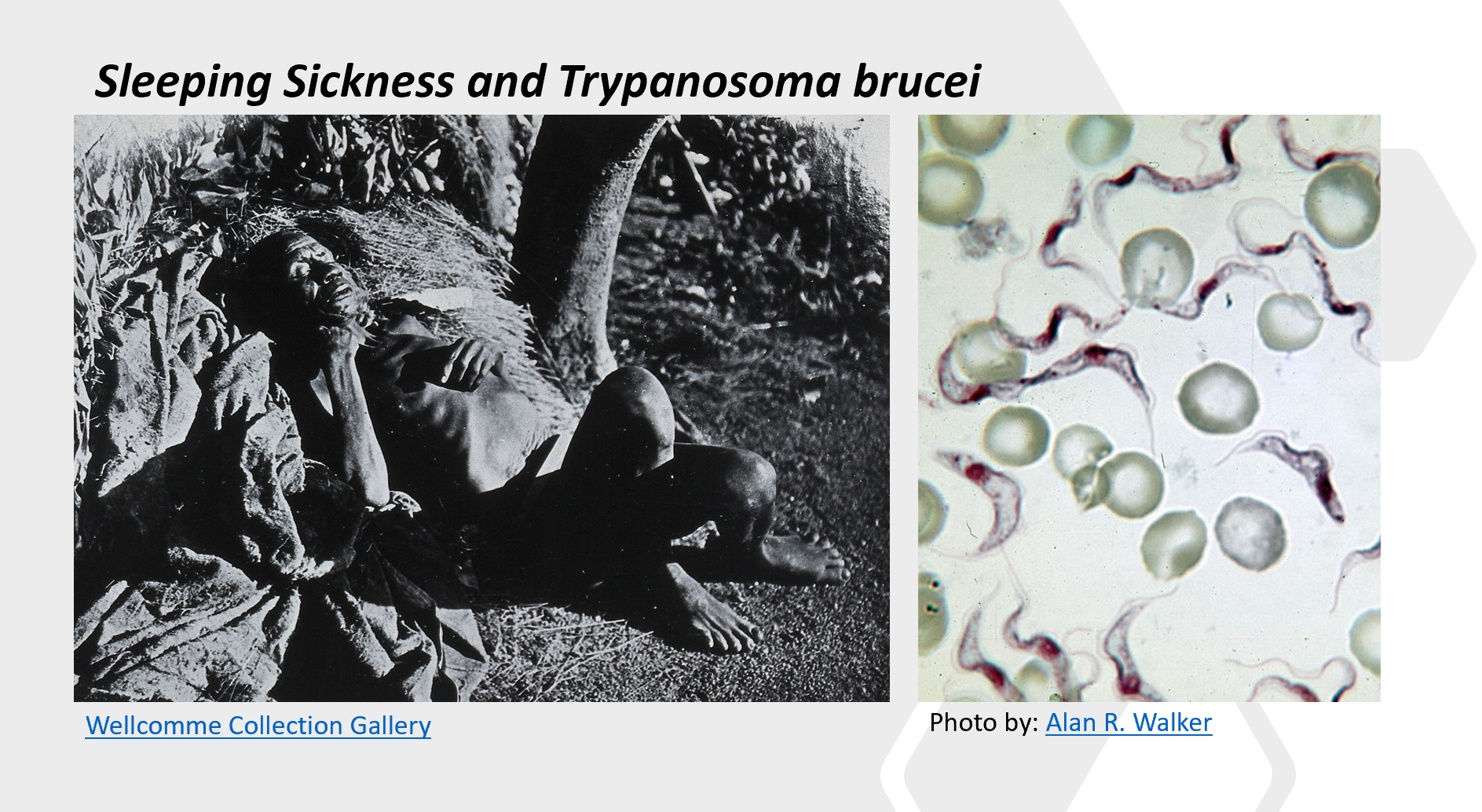
Trypanosomiasis is a disease usually referring to African human trypanosomiasis or African sleeping sickness. This infectious disease is caused by the parasites Trypanosoma brucei gambiense or Trypanosoma brucei rhodesiense cause this infectious disease, and the tsetse fly transmits the disease.
The vector tsetse fly, Glossina, carries the trypanosome within the midgut after a blood meal. These protozoa then migrate to the salivary glands of the fly whereby they can be transmitted during the next feeding. After inoculation within the host, the parasite can live freely within the bloodstream and evade mammalian host defenses through variable surface glycoproteins (VSG). The clinical disease has 2 stages. These are characterized by an early/first hemolymphatic stage and a late /second meningoencephalitis stage with an invasion of the central nervous system (CNS). In stage 1, systemic symptoms develop including intermittent fever, headache, pruritus, and lymphadenopathy. Undulating fevers reflect parasites multiplying within the blood. Less frequent hepatosplenomegaly may occur in the early stage. In the late/second stage, CNS symptoms manifest as sleep disturbances or neuropsychiatric disorders. A sleep disorder is the most common symptom of the second stage, and it is from this that the term “African sleeping sickness” was ascribed.
New Drug Targets:
Enzymes involved in the pentose phosphate pathway provide a new set of drug targets to help combat parasitic infections such as those caused by trypanosomes. In this context, a focus for new therapeutics has been on the disruption of glycosomes within these organisms. Glyoxosomes are specialized peroxisomes found in plants and some fungi. They all have enzymes for the glyoxylate shunt and can hence create some glucogenic intermediates. In trypanosomes and Leishmania, another internal organelle called the glycosome is found. This houses most of the glycolytic enzymes and also the oxidative enzymes of the pentose phosphate pathway. This makes the enzymes in the glycosome unique drug targets.
The third enzyme utilized in the oxidative branch of the PPP is 6-phosphogluconate dehydrogenase (6PGDH). The overall reaction is shown here:
3. 6-phosphogluconate dehydrogenase: 6-phosphogluconate + NADP+ ↔ ribulose-5-phosphate + NADPH + CO2
6PGDH catalyzes a reversible oxidative decarboxylation reaction, as shown in Figure \(\PageIndex{10}\). Oxidation, followed by decarboxylation forms an endiol intermediate followed by conversion to product. Notably, NADP+ serves as an electron acceptor in the reaction, leading to the production of a second molecule of NADPH. Essentially, an active center lysine abstracts a proton from the substrate, 6-phosphogluconate, and NADP+ is reduced, causing the formation of a ketone-intermediate. Decarboxylation leads to the formation of the enediol intermediate. The active site lysine abstracts a proton in the second half of the reaction causing the formation of ribulose 5-phosphate. The abstracted proton is transferred to the active site Glu190 resetting the enzyme for another catalytic cycle.
Figure \(\PageIndex{10}\): Mechanism of 6-phosphogluconate dehydrogenase.
Figure \(\PageIndex{11}\) shows an interactive iCn3D model of sheep 6-phosphogluconate dehydrogenase (2PGD).
Figure \(\PageIndex{11}\): Sheep 6-phosphogluconate dehydrogenase (2PGD). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...DqosWmFg1DqK16
The protein is a homodimer in which the monomers act independently: each contains a large, mainly alpha-helical domain and a smaller beta-alpha-beta domain, containing a mixed parallel and anti-parallel 6-stranded beta-sheet. NADP+ is bound in a cleft in the small domain, the substrate binding in an adjacent pocket.
Nonoxidative branch
The non-oxidative branch of the pentose phosphate pathway (nonox-PPP) is responsible for the formation of simple sugars within the cell and maintaining these building blocks in appropriate concentrations. The enzymes utilized in this pathway mediate a set of reversible reactions that lead to the production of ribose-5-phosphate, xylulose-5-phosphate, sedoheptulose 7-phosphate, and erythrose 4-phosphate, as well as intermediates utilized in the glycolytic pathway, including glyceraldehyde-3-phosphate and fructose-6-phosphate Major enzymes utilized include transketolase (TKT) and transaldolase (TALDO), as well as important isomerase and epimerase enzymes.
Ribose 5-phosphate is an important building block for the biosynthesis of nucleotides, and erythrose 4-phosphate is used in the synthesis of aromatic amino acids. When these simple sugars are in excess, they can also be converted into glycolytic intermediates and utilized for energy production.
The nonoxidative branch of the phosphopentose pathway is shown in Figure \(\PageIndex{12}\).
Figure \(\PageIndex{12}\) :Nonoxidative branch of the phosphopentose pathway
Now let's examine the individual enzymes. Ribulose 5-phosphate is the starting place for the non-oxidative portion of the PPP. It can be converted down two different pathways, either to ribose-5-phosphate or to xylulose-5-phosphate. The ribose 5-phosphate isomerase is involved in the ketose-aldose conversion and will be the first enzyme discussed.
Ribose-5-phosphate isomerase (Rpi)
This enzyme is fully reversible and is usually referred to as ribose-5-phosphate isomerase, although it can also be known as ribulose 5-phosphate isomerase, as it mediates the transition between this aldose-ketose pair. In addition to the PPP, this enzymatic reaction is also required in the Calvin cycle in photosynthesis.
There are two different Rpi enzymes, RpiA and RpiB, which have little sequence or structural similarities as well as different mechanisms. RpiA is found in all three kingdoms of life and is highly conserved due to its role in the PPP and the Calvin Cycle of photosynthesis. RpiB, on the other hand, is only found in some bacteria and protozoans. Thus, RpiB is a potential therapeutic target for the treatment of diseases such as African Sleeping Sickness, Chagas disease, and leishmaniasis. While RpiA and RpiB are structurally very different, both enzymes catalyze the isomerization reaction through an enediol intermediate using the linear form of the sugar. The reaction mechanism of RpiB is detailed below.
Figure \(\PageIndex{13}\) shows a mechanism for Escherichia coli RpiB.
Figure \(\PageIndex{13}\): Mechanism for Escherichia coli RpiB. https://www.ebi.ac.uk/thornton-srv/m-csa/entry/680/. Creative Commons Attribution 4.0 International (CC BY 4.0) License.
Ribulose-5-phosphate (3) epimerase (RPE)
This enzyme is also called pentose-5-phosphate 3-epimerase or PPE. Sugar epimerase enzymes alter the stereochemistry of the sugar at one of the alcohol positions creating a different epimer. The RPE enzyme has a Zn2+ cofactor, however, its requirement for enzyme activity is not essential. A Zn2+ independent form can still function and stabilize an oxyanion intermediate with adjacent and conserved methionines. A mechanism of the Zn2+ dependent form of the enzyme is shown in Figure \(\PageIndex{15}\).
Figure \(\PageIndex{15}\): Mechanism for rice ribulose-5-phosphate epimerase
The enzyme utilizes an acid/base catalytic mechanism that mediates the formation of a trans-2,3-enediol phosphate intermediate. Key aspartic acid residues act as proton donors and acceptors during the reaction. A zinc metal cofactor helps stabilize charges during the reaction.
Figure \(\PageIndex{16}\) shows an interactive iCn3D model of cytosolic D-ribulose-5-phosphate 3-epimerase from rice (1h1z).
Figure \(\PageIndex{16}\): Cytosolic D-ribulose-5-phosphate 3-epimerase from rice (1h1z). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...AH7uS2T8hsnpXA
Transketolase (TK)
The pool of ribose 5-phosphate that is created can be utilized for nucleotide production, or some of it can undergo a reaction with the xylulose 5-phosphate created in the RPE reaction. The transketolase enzyme converts two five-carbon sugars (ribose 5-phosphate and xylulose 5-phosphate) into a 3-carbon and a 7-carbon sugar (glyceraldehyde 3-phosphate and sedoheptulose 7-phosphate). Essentially, transketolase enzymes transfer ketone functional groups from ketoses to aldoses, effectively creating a new ketose that is two carbons larger. The ketose donor then becomes an aldose with two fewer carbons. The basic transketolase reaction shown in Figure \(\PageIndex{17}\)
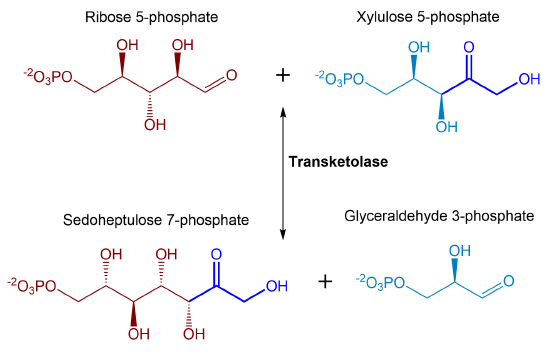
Figure \(\PageIndex{17}\): Transketolase reaction. The reaction shows the reversible conversion of ribose 5-phosphate and xylulose 5-phosphate to sedoheptulose 7-phosphate and glyceraldehyde 3-phosphate.
Note the number of carbons in the reactants and products: 5C + 5C ↔ 3C + 7C. In this reversible reaction, the enzyme uses the cofactor thiamine pyrophosphate (TPP) and a divalent cation. The enzyme transfers a 2C ketol group from xylulose-5-phosphate to ribose-5-phosphate, an aldose. The product glyceraldehyde-3-phosphate is a glycolytic intermediate and can be used in the glycolytic pathway.
The mechanism for the reverse reaction (yeast numbering system) is shown in Figure \(\PageIndex{18}\). In this reaction, sedoheptulose 7-phosphate binds with the enzyme and the TPP cofactor is activated to form a carbanion. The carbanion mediates nucleophilic attack on the carbonyl carbon of the substrate forming a covalent intermediate. His263 serves as a base and abstracts a proton, enabling bond cleavage and the formation of ribose 5-phosphate. Ribose 5-phosphate leaves the active site and glyceraldehyde 3-phosphate enters. The two carbon intermediate covalently bound to the TPP mediates nucleophilic attack on the glyceraldehyde 3-phosphate enabling the formation of xylulose 5-phosphate and the restoration of the enzyme.
Figure \(\PageIndex{18}\): Mechanism of the transketolase reaction during the pentose phosphate pathway.
https://www.ebi.ac.uk/thornton-srv/m-csa/entry/219/
Figure \(\PageIndex{19}\) shows an interactive iCn3D model of Human transketolase in a covalent complex with donor ketose D-xylulose-5-phosphate (4kxv).
Figure \(\PageIndex{19}\): Human transketolase in covalent complex with donor ketose D-xylulose-5-phosphate (4kxv). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...WtGtQ7mKA6UYx7
The enzyme is a dimer (gray and cyan coloring). Numbering of key residues compared to yeast mechanism:
| Human | Yeast |
| E366 | E418 |
| Q428 | H481 |
| H258 | H262 |
| H38 | H30 |
Also shown are human active site residues H474, R318, and S345. The covalent xylulose-5-phosphate adduct is shown in spacefill and labeled DX5 (xylitol-5-phosphate). TPP is shown in sticks and labeled. The metal ion is Ca2+. Structural analyzes show a 200 distortion in the planarity of the cofactor-substrate bond and a lengthening of the C-C bond in the substrate which breaks.
Transaldolase
In addition to being used as a resource in the glycolytic pathway, glyceraldehyde 3-phosphate can also be utilized as a substrate in a transaldolase reaction along with the sedoheptulose 7-phosphate that is produced in the previous reaction. This results in the formation of erythrose 4-phosphate and fructose 6-phosphate. A summary of the reaction is shown in Figure \(\PageIndex{20}\).
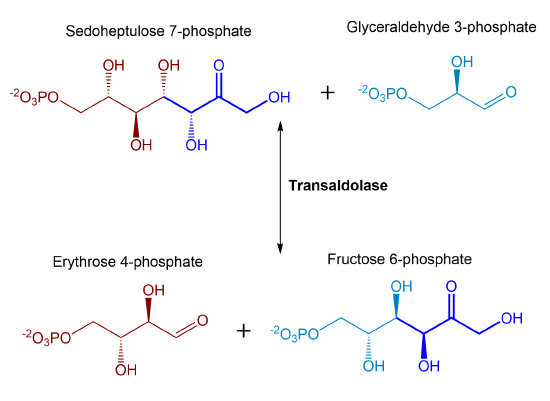
Figure \(\PageIndex{20}\): Summary of transaldolase reaction
As with transketolase, the transaldolase enzyme is reversible. Again, note the number of carbons in the reactants and products: 3C + 7C ↔ 4C + 6C. Unlike the transketolase used in the last reaction, the transaldolase enzyme does NOT use TPP as a cofactor. Instead, it forms a Schiff base intermediate similar to that of the aldolase enzyme in the glycolytic pathway. The enzyme removes a 3C ketol group (dihydroxyacetone) from sedoheptulose 7-phosphate and transfers it to glyceraldehyde 3-phosphate forming fructose 6-phosphate. Erythrose 4-phosphate is left from the original sedoheptulose 7-phosphate.
We will explore the mechanism for the E. Coli enzyme in the reverse direction. The first part of the mechanism of transaldolase is shown in Figure \(\PageIndex{21}\).
Figure \(\PageIndex{21}\): Mechanism for the first half of the transaldolase reaction. https://www.ebi.ac.uk/thornton-srv/m-csa/entry/148/
In the first part of the reaction, fructose 6-phosphate binds to the active site of the enzyme, where an active site lysine residue mediates nucleophilic attack on the carbonyl carbon and forms a covalent intermediate with the enzyme. Formation of the Schiff base leads to dehydration of the intermediate. The Schiff base nitrogen becomes protonated and this leads to the oxidation of the C4 hydroxyl leads and subsequent bond cleavage releasing glyceraldehyde 3-phosphate. The remaining ES-complex rearranges to form an enol intermediate.
The mechanism for the second half of the transaldolase reaction is shown in Figure \(\PageIndex{22}\).
Figure \(\PageIndex{22}\): Mechanism for the second half of the transaldolase reaction. https://www.ebi.ac.uk/thornton-srv/m-csa/entry/148/
Once glyceraldehyde 3-phosphate has left the active site of the enzyme, erythrose 4-phosphate can bind. The enol from the ES intermediate mediates nucleophilic attack on the aldehyde carbonyl group of erythrose 4-phosphate. This results in the formation of a Schiff base intermediate. Hydration at the Schiff base carbon atom ensues followed by the oxidation of the newly incorporated alcohol to form a ketone functional group. Formation of the ketone causes bond cleavage between the enzyme and the newly formed ketose, sedoheptulose 7-phosphate.
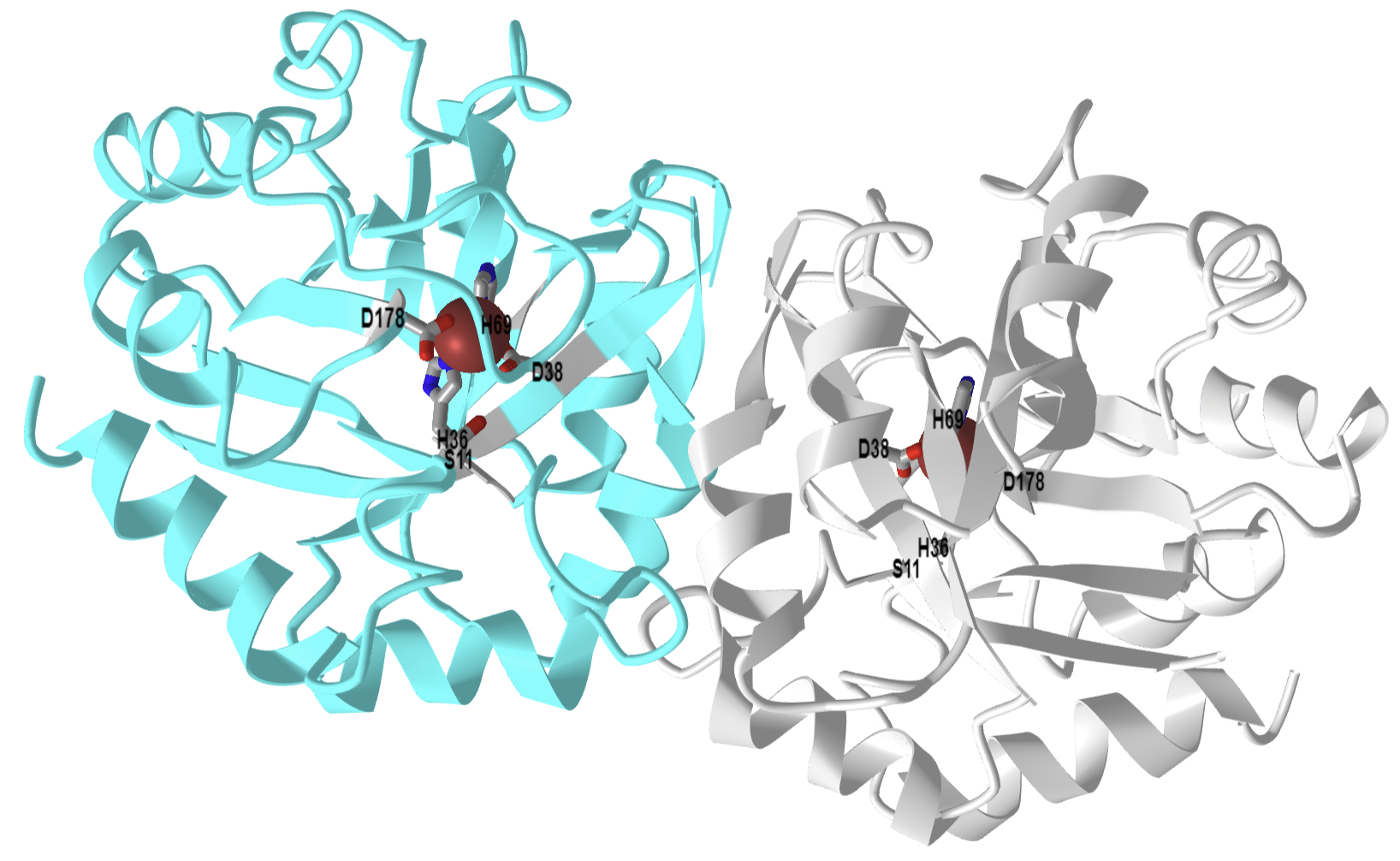
Figure \(\PageIndex{23}\) shows an interactive iCn3D model of transaldolase B from Escherichia coli (1ONR).
Figure \(\PageIndex{23}\): 3D structure of Transaldolase B from E. coli (1ONR). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...7swLZVpNPe3eU9
Transketolase
As seen above, fructose 6-phosphate is a product of the transaldolase enzyme during the nonoxidative branch of the PPP. This sugar can feed back into the glycolytic cycle. The formation of fructose 6-phosphate can also occur using the transketolase enzyme when it transfers a 2C ketol group from xylulose 5-phosphate to the aldose, erythrose 4-phosphate. This reaction is summarized in Figure \(\PageIndex{24}\).
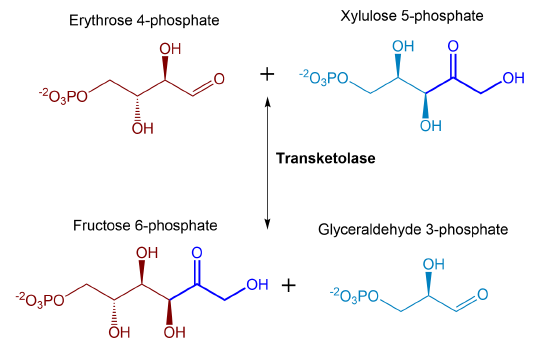
Figure \(\PageIndex{24}\) Summary of second transketolase reaction during the pentose phosphate pathway.
Both products of this final transketolase reaction can be utilized in energy formation through glycolysis. Thus, there are clear metabolic ties of the PPP with the energy-producing glycolytic pathway and intermediates from both pathways can easily be exchanged through the activity of these reversible enzymes. The PPP provides key intermediates, including ribose 5-phosphate that is used in the biosynthesis of nucleotides and ultimately nucleic acids, as well as, other important metabolic molecules such as FAD, NAD+, NADP+, and Coenzyme A. In addition to being a biosynthetic source for these molecules, the PPP also is the primary source for reducing NADP+ to NADPH, during the oxidative reactions. NADPH is utilized in many biosynthetic processes as an electron donor. For example, the biosynthesis of many lipid molecules requires NADPH, including the production of triacylglycerols, phospholipids, and steroids. NADPH is also required for the biosynthesis of some amino acids (such as glutamate and proline) and is also involved in the reduction of ribonucleotides and deoxyribonucleotides during the synthesis of RNA and DNA. NADPH is also utilized by a number of oxidoreductases involved in detoxification reactions within the body. Within the immune system, NADPH oxidases or NOX enzymes are involved in the production of superoxide and utilized to damage invading pathogens nonspecifically. NADP+/NADPH ratios also can play a regulatory role in cellular metabolic processes and are utilized as allosteric effectors for several enzymes and oxidation sensor proteins.
Regulation of the Pentose Phosphate Pathway
The cellular demand for the two major products of the PPP (ribose 5-phosphate and reduced NADPH) can be different depending on the cell type or the current cellular environment, such as times of increased metabolic demand or oxidative stress. Thus, the two major products may need to be produced in different quantities and independently of one another. For example, we can imagine times when the needs for ribose 5-phosphate and NADPH are in balance with the standard PPP reactions. However, we can also imagine times when the demand for ribose 5-phosphate may be much higher than the demand for NADPH, or vice versa, the demand for NADPH may be much greater than the demand for ribose 5-phosphate. Thus, there are regulatory strategies in place that enable the ability to regulate the production of these different pools independently of one another and adapt to cellular needs. This ability is largely dependent on the production of metabolic intermediates that can be easily interchanged within the glycolytic pathway.
For example, the ratio of NADP+/NADPH serves as a key regulator of the oxidative branch of the PPP. The first enzymatic step of the pathway mediated by the glucose 6-phosphate dehydrogenase (G6PD) is regulated in this fashion and helps control the pool of glucose 6-phosphate that will be utilized within the PPP to produce reduced NADPH. Low levels of NADP+ inhibit the G6PD enzyme. The G6PD reaction is essentially irreversible and serves as the committed step for glucose to enter into the oxidative portion of the PPP. Thus the regulation of this enzymatic step is key in the regulation of NADPH levels within the cell. When the ratio of NADP+/NADPH increases, G6PD becomes more active and the reduction of NADP+ to NADPH increases.
When ribose 5-phosphate is also in high demand, the pool of ribose 5-phosphate will be low and increase the activity of the ribose 5-phosphate isomerase in the forward direction to convert the ribulose 5-phosphate produced in the oxidative branch to generate more ribose 5-phosphate.
However, if more ribose 5-phosphate is required than can be supplied from the activity of the oxidative branch, the pool of ribose 5-phosphate can be fed from the conversion of intermediates from the glycolytic pathway in the nonoxidative branch of the PPP. The opposite occurs as well when there is a high need for NADPH in the cell but a low need for ribose 5-phosphate. In this situation, ribose 5-phosphate is converted to fructose and glyceraldehyde 3-phosphate and can be incorporated into the glycolytic pathway.
In addition, the activity of the PPP varies depending on the tissue type and location within the body. For example, skeletal muscle has very low PPP activity, as this tissue requires more energy production and activity of the glycolytic pathway. On the other hand, the PPP is highly active in adipose tissue due to the heightened requirement for intermediates needed for lipid biosynthesis.
References:
Edwards et al.: Structural characterization of a ribose5-phosphate isomerase B from the pathogenic fungus Coccidioides immitis. BMC Structural Biology 2011 11:39. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3212906/
Ge T, Yang J, Zhou S, Wang Y, Li Y and Tong X (2020) The Role of the Pentose Phosphate Pathway in Diabetes and Cancer. Front. Endocrinol. 11:365. doi: 10.3389/fendo.2020.00365
Polat, I.H., Tarrado-Castellarnau, M., Bharat, R., Perarnau, J., Benito, A., Cortes, R., Sabatier, P., and Cascante, M. (2021) Oxidative Pentose Phosphate Pathway Enzyme 6-Phosphogluconate Dehydrogenase Plays a Key Role in Breast Cancer Metabolism. Biology 10(2):85. https://doi.org/10.3390/biology10020085
Wikipedia contributors. (2022, March 6). Glucose-6-phosphate dehydrogenase. In Wikipedia, The Free Encyclopedia. Retrieved 22:01, August 12, 2022, from https://en.wikipedia.org/w/index.php?title=Glucose-6-phosphate_dehydrogenase&oldid=1075517394
Wikipedia contributors. (2021, August 29). 6-phosphogluconolactonase. In Wikipedia, The Free Encyclopedia. Retrieved 23:05, August 12, 2022, from https://en.wikipedia.org/w/index.php?title=6-phosphogluconolactonase&oldid=1041297267
Wikipedia contributors. (2022, March 28). 6-Phosphogluconate dehydrogenase. In Wikipedia, The Free Encyclopedia. Retrieved 05:50, August 14, 2022, from https://en.wikipedia.org/w/index.php?title=6-Phosphogluconate_dehydrogenase&oldid=1079702040
Wikipedia contributors. (2022, July 4). Ribose-5-phosphate isomerase. In Wikipedia, The Free Encyclopedia. Retrieved 05:33, August 15, 2022, from https://en.wikipedia.org/w/index.php?title=Ribose-5-phosphate_isomerase&oldid=1096406233




.png?revision=1&size=bestfit&width=520&height=329)
.png?revision=1&size=bestfit&width=560&height=322)
.png?revision=1&size=bestfit&width=343&height=361)
.png?revision=1&size=bestfit&width=341&height=277)
.png?revision=1&size=bestfit&width=404&height=394)
.png?revision=1&size=bestfit&width=510&height=370)