7.13A: Strategies Used in Sequencing Projects
- Page ID
- 9310
The strategies used for sequencing genomes include the Sanger method, shotgun sequencing, pairwise end, and next-generation sequencing.
LEARNING OBJECTIVES
Compare the different strategies used for whole-genome sequencing: Sanger method, shotgun sequencing, pairwise-end sequencing, and next-generation sequencing
Key Takeaways
Key Points
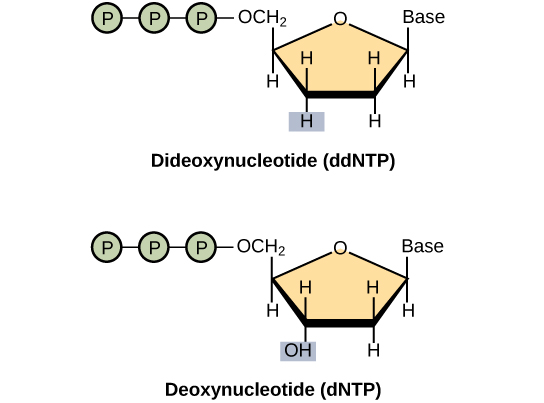
- The Sanger method is a basic sequencing technique that uses fluorescently-labeled dideoxynucleotides (ddNTPs) during DNA replication which results in multiple short strands of replicated DNA that terminate at different points, based on where the ddNTP was incorporated.
- Shotgun sequencing is a method that randomly cuts DNA fragments into smaller pieces and then, with the help of a computer, takes the DNA fragments, analyzes them for overlapping sequences, and reassembles the entire DNA sequence.
- Pairwise-end sequencing is a type of shotgun sequencing which is used for larger genomes and analyzes both ends of the DNA fragments for overlap.
- Next-generation sequencing is a type of sequencing which is automated and relies on sophisticated software for rapid DNA sequencing.
Key Terms
- fluorophore: a molecule or functional group which is capable of fluorescence
- contig: a set of overlapping DNA segments, derived from a single source of genetic material, from which the complete sequence may be deduced
- dideoxynucleotide: any nucleotide formed from a deoxynucleotide by loss of an a second hydroxyl group from the deoxyribose group
Strategies Used in Sequencing Projects
The basic sequencing technique used in all modern day sequencing projects is the chain termination method (also known as the dideoxy method), which was developed by Fred Sanger in the 1970s. The chain termination method involves DNA replication of a single-stranded template with the use of a primer and a regular deoxynucleotide (dNTP), which is a monomer, or a single unit, of DNA. The primer and dNTP are mixed with a small proportion of fluorescently-labeled dideoxynucleotides (ddNTPs). The ddNTPs are monomers that are missing a hydroxyl group (–OH) at the site at which another nucleotide usually attaches to form a chain. Each ddNTP is labeled with a different color of fluorophore. Every time a ddNTP is incorporated in the growing complementary strand, it terminates the process of DNA replication, which results in multiple short strands of replicated DNA that are each terminated at a different point during replication. When the reaction mixture is processed by gel electrophoresis after being separated into single strands, the multiple, newly-replicated DNA strands form a ladder due to their differing sizes. Because the ddNTPs are fluorescently labeled, each band on the gel reflects the size of the DNA strand and the ddNTP that terminated the reaction. The different colors of the fluorophore-labeled ddNTPs help identify the ddNTP incorporated at that position. Reading the gel on the basis of the color of each band on the ladder produces the sequence of the template strand.

Early Strategies: Shotgun Sequencing and Pair-Wise End Sequencing
In the shotgun sequencing method, several copies of a DNA fragment are cut randomly into many smaller pieces (somewhat like what happens to a round shot cartridge when fired from a shotgun). All of the segments are then sequenced using the chain-sequencing method. Then, with the help of a computer, the fragments are analyzed to see where their sequences overlap. By matching overlapping sequences at the end of each fragment, the entire DNA sequence can be reformed. A larger sequence that is assembled from overlapping shorter sequences is called a contig. As an analogy, consider that someone has four copies of a landscape photograph that you have never seen before and know nothing about how it should appear. The person then rips up each photograph with their hands, so that different size pieces are present from each copy. The person then mixes all of the pieces together and asks you to reconstruct the photograph. In one of the smaller pieces you see a mountain. In a larger piece, you see that the same mountain is behind a lake. A third fragment shows only the lake, but it reveals that there is a cabin on the shore of the lake. Therefore, from looking at the overlapping information in these three fragments, you know that the picture contains a mountain behind a lake that has a cabin on its shore. This is the principle behind reconstructing entire DNA sequences using shotgun sequencing.
Originally, shotgun sequencing only analyzed one end of each fragment for overlaps. This was sufficient for sequencing small genomes. However, the desire to sequence larger genomes, such as that of a human, led to the development of double-barrel shotgun sequencing, more formally known as pairwise-end sequencing. In pairwise-end sequencing, both ends of each fragment are analyzed for overlap. Pairwise-end sequencing is, therefore, more cumbersome than shotgun sequencing, but it is easier to reconstruct the sequence because there is more available information.
Next-generation Sequencing
Since 2005, automated sequencing techniques used by laboratories are under the umbrella of next-generation sequencing, which is a group of automated techniques used for rapid DNA sequencing. These automated, low-cost sequencers can generate sequences of hundreds of thousands or millions of short fragments (25 to 500 base pairs) in the span of one day. Sophisticated software is used to manage the cumbersome process of putting all the fragments in order.


