4.11: Enzyme Inhibition
- Page ID
- 3040
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Inhibition of specific enzymes by drugs can be medically useful. Understanding the mechanisms of enzyme inhibition is therefore of considerable importance. We will discuss four types of enzyme inhibition – competitive, non- competitive, uncompetitive, and suicide. Of these, the first three types are reversible. The last one is not.
Competitive Inhibition
Probably the easiest type of enzyme inhibition to understand is competitive inhibition and it is the one most commonly exploited pharmaceutically. Molecules that are competitive inhibitors of enzymes resemble one of the normal substrates of an enzyme. An example is methotrexate, which resembles the folate substrate of the enzyme dihydrofolate reductase (DHFR). This enzyme normally catalyzes the reduction of folate, an important reaction in the metabolism of nucleotides. When the drug methotrexate is present, some of the enzyme binds to it instead of to folate and during the time methotrexate is bound, the enzyme is inactive and unable to bind folate. Thus, the enzyme is inhibited. Notably, the binding site on DHFR for methotrexate is the active site, the same place that folate would normally bind. As a result, methotrexate ‘competes’ with folate for binding to the enzyme. The more methotrexate there is, the more effectively it competes with folate for the enzyme’s active site. Conversely, the more folate there is, the less of an effect methotrexate has on the enzyme because folate outcompetes it.

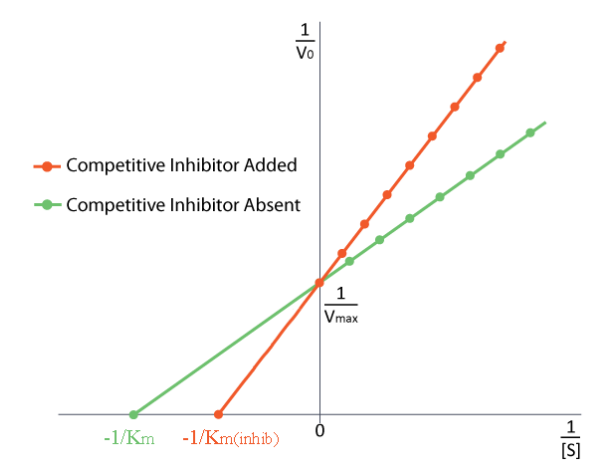
No Effect On \(V_{MAX}\)
How do we study competitive inhibition. It is typically done as follows. First one performs a set of V vs. [S] reactions without inhibitor (20 or so tubes, with buffer and constant amounts of enzyme, varying amounts of substrate, equal reaction times). V vs. [S] is plotted, as well as 1/V vs. 1/[S], if desired. Next, a second set of reactions is performed in the same manner as before, except that a fixed amount of the methotrexate inhibitor is added to each tube. At low concentrations of substrate, the inhibitor competes for the enzyme effectively, but at high concentrations of substrate, the inhibitor will have a much reduced effect, since the substrate outcompetes it, due to its higher concentration (remember that the inhibitor is at fixed concentration). Graphically, the results of these experiments are shown above. Notice that at high substrate concentrations, the competitive inhibitor has essentially no effect, causing the Vmax for the enzyme to remain unchanged. To reiterate, this is due to the fact that at high substrate concentrations, the inhibitor doesn’t compete well. However, at lower substrate concentrations it does.
Increased KM
Note that the apparent KM of the enzyme for the substrate increases (-1/KM gets closer to zero - red line above) when the inhibitor is present, thus illustrating the better competition of the inhibitor at lower substrate concentrations. It may not be obvious why we call the changed KM the apparent KM of the enzyme. The reason is that the inhibitor doesn’t actually change the enzyme’s affinity for the folate substrate. It only appears to do so. This is because of the way that competitive inhibition works. When the competitive inhibitor binds the enzyme, it is effectively ‘taken out of action.’ Inactive enzymes have NO affinity for substrate and no activity either. We can’t measure KM for an inactive enzyme.
The enzyme molecules that are not bound by methotrexate can, in fact, bind folate and are active. Methotrexate has no effect on them and their KM values are unchanged. Why then, does KM appear higher in the presence of a competitive inhibitor. The reason is that the competitive inhibitor is reducing the amount of active enzyme at lower concentrations of substrate. When the amount of enzyme is reduced, one must have more substrate to supply the reduced amount of enzyme sufficiently to get to Vmax/2.
It is worth noting that in competitive inhibition, the percentage of
inactive enzyme changes drastically over the range of [S] values
used. To start, at low [S] values, the greatest percentage of the
enzyme is inhibited. At high [S], no significant percentage of
enzyme is inhibited. This is not always the case, as we shall see
in non-competitive inhibition.
Non-Competitive Inhibition
A second type of inhibition employs inhibitors that do not resemble the substrate and bind not to the active site, but rather to a separate site on the enzyme (rectangular site below). The effect of binding a non-competitive inhibitor is significantly different from binding a competitive inhibitor because there is no competition. In the case of competitive inhibition, the effect of the inhibitor could be reduced and eventually overwhelmed with increasing amounts of substrate. This was because increasing substrate made increasing percentages of the enzyme active. With non-competitive inhibition, increasing the amount of substrate has no effect on the percentage of enzyme that is active. Indeed, in non-competitive inhibition, the percentage of enzyme inhibited remains the same through all ranges of [S].

This means, then, that non-competitive inhibition effectively reduces the amount of enzyme by the same fixed amount in a typical experiment at every substrate concentration used The effect of this inhibition is shown above. As you can see, Vmax is reduced in non-competitive inhibition compared to uninhibited reactions. This makes sense if we remember that Vmax is dependent on the amount of enzyme present. Reducing the amount of enzyme present reduces Vmax. In competitive inhibition, this doesn’t occur detectably, because at high substrate concentrations, there is essentially 100% of the enzyme active and the Vmax appears not to change. Additionally, KM for non-competitively inhibited reactions does not change from that of uninhibited reactions. This is because, as noted previously, one can only measure the KM of active enzymes and KM is a constant for a given enzyme.

Uncompetitive Inhibition
A third type of enzymatic inhibition is that of uncompetitive inhibition, which has the odd property of a reduced Vmax as well as a reduced KM. The explanation for these seemingly odd results is rooted in the fact that the uncompetitive inhibitor binds only to the enzyme-substrate (ES) complex. The inhibitor-bound complex forms mostly under concentrations of high substrate and the ES-I complex cannot release product while the inhibitor is bound, thus explaining the reduced Vmax.
The reduced KM is a bit harder to conceptualize. The answer lies in the fact that the inhibitor-bound complex effectively reduces the concentration of the ES complex. By Le Chatelier’s Principle, a shift occurs to form additional ES complex, resulting in less free enzyme and more enzyme in the forms ES and ESI (ES with inhibitor). Decreases in free enzyme correspond to an enzyme with greater affinity for its substrate. Thus, paradoxically, uncompetitive inhibition both decreases Vmax and increases an enzyme’s affinity for its substrate.
Suicide Inhibition
In contrast to the first three types of inhibition, which involve reversible binding of the inhibitor to the enzyme, suicide inhibition is irreversible because the inhibitor becomes covalently bound to the enzyme during the inhibition and thus cannot be removed. Suicide inhibition rather closely resembles competitive inhibition because the inhibitor generally resembles the substrate and binds to the active site of the enzyme. The primary difference is that the suicide inhibitor is chemically reactive in the active site and makes a bond with it that precludes its removal. Such a mechanism is that employed by penicillin (Figure 4.10.5), which covalently links to the bacterial enzyme, D-D transpeptidase and stops it from functioning. Since the normal function of the enzyme is to make a bond necessary for the peptido-glycan complex of the bacterial cell wall, the cell wall cannot properly form and bacteria cannot reproduce. If one were to measure the kinetics of suicide inhibitors under conditions where there was more enzyme than inhibitor, they would resemble non-competitive inhibition’s kinetics because both involve reducing the amount of active enzyme by a fixed amount in a set of reactions.