W2017_Lecture_19_reading
- Page ID
- 8268
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)The scope of the problem
In this module we discuss the replication of DNA - one of the key requirements for a living system to regenerate and create the next generation. Let us first briefly consider the scope of the problem by way of a literary analogy.
The human genome consists of roughly 6.5 billion base pairs of DNA if one considers the full diploid genome (i.e. if you count the DNA inherited from both parents). Six point five billion looks like this: 6,500,000,000. That's a large number. To get a better idea of what that number means, imagine that our DNA is a set of written instructions for constructing one of us. By analogy we can then compare it to another written document. For this example we begin by considering Tolstoy's "War and Peace", a novel many people are familiar with for its voluminous nature. Data from wikipedia estimates that "War and Peace" contains about 560,000 words. A second written work many are familiar with are the seven volumes of J.K. Rowling's "Harry Potter". This work checks in at ~1,080,000 words (Referenced Statistics on Wikipedia). If we assume that the length of the average English word is 5 characters, the two literary works are 2.8 million and 5.4 million characters in length, respectively. Therefore, even all seven volumes of "Harry Potter" have over 1000x fewer characters than our own genomes. The number of characters in these novels are, however, much closer to the number of nucleotides in a typical bacterial genome.
Now imagine for a moment developing a machine or mechanical process (not an electronic process) that is responsible for reading and copying these books. Or imagine yourself copying these texts. How fast could you do it? How many mistakes are you likely to make? Do you expect there to be a trade-off between the speed at which you can copy and the accuracy? What type of resources does this process need? How much energy is required? Now imagine copying something 1000x larger! Oh, and just for good measure, your imaginary mechanical device needs to do its work on text that is ~25Å wide (i.e. 0.0000000025 meters wide). By comparison, a typical 10 point font is ~0.00025 meters wide, about 100,000x larger than with width of a DNA base pair.
With that in mind, it is worth noting that a human cell can take about 24 hours to divide (DNA replication must therefore be a little faster). A healthy E. coli cell may take only 20 minutes to divide (including replicating its ~4.5 million base pair genome). Both the human and bacterium do this while typically making few enough mistakes that the subsequent generation remains viable and recognizable. That should seem rather amazing! Now consider that the human body is estimated to consist of ~ 10 trillion cells (10,000,000,000,000) and that it may have between 2 and 10 times that number of microbial residents and that's a lot of cell division to consider.
Design challenge
If the cell is to replicate - its ultimate goal - a copy of the DNA must be created. So one clear problem statement/question is "how can the cell effectively copy its DNA?" Given the analogy above, some relevant subquestions of relevance might be: What are the chemical and physical properties that enable DNA to be copied? With what fidelity must the DNA be copied? What speed must it be copied at? Where does the energy come from for this task and how much is necessary? Where do the "raw materials" come from? How do the molecular machines involved in this process couple the assembly of raw materials and the energy required to build a new DNA molecule together? The list could, of course, go on.
In the following and in lecture we will be interested in starting to examine how the process of DNA replication is accomplished while keeping in mind some of the driving questions. As you go through the reading and lecture materials try to be constantly aware of these and other questions associated with this process. Use these questions as guideposts for organizing your thoughts and try to find matches between the "facts" that you think you might be expected to know and the driving questions.
The DNA Double Helix
To build some extra context we also need a little bit of empirically determined knowledge. Perhaps one of the best known and popular features of the hereditary form of the DNA molecule is that it has a double helical tertiary structure. The appreciation of this dates to the the 1950s. The story of this discovery has been widely recounted - and the details are beyond the scope of this text. Briefly, Francis Crick and James Watson are credited with determining the structure of DNA. Rosalind Franklin is now also widely credited with generating critical X-ray diffraction data that enabled Watson and Crick to piece together the puzzle of the DNA molecule.
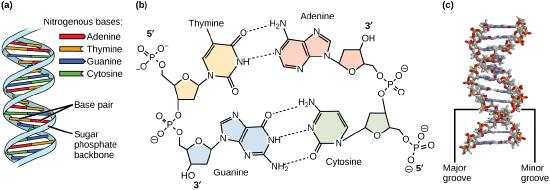
Models of the structure of DNA revealed the molecule is made up of two strands of covalently linked nucleotides that are twisted around each other to form a right-handed helix. In each strand, nucleotides are covalently joined to two other nucleotides (except at the very ends of a linear strand) via phosphodiester bonds that link the sugars via the 5' and 3' hydroxyl groups (see panel b in the figure below) - recall that the labels 5' and 3' refer to the carbons on the sugar molecule. These sugar and phosphate chains form a contiguous set of covalent links that are often referred to as the "backbone" of the structure. In a linear molecule, each strand has two free ends. One is termed the 5'-end because the unlinked functional group that is typically involved in joining nucleotides is the phosphate linked to the 5' carbon. The other end of the strand is called the 3'-end because the unlinked functional group that is typically involved in joining nucleotides is the hydroxyl group linked to the 3' carbon of the sugar. Since the two ends of the strand are not symmetric this makes it easy to designate or describe a direction one the strand - one can, for instance, say that they are reading from the 5'-end to 3'-end to indicate that they are "walking" along the strand starting at the 5'-end and and moving towards the 3'-end. This direction (5' to 3') is, by the way, the convention used by most biologists. One can read in the opposite direction (3' to 5') provided this is made explicit. The two strands of covalently linked nucleotides are found to be anti-parallel to one another in the double-helix; that is, the orientation/direction of one strand is opposite to that of the other strand (see panel b in the figure below). The backbone is structurally on the "outside" of the double helix, creating a band of negative charges on the surface. By contrast, the nitrogenous bases of each of the antiparallel strands stack on the inside of the structure and oppose one another in a way that allows hydrogen bonds between unique purine/pyrimidine pairs (A pairing with T and G pairing with C) to form. These specific pairings are said to be complementary to on another and thus the opposite strands of a double helix are often referred to one another as complementary strands.
Complementary strands carry redundant information. Because of the strict chemical pairing, if you know the sequence of one strand you obligatorily know the strand of its complement. Take for example the sequence 5′- C A T A T G G G A T G - 3′. Note how the sequence is annotated with the orientation (indicated by 5' and 3' labels). The complement of this sequence - written according to the 5' to 3' convention is: 5′- C A T C C C A T A T G - 3′. If you aren't convinced, write these two sequences out across from one another in your notes, making sure to write them as antiparallel strands. Note that the twisting of the two complementary strands around each other results in the formation of structural features called the major and minor grooves that will become more important when we discuss the binding of proteins to DNA (see panel c in the figure below).
Most of the Bis2a instructors will expect you to recognize key structural features depicted in the figure below and that you will be able to create a basic figure of the structure of DNA yourself.
DNA has (a) a double helix structure and (b) phosphodiester bonds. The (c) major and minor grooves are binding sites for DNA binding proteins during processes such as transcription (the creation of RNA from a DNA template) and replication.
At around the same time the structure was being solved three hypotheses for modes of DNA replication were being considered. The models for replication were known as: the conservative model, the semi-conservative model and the dispersive model.
1. Conservative: The conservative model of replication postulated that each whole double-stranded molecule could act as a template for the synthesis of a completely new double-stranded molecule. That is, if one were to put a chemical tag on the template DNA molecule after replication none of that tag would be found on the new copy.
2. Semi-conservative: This hypothesis stipulated that each individual strand of a DNA molecule could serve as a template for a new strand to which it would now associate with. in this case, if a chemical label were placed on a double stranded DNA molecule, one strand on each of the copies would retain the label.
3. Dispersive: This model proposed that a copied double-helix would be a piecewise combination of continuous segments of "old" and "new" strands. If a chemical label were placed on a DNA molecule that were copied using a dispersive mechanism, one would find discrete segments of the resulting copy that were labeled on both strands separated by completely unlabeled parts.
Meselson and Stahl resolved the issue in 1958 when they reported results of a now famous experiment (describe on Wikipedia) which showed that DNA replication is semi-conservative (see figure below), where each strand is used as a template for the creation of the new strand. To learn more about this experiment watch The Meselson-Stahl Experiment.
DNA has an anti-parallel double helix structure, the nucleotide bases are hydrogen bonded together and each strand complements the other. DNA is replicated in a semi-conservative manner, each strand is used as the template for the newly made strand.
DNA Replication
Having established some basic structural features and the need for a semi-conservative mechanism it is important to begin understanding some of what is known about the process and to think about what questions one might want to answer if they are to better understand what is going on.
Since DNA replication is a process we can invoke the energy story rubric to think about it. Recall that the energy story rubric is there to help us think systematically about processes (how things go from A to B). In this case the process in question is the act of starting with one double-stranded DNA molecule and ending up with two double-stranded molecules. So, we will ask things like: What does the system look like at the beginning (matter and energy) of replication? How are matter and energy transferred in the system and what catalyzes the transfers? What does the system look like at the end of the process? We can also ask questions regarding specific events that MUST happen during the process. For instance, since DNA is a long molecule and it is sometimes circular, we can ask basic questions like, where does the process of replication start? Where does it end? We can also ask practical questions about the process like, what happens when a double-stranded structure is unwound?
We consider some of these key questions in the text and in class, and encourage you to do the same.
Requirements for DNA replication
Let's start by listing some basic functional requirements for DNA replication that we can infer just by thinking about the process that must happen and/or be required for the replication to happen. So, what do we need?
• We know that DNA is composed of nucleotides. If we are going to create a new strand we will need a source of nucleotides.
• We can infer that building a new strand of DNA will require an energy source - we should try to find this.
• We can infer that that there must be a process for finding a place to start replication.
• We can infer that there will be one or more enzymes that help catalyze the process of replication.
• We can also infer that since this is a biochemical process, that some mistakes will be made.
Nucleotide Structure Review
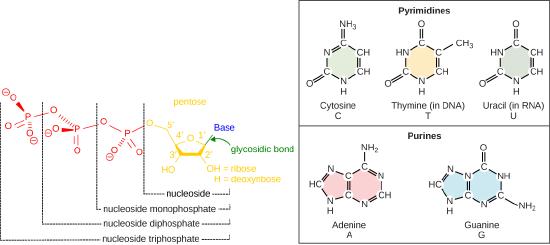
Recall some basic structural features of the nucleotide building blocks of DNA including: the nucleotides start as off as nucleotide triphosphates. Nucleotide are composed of a nitrogenous base, deoxyribose (5-carbon sugar), and a phosphate group. The nucleotide is named according to its nitrogenous base, purines such as adenine (A) and guanine (G), or pyrimidines such as cytosine (C) and thymine (T). Recall the structures below. Note that the nucleotide Adenosine triphosphate (ATP) is a precursor of the deoxyribonucleotide (dATP) which is incorporated into DNA.
Initiation of Replication
Where along the DNA does the replication machinery start DNA replication?
With millions, if not billions, of nucleotides to copy how does the DNA polymerase know where to start? Not surprisingly this process turns out to not be random. There are specific nucleotide sequences called origins of replication along the length of the DNA at which replication begins. Once this site is identified, however, there is a problem. The DNA double helix is held together by base stacking interactions and hydrogen bonds. If each strand is to be read and copied individually, there must be some mechanism responsible for helping to dissociate the two strands from one another. Energetically, this is an endergonic process. Where does the energy come from and how is this reaction catalyzed? Basic reasoning should, at this point, lead to the hypothesis that an protein catalyst is likely involved and that this enzyme either creates new bonds that are energetically more favorable (exergonic) than the bonds it breaks AND/OR it is able to couple the use of an external energy source to help dissociate the strands.
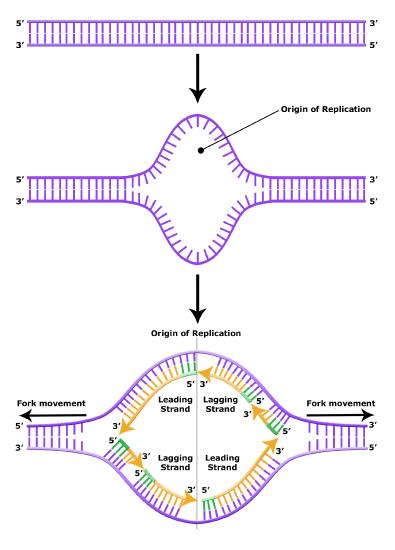
It turns out that the details of this process and the proteins involved differ depending on the specific organism in question and many of the molecular level details are yet to be completely understood. There are, however, some common features in the replication of eukaryotes, bacteria and archaea and one of these features is that multiple different types of proteins are involved in replicating DNA. First, proteins generally called "initiators" have the capacity to bind DNA at or very near origins of replication. The interaction of the initiator proteins with the DNA helps to destabilize the double helix and also help to recruit other proteins, including an enzyme called a DNA helicase to the DNA. In this case the energy required to destabilize the DNA double helix seems to come from the formation of new associations between DNA and the initiator proteins and the proteins themselves. The DNA helicase is a multi-subunit protein that is important in the process of replication because it couples the exergonic hydrolysis of ATP to the unwinding of the DNA double helix. Additional proteins must be recruited to the initiation complex (the collection of proteins involved in initiating transcription). These include, but are not limited to, additional enzymes called primase and DNA polymerase. While the initiators are lost soon after the initiation of replication, the rest of the proteins work in concert to execute the process of DNA replication. This complex of enzymes function at Y-shaped structures in the DNA called replication forks (see figure below). For any replication event two replication forks may be formed at each origin of replication, extending in both directions. Multiple origins of replication can be found on eukaryotic chromosomes and some archaea, while the the genome of the bacterium, E. coli, seems to encode one origin of replication.
Suggested discussion
Why would different organisms have different numbers of replication origins? What could the benefit be to having more than one? Is there a drawback to having more than one?
Suggested discussion
Given what needs to happen at origins of replication, can you use logic to infer and propose for discussion some potential features that distinguish replication origins from other segments of DNA?
Elongation of Replication
The melting open of the DNA double helix and the assembling the DNA replication complex is just the first step in the process of replication. Now the process of creating a new strand actually needs to get started. Here additional challenges are encountered. The first obvious issue is that of determining which of the two strands should be copied at any replication fork (i.e. which strand will serve as a template for semi-conservative synthesis)? Are both strands equally viable alternatives? There is also the problem of actually getting the process of new strand synthesis started. Can the DNA polymerase initiate new strand on its own? The answer to the latter question, and some of the rationale and consequences, will be discussed later. The key idea to note at this point is that it has been experimentally determined that DNA polymerase can NOT initiate strand synthesis on its own. Rather, DNA polymerase requires a short stretch of double-stranded structure followed by single stranded template. The creation of a short oligonucleotide is carried out by the enzyme primase. This protein creates a short polymer of RNA (not DNA) called a primer (these are depicted by short green lines in the figures above and below) that can be used by DNA polymerase to nucleate a new growing strand.
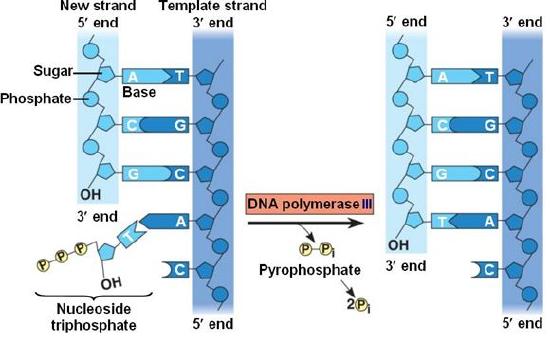
During the process of strand elongation, the DNA polymerase polymerizes a new covalently-linked strand of DNA nucleotides (in bacteria this specific enzyme may be called DNA polymerase III; in eukaryotes polymerase nomenclature is more complex and the roles of several polymerase proteins are not completely understood). It turns out that one of the strands is favored exclusively over the other to serve as a template. DNA polymerase will 'read' the template strand from 3' to 5' and synthesize a new strand in the 5' to 3' direction. Hypotheses to explain this universal observation usually center around the energetics associated with the addition of a new nucleotide and arguments associated with DNA repair, that we will describe shortly. Let us therefore briefly consider the reaction involving the addition of a single nucleotide. The primer provides an important 3' hydroxyl on which to begin synthesis. The next deoxyribonucleotide triphosphate enters the binding site of the DNA polymerase and as shown in the figure below is oriented by the polymerase such that a hydrolysis of the 5' triphosphate can occur, releasing pyrophosphate and coupling this exergonic reaction to the synthesis of a phosphodiester bond between the 5' phosphate of the incoming nucleotide and the 3' hydroxyl group of the primer. This process can be repeated until deoxyribonucleotide triphosphates run out or the replication complex falls off of the DNA. In effect, DNA polymerase adds the phosphate group (5') from the incoming nucleotide to the existing hydroxyl group (3') of the previously added nucleotide.
Correct base pairing, or selection of correct nucleotide to add at each step, is accomplished by structural constraints felt by the DNA polymerase and the energetically favorable hydrogen bonds formed between complementary nucleotides. The process is energetically driven by the hydrolysis of the incoming 5' triphosphate and the energetically favorable interactions formed by the inter-nucleotide interactions in the growing double helix (base stacking and complementary base pairing hydrogen bonds). Note that the energetics of nucleotide addition do not technically preclude a strand growing in the 3' to 5' direction, the key difference in this scheme is that the energy "source" for synthesis would need to come from a nucleotide already incorporated into the growing strand rather than the new incoming nucleotide (which this might be an important selective disadvantage is discussed briefly). After elongation has started a different DNA polymerase (in bacteria this is usually called DNA Polymerase I) comes in to remove the RNA primer and to synthesize the remaining bit of missing DNA.
As will be discussed in more detail in class, the movement of the replication fork induces winding of the DNA in both directions of replication. Another ATP consuming enzyme called topoisomerase helps to relieve this stress.
DNA polymerase catalyzes the addition of the 5' phosphate group from an incoming nucleotide to the 3' hydroxyl group of the previous nucleotide. This process creates a phosphodiester bond between the nucleotides while hydrolyzing the phosphoanhydride bond in the nucleotide.
Source: http://bio1151.nicerweb.com/Locked/m...h16/elong.html
Suggested discussion
Create an energy story for the addition of a nucleotide onto a polymer as shown in the figure above. This will be an explicit learning goal from some of your Bis2A instructors.
Leading and Lagging Strand
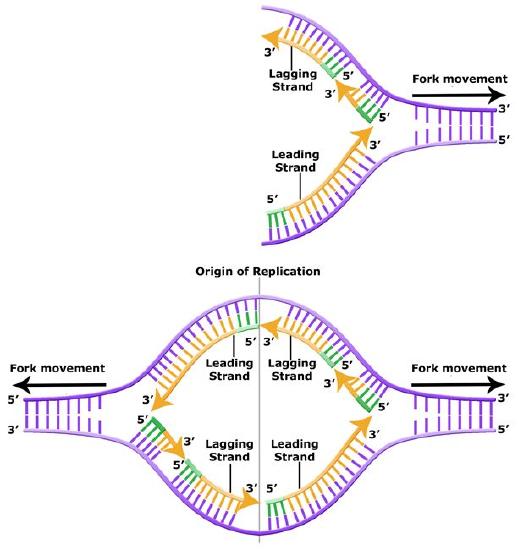
The discussion above about strand elongation describes the process of new strand synthesis if that strand happens to be synthesized in the same direction as the replication fork is or appears to be moving along the DNA. This strand can be synthesized continuously and is called the leading strand. However, both strands of the original DNA double helix must be copied. Since the DNA polymerase can only synthesize DNA in a 5' to 3' direction, the polymerization of the strand opposite of the leading strand must occur in the opposite direction that helicase, or front of the replication fork, is traveling. This strand is called the lagging strand and due to geometric constraints must be synthesized through a series of RNA priming and DNA synthesis events into short segments called Okazaki fragments. As noted, the initiation of synthesis of each Okazaki fragment requires an primase to synthesize an RNA primer and each of these RNA primers must be ultimately removed and replaced with DNA nucleotides by a different DNA polymerase. The covalent bonds between each of the Okazaki fragment can not be made by the DNA polymerase and must therefore be formed by yet another enzyme called DNA ligase. The geometry of lagging strand synthesis is difficult to visualize and will be covered in class.
The lagging strand is created in multiple segments. A replication fork showing the leading and lagging strand. A replication bubble showing the leading and lagging strands.
Bis2A Team original image
Termination of Replication
Telomeres and Telomerase
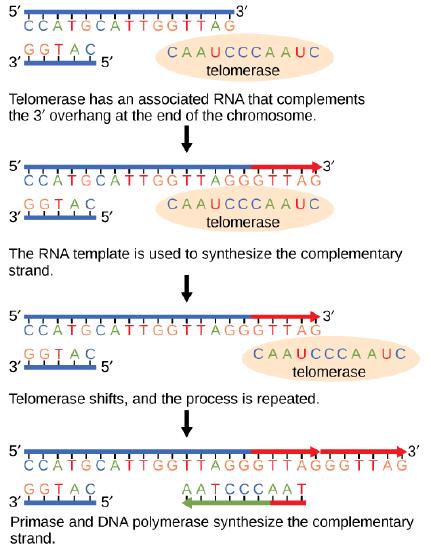
The ends of replication in circular bacterial chromosomes poses few practical problems. However, the ends of linear eukaryotic chromosomes pose a specific problem for DNA replication. Because DNA polymerase can add nucleotides in only one direction (5' to 3'), the leading strand allows for continuous synthesis until the end of the chromosome is reached; however, as the replication complex arrives at the end of the lagging strand there is no place for the primase to "land" and synthesize an RNA primer so that the synthesis of the missing lagging strand DNA fragment at the end of the chromosome can be initiated by the DNA polymerase. Without some mechanism to help fill this gap, this chromosomal end will remain unpaired and will be lost to nucleases. Over time, and several rounds of replication, this would result in the ends of linear chromosomes getting progressively shorter, ultimately compromising the ability of the organism to survive. These ends of the linear chromosomes are known as telomeres and nearly all eukaryotic species have evolved repetitive sequences that do not code for a specific gene. As a consequence, these "non-coding" telomeres act as replication buffers and are shortened with each round of DNA replication instead of critical genes. For example, in humans, a six base-pair sequence, TTAGGG, is repeated 100 to 1000 times at the end of most chromosomes. In addition to acting as a potential buffer, the discovery of the enzyme telomerase helped in the understanding of how chromosome ends are maintained. Telomerase is an enzyme composed of protein and RNA. Telomerase attaches to the end of the chromosome by complementary base pairing between the RNA component of telomerase and the DNA template. The RNA is used as a complementary strand for the short elongation of its complement. This process can be repeated numerous times. Once the lagging strand template is sufficiently elongated by telomerase, primase will create a primer followed by DNA polymerase which can now add nucleotides that are complementary to the ends of the chromosomes. Thus, the ends of the chromosomes are replicated.
The ends of linear chromosomes are maintained by the action of the telomerase enzyme.
Telomerase is not active in adult somatic cells. Adult somatic cells that undergo cell division continue to have their telomeres shortened. This essentially means that telomere shortening is associated with aging. In 2010, scientists found that telomerase can reverse some age-related conditions in mice, and this may have potential in regenerative medicine.1 Telomerase-deficient mice were used in these studies; these mice have tissue atrophy, stem-cell depletion, organ system failure, and impaired tissue injury responses. Telomerase reactivation in these mice caused extension of telomeres, reduced DNA damage, reversed neurodegeneration, and improved functioning of the testes, spleen, and intestines. Thus, telomere reactivation may have potential for treating age-related diseases in humans.
Differences in DNA Replication Rates Between Bacteria and Eukaryotes
DNA replication has been extremely well-studied in bacteria, primarily because of the small size of the genome and large number of variants available. Escherichia coli has 4.6 million base pairs in a single circular chromosome, and all of it gets replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the chromosome in both directions. This means that approximately 1000 nucleotides are added per second. The process is much more rapid than in eukaryotes. Table 1 summarizes the differences between bacterial and eukaryotic replications.
| Differences between Prokaryotic and Eukaryotic Replication | ||
|---|---|---|
| Property | Prokaryotes | Eukaryotes |
| Origin of replication | Single | Multiple |
| Rate of polymerization per polymerase | 1000 nucleotides/s | 50 to 100 nucleotides/s |
| Chromosome structure | circular | linear |
| Telomerase | Not present | Present |
Link to external resource
Click through a tutorial on DNA replication.
Replication Design Challenge: Proofreading
When the cell begins the task of replicating the DNA, it does so in response to environmental signals that tell the cell it is time to divide. The ideal goal of DNA replication is to produce two identical copies of the double-stranded DNA template and to do it in an amount of time that does not pose an unduly high evolutionarily selective cost. This is a daunting task when you consider that there are ~6,500,000,000 base pairs in the human genome and ~4,500,000 base pairs in the genome of a typical E. coli strain and that Nature has determined that the cells must replicate within 24 hours and 20 minutes, respectively. In either case many individual biochemical reactions need to take place.
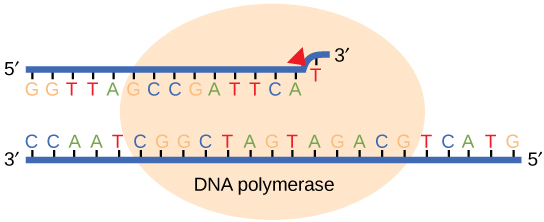
While ideally replication would happen with perfect fidelity, DNA replication, like all other biochemical processes, is imperfect - bases may be left out, extra bases added, or bases may be added that do not properly base-pair. In many organisms, many of the mistakes that occur during DNA replication are promptly corrected by DNA polymerase itself via a mechanism known as proofreading. In proofreading, the DNA polymerase "reads" each newly added base via sensing the presence or absence of small structural anomalies before adding the next base to the growing strand. In so doing, a correction can be made.
If the polymerase detects that a newly added base has paired correctly with the base in the template strand, the next nucleotide is added. If, however, a wrong nucleotide is added to the growing polymer, the misshaped double helix will cause DNA polymerase to stall, the newly made strand will be ejected from the polymerizing site on the polymerase and will enter into an exonuclease site. In this site, DNA polymerase is able to cleave off the last several nucleotides that were added to the polymer. Once the incorrect nucleotides have been removed, new ones will be added again. This proofreading capability comes with some trade-offs: Using an error correcting/more accurate polymerase requires time (the trade-off is speed of replication) and energy (always an important cost to consider). The slower you go the more accurate you can be. Going too slow, however, may keep you from replicating as fast as your competition so figuring out the balance ifs key.
Errors that are not corrected by proofreading become what are known as mutations.
Suggested discussion
Why would DNA replication need to be fast? Consider the environment the DNA is in, and compare that to the structure of DNA while being replicated
Suggested discussion
What are the pros and cons for DNA polymerases' proofreading capabilities?
Replication Mistakes and DNA Repair
Although DNA replication is typically a highly accurate process and proofreading DNA polymerases help to keep the error rate low, mistakes still occur. In addition to errors of replication environmental damage may also occur to the DNA. Such uncorrected errors of replication or environmental DNA damage may lead to serious consequences. Therefore, Nature has evolved several mechanisms for repairing damaged or incorrectly synthesized DNA.
Mismatch Repair
Some errors are not corrected during replication, but are instead corrected after replication is completed; this type of repair is known as mismatch repair. Specific enzymes recognize the incorrectly added nucleotide and excise it; replacing it with the correct base. But, how do mismatch repair enzymes recognize which of the two bases is the incorrect one?
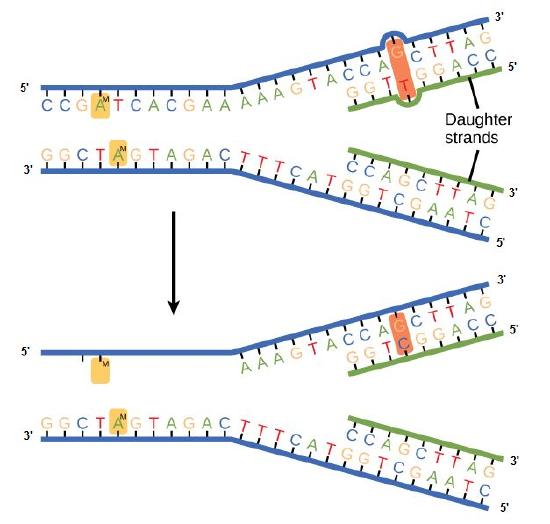
In E. coli, after replication, the nitrogenous base adenine acquires a methyl group; this means that directly after replication the parental DNA strand will have methyl groups, whereas the newly synthesized strand lacks them. Thus, mismatch repair enzymes are able to scan the DNA remove the wrongly incorporated bases from the newly synthesized, the non-methylated strand by using the methylated strand as the "correct" template from which to incorporate a new nucleotide. In eukaryotes, the mechanism is not as well understood, but it is believed to involve recognition of unsealed nicks in the new strand, as well as a short-term continuing association of some of the replication proteins with the new daughter strand after replication has completed.
In mismatch repair, the incorrectly added base is detected after replication. The mismatch repair proteins detect this base and remove it from the newly synthesized strand by nuclease action. The gap is now filled with the correctly paired base.
Nucleotide Excision Repair
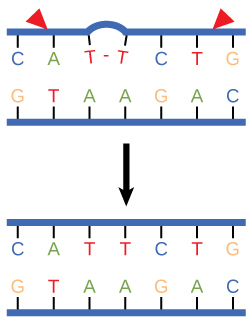
Nucleotide excision repair enzymes replace incorrect bases by making a cut on both the 3' and 5' ends of the incorrect base. The entire segment of DNA is removed and replaced with correctly paired nucleotides by the action of a DNA polymerase. Once the bases are filled in, the remaining gap is sealed with a phosphodiester linkage catalyzed by the enzyme DNA ligase. This repair mechanism is often employed when UV exposure causes the formation of pyrimidine dimers.
Nucleotide excision repairs thymine dimers. When exposed to UV, thymines lying adjacent to each other can form thymine dimers. In normal cells, they are excised and replaced.
Consequences of errors in replication, transcription and translation
Something key to think about:
Cells have evolved a variety of ways to make sure DNA errors are both detected and corrected. We have already discussed several of them. The question becomes, why? Why did so many different mechanisms evolve? From proofreading by the various DNA-dependent DNA polymerases, to the complex repair systems. Such mechanisms did not evolve for errors in transcription or translation. If you are familiar with the processes of transcription and/or translation think about what the consequences would be of an error in transcription? Would such an error effect the offspring? Would it be lethal to the cell? What errors in translation? Ask the same questions about the process of translation? What would happen if the wrong amino acid was accidentally put into the growing polypeptide during translation? How do these contrast with DNA replication? If you are not familiar with transcription or translation, don't fret. We'll learn those soon and return to this question again.