S2018_Lecture09_Reading
- Page ID
- 11395
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Introduction to bacterial and archaeal diversity
Perhaps bacteria may tentatively be regarded as biochemical experiments; owing to their relatively small size and rapid growth, variations must arise much more frequently than in more differentiated forms of life, and they can in addition afford to occupy more precarious positions in natural economy than larger organisms with more exacting requirements.
Marjory Stephenson, in Bacterial Metabolism, (1930)
Prokaryotes are single-celled organisms with neither a membrane-bound nucleus nor other lipid membrane-bound organelles. They are composed of two phylogenetically distinct groups of organisms: Bacteria and Archaea. In recent years, the term prokaryote has fallen out of favor for many microbiologists. The reason is that while bacteria and archaea share many morphological characteristics, they nevertheless, represent evolutionarily distinct domains of life. The figure below shows a simple phylogenetic tree with the three main domains of life: Bacteria, Archaea, and Eukarya. This means that the use of the term prokaryote should not be used with the intention to group the bacteria and archaea on the basis of shared evolutionary history. It is, however, convenient to use the term "prokaryote" when describing the groups of organisms that share the common morphological characteristics (i.e. no nucleus) and some of your instructors will likely do so. When you hear or use the term "prokaryote", therefore, make sure that it is not being used to or implying that the bacteria and archaea are part of the same phylogenetic group. Rather make sure that the use of the term "prokaryote" is limited to describing the common physical characteristics of these two microbial groups.

Figure 1. Although bacteria and archaea are both described as prokaryotes, they have been placed in separate domains of life. An ancestor of modern archaea is believed to have given rise to Eukarya, the third domain of life. Archaeal and bacterial phyla are shown; the exact evolutionary relationship between these phyla is still open to debate.
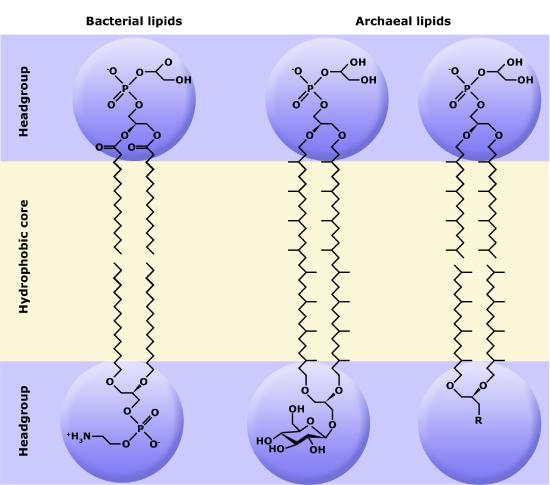
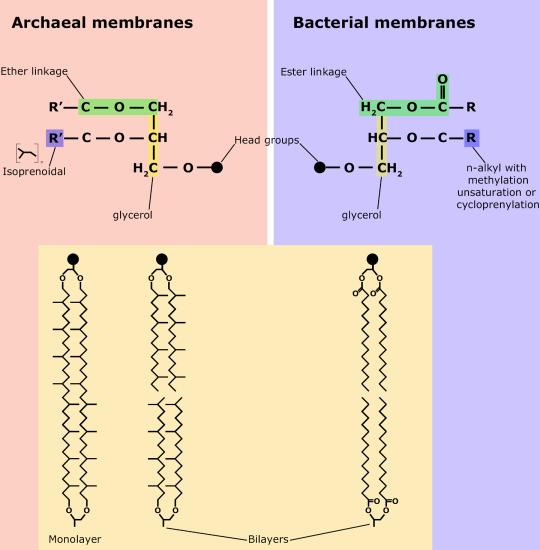
Although bacteria and archaea share many morphological, structural, and metabolic attributes, there are numerous differences between the organisms in these two clades. The most notable differences are in the chemical structure and compositions of membrane lipids, the chemical composition of the cell wall, and the makeup of the information processing machinery (e.g., replication, DNA repair, and transcription).
Bacterial and archaeal diversity
Bacteria and archaea were on Earth long before multicellular life appeared. They are ubiquitous and have highly diverse metabolic activities. This diversity allows different species within clades to inhabit every imaginable surface where there is sufficient moisture. For example, some estimates suggest that in the typical human body, bacterial cells outnumber human body cells by about ten to one. Indeed, bacteria and archaea comprise the majority of living things in all ecosystems. Certain bacterial and archaeal species can thrive in environments that are inhospitable for most other life. Bacteria and archaea, along with microbial eukaryotes, are also critical for recycling the nutrients essential for creating new biomolecules. They also drive the evolution of new ecosystems (natural or man-made).
The first inhabitants of Earth
The Earth and its moon are thought to be about 4.54 billion years old. This estimate is based on evidence from radiometric dating of meteorite material, together with other substrate material from Earth and the moon. Early Earth had a very different atmosphere (contained less molecular oxygen) than it does today and was subjected to strong radiation; thus, the first organisms would have flourished in areas where they were more protected, such as in ocean depths or beneath the Earth's surface. During this time period, strong volcanic activity was common on Earth, so it is likely that these first organisms were adapted to very high temperatures. Early Earth was also bombarded with mutagenic radiation from the sun. The first organisms, therefore, needed to be able to withstand all these harsh conditions.
So, when and where did life begin? What were the conditions on Earth when life began? What did LUCA (the Last Universal Common Ancestor), the predecessor to bacteria and archaea look like? While we don't know exactly when and how life arose and what it looked like when it did, we do have a number of hypotheses based on various biological and geological data that we briefly describe below.
The ancient atmosphere
Evidence indicates that during the first two billion years of Earth’s existence, the atmosphere was anoxic, meaning that there was no molecular oxygen. Therefore, only those organisms that can grow without oxygen—anaerobic organisms—were able to live. Autotrophic organisms that convert solar energy into chemical energy are called phototrophs, and they appeared within one billion years of the Earth's formation. Then, cyanobacteria, also known as blue-green algae, evolved from these simple phototrophs one billion years later. Cyanobacteria began oxygenating the atmosphere. Increased atmospheric oxygen allowed the development of more efficient O2-utilizing catabolic pathways. It also opened up the land to increased colonization, because some O2 is converted into O3 (ozone), and ozone effectively absorbs the ultraviolet light that would otherwise cause lethal mutations in DNA. Ultimately, the increase in O2 concentrations allowed the evolution of other life forms.
Note:
The evolution of bacteria and archaea:
How do scientists answer questions about the evolution of bacteria and archaea? Unlike with animals, artifacts in the fossil record of bacteria and archaea offer very little information. Fossils of ancient bacteria and archaea look like tiny bubbles in rock. Some scientists turn to comparative genetics which, as its name suggests, is a domain of biology that makes quantitative comparisons of the genetic information between two or more species. A core assumption in the field of comparative genetics is that the more recently two species have diverged, the more similar their genetic information will be. Conversely, species that diverged long ago will have more genes that are dissimilar. Therefore, by comparing genetic sequences between organisms can shed light on their evolutionary relationships and allow scientists to create models of what the genetic makeup of the ancestors of the organisms being compared might have looked like.
Scientists at the NASA Astrobiology Institute and at the European Molecular Biology Laboratory collaborated to analyze the molecular evolution of 32 specific proteins common to 72 species of bacteria. The model they derived from their data indicates that three important groups of bacteria—Actinobacteria, Deinococcus, and Cyanobacteria (which the authors call Terrabacteria)—were likely the first to colonize land. Organisms in the genus Deinococcus are bacteria that tend to be highly resistant to ionizing radiation. Cyanobacteria are photosynthesizers, while Actinobacteria are a group of very common bacteria that include species important in decomposition of organic wastes.
The timelines of species divergence suggest that bacteria (members of the domain Bacteria) diverged from common ancestral species between 2.5 and 3.2 billion years ago, whereas archaea diverged earlier: between 3.1 and 4.1 billion years ago. Eukarya diverged off the Archaean line later. Furthermore, there were bacteria able to grow in the anoxic environment that existed prior to the advent of cyanobacteria (about 2.6 billion years ago). These bacteria needed to be resistance to drying and to possess compounds that protect the organism from radiation. It has been proposed that the emergence of cyanobacteria with its ability to conduct photosynthesis and produce oxygen was a key event in the evolution of life on Earth.
Microbial mats
Microbial mats (large biofilms) may be representative of the earliest visible structure formed by life on Earth; there is fossil evidence of their presence starting about 3.5 billion years ago. A microbial mat is a multi-layered sheet of microbes composed mostly of bacteria but that may also include archaea. Microbial mats are a few centimeters thick, and they typically grow at the interface between two materials, mostly on moist surfaces. Organisms in a microbial mat are held together by a glue-like, sticky substance that they secrete, forming an extracellular matrix. The species within the mat carry out different metabolic activities depending on their environment. As a result, microbial mats have been identified that have different textures and colors reflecting the mat composition and the metabolic activities conducted by the microorganisms that make up the mat.
The first microbial mats likely harvested energy through redox reactions (discussed elsewhere) from chemicals found near hydrothermal vents. A hydrothermal vent is a breakage or fissure in the Earth’s surface that releases geothermally heated water. With the evolution of photosynthesis about 3 billion years ago, some organisms in microbial mats came to use a more widely available energy source—sunlight—whereas others depended on chemicals from hydrothermal vents for energy and food.

Figure 2. (a) This microbial mat, about one meter in diameter, grows over a hydrothermal vent in the Pacific Ocean in a region known as the “Pacific Ring of Fire.” Chimneys, such as the one indicated by the arrow, allow gases to escape. (b) In this micrograph, bacteria within a mat are visualized using fluorescence microscopy. (credit a: modification of work by Dr. Bob Embley, NOAA PMEL, Chief Scientist; credit b: modification of work by Ricardo Murga, Rodney Donlan, CDC; scale-bar data from Matt Russell)
Stromatolites
A stromatolite is a sedimentary structure formed when minerals precipitate out of water due to the metabolic activity of organisms in a microbial mat. Stromatolites form layered rocks made of carbonate or silicate. Although most stromatolites are artifacts from the past, there are places on Earth where stromatolites are still forming. For example, growing stromatolites have been found in the Anza-Borrego Desert State Park in San Diego County, California.

Figure 3. (a) These living stromatolites are located in Shark Bay, Australia. (b) These fossilized stromatolites, found in Glacier National Park, Montana, are nearly 1.5 billion years old. (credit a: Robert Young; credit b: P. Carrara, NPS).
Bacteria and archaea are adaptable: life in moderate and extreme environments
Some organisms have developed strategies that allow them to survive harsh conditions. Bacteria and archaea thrive in a vast array of environments: some grow in conditions that would seem very normal to us, whereas others are able to thrive and grow under conditions that would kill a plant or an animal. Almost all bacteria and archaea have some form of a cell wall, a protective structure that allows them to survive in both hyper- and hypo-osmotic conditions. Some soil bacteria are able to form endospores that resist heat and drought, thereby allowing the organism to survive until more favorable conditions recur. These adaptations, along with others, allow bacteria to be the most abundant life forms in all terrestrial and aquatic ecosystems.
Some bacteria and archaea are adapted to grow under extreme conditions and are called extremophiles, meaning “lovers of extremes.” Extremophiles have been found in all kinds of environments, such as in the depths of the oceans and the earth; in hot springs, the Artic, and the Antarctic; in very dry places; in harsh chemical environments; and in high-radiation environments, just to mention a few. These organisms help to give us a better understanding of the diversity of life and open up the possibility of finding microbial species that may lead to the discovery of new therapeutic drugs or have industrial applications. Because they have specialized adaptations that allow them to live in extreme conditions, many extremophiles cannot survive in moderate environments. There are many different groups of extremophiles. They are categorized based on the conditions in which they grow best, and several habitats are extreme in multiple ways. For example, a soda lake is both salty and alkaline, so organisms that live in a soda lake must be both alkaliphiles and halophiles. Other extremophiles, like radioresistant organisms, do not prefer an extreme environment (in this case, one with high levels of radiation) but have adapted to survive in it.
Table 1. This table lists some extremophiles and their preferred conditions.
| Extremophile Type | Conditions for Optimal Growth |
| Acidophiles | pH 3 or below |
| Alkaliphiles | pH 9 or above |
| Thermophiles | Temperature of 60–80 °C (140–176 °F) |
| Hyperthermophiles | Temperature of 80–122 °C (176–250 °F) |
| Psychrophiles | Temperature of -15 °C (5 °F) or lower |
| Halophiles | Salt concentration of at least 0.2 M |
| Osmophiles | High sugar concentration |

Figure 4. Deinococcus radiodurans, visualized in this false-color transmission electron micrograph, is a bacterium that can tolerate very high doses of ionizing radiation. It has developed DNA repair mechanisms that allow it to reconstruct its chromosome even if it has been broken into hundreds of pieces by radiation or heat. (credit: modification of work by Michael Daly; scale-bar data from Matt Russell)
Footnotes
1. Battistuzzi, FU, Feijao, A, and Hedges, SB. A genomic timescale of prokaryote evolution: Insights into the origin of methanogenesis, phototrophy, and the colonization of land. BioMed Central: Evolutionary Biology 4 (2004): 44, doi:10.1186/1471-2148-4-44.
Cellular structure of bacteria and archaea
In this section, we will discuss the basic structural features of both bacteria and archaea. There are many structural, morphological, and physiological similarities between bacteria and archaea. As discussed in the previous section, these microbes inhabit many ecological niches and carry out a great diversity of biochemical and metabolic processes. Both bacteria and archaea lack a membrane-bound nucleus and membrane-bound organelles, which are hallmarks of eukaryotes.
While Bacteria and Archaea are separate domains, morphologically they share a number of structural features. As a result, they face similar problems, such as the transport of nutrients into the cell, the removal of waste material from the cell, and the need to respond to rapid local environmental changes. In this section, we will focus on how their common cell structure allows them to thrive in various environments and simultaneously puts constraints on them. One of the biggest constraints is related to cell size.
Although bacteria and archaea come in a variety of shapes, the most common three shapes are as follows: cocci (spherical), bacilli (rod-shaped), and spirilli (spiral-shaped) (figure below). Both bacteria and archaea are generally small compared to typical eukaryotes. For example, most bacteria tend to be on the order of 0.2 to 1.0 µm (micrometers) in diameter and 1-10 µm in length. However, there are exceptions. Epulopiscium fishelsoni is a bacillus-shaped bacterium that is typically 80 µm in diameter and 200-600 µm long. Thiomargarita namibiensis is a spherical bacterium between 100 and 750 µm in diameter and is visible to the naked eye. For comparison, a typical human neutrophil is approximately 50 µm in diameter.
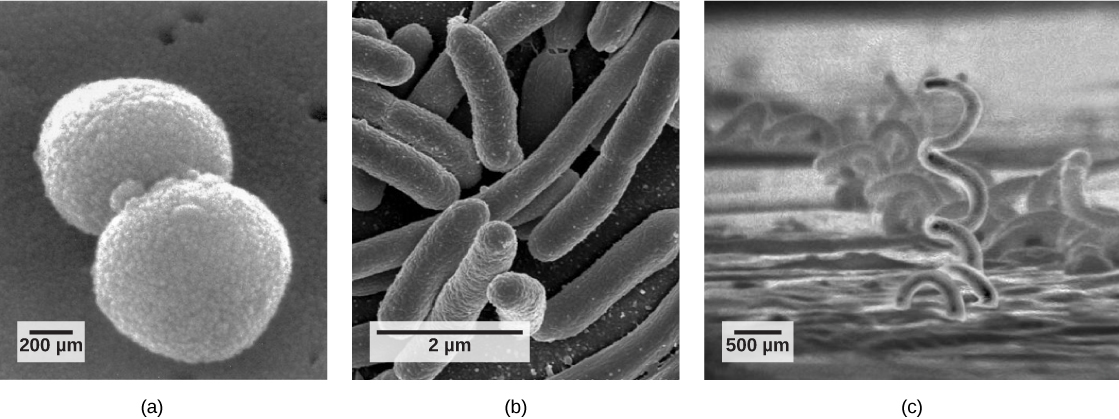

Figure 1. This figure shows the three most common shapes of bacteria and archaea: (a) cocci (spherical), (b) bacilli (rod-shaped), and (c) spirilli (spiral-shaped).
A thought question:
One question that comes to mind is why are bacteria and archaea typically so small? What are the constraints that keep them microscopic? How could bacteria such as Epulopiscium fishelsoni and Thiomargarita namibiensis overcome these constraints? Think of possible explanations or hypotheses that might answer these questions. We'll explore and develop an understanding of these questions in more detail below and in class.
The bacterial and archaeal cell: common structures
Introduction to the basic cell structure
Bacteria and archaea are unicellular organisms, which lack internal membrane-bound structures that are disconnected from the plasma membrane, a phospholipid membrane that defines the boundary between the inside and outside of the cell. In bacteria and archaea, the cytoplasmic membrane also contains all membrane-bound reactions, including those related to the electron transport chain, ATP synthase, and photosynthesis. By definition, these cells lack a nucleus. Instead, their genetic material is located in a self-defined area of the cell called the nucleoid. The bacterial and archaeal chromosome is often a single covalently closed circular double-stranded DNA molecule. However, some bacteria have linear chromosomes, and some bacteria and archaea have more than one chromosome or small non-essential circular replicating elements of DNA called plasmids. Besides the nucleoid, the next common feature is the cytoplasm (or cytosol), the "aqueous," jelly-like region encompassing the internal portion of the cell. The cytoplasm is where the soluble (non-membrane-associated) reactions occur and contains the ribosomes, the protein-RNA complex where proteins are synthesized. Finally, many bacteria and archaea also have cell walls, the rigid structural feature surrounding the plasma membrane that helps provide protection and constrain the cell shape. You should learn to create a simple sketch of a general bacterial or archaeal cell from memory.

Figure 2. The features of a typical prokaryotic cell are shown.
Constraints on the bacterial and archaeal cell
One common, almost universal, feature of bacteria and archaea is that they are small, microscopic to be exact. Even the two examples given as exceptions, Epulopiscium fishelsoni and Thiomargarita namibiensis, still face the basic constraints all bacteria and archaea face; they simply found unique strategies around the problem. So what is the largest constraint when it comes to dealing with the size of bacteria and archaea? Think about what the cell must do to survive.
Some basic requirements
So what do cells have to do to survive? They need to transform energy into a usable form. This involves making ATP, maintaining an energized membrane, and maintaining productive NAD+/NADH2 ratios. Cells also need to be able to synthesize the appropriate macromolecules (proteins, lipids, polysaccharides, etc.) and other cellular structural components. To do this, they need to be able to either make the core, key precursors for more complex molecules or get them from the environment.
Diffusion and its importance to bacteria and archaea
Movement by diffusion is passive and proceeds down the concentration gradient. For compounds to move from the outside to the inside of the cell, the compound must be able to cross the phospholipid bilayer. If the concentration of a substance is lower inside the cell than outside and it has chemical properties that allow it to move across the cell membrane, that compound will energetically tend to move into the cell. While the "real" story is a bit more complex and will be discussed in more detail later, diffusion is one of the mechanisms bacteria and archaea use to aid in the transport of metabolites.
Diffusion can also be used to get rid of some waste materials. As waste products accumulate inside the cell, their concentration rises compared to that of the outside environment, and the waste product can leave the cell. Movement within the cell works the same way: compounds will move down their concentration gradient, away from where they are synthesized to places where their concentration is low and therefore may be needed. Diffusion is a random process—the ability of two different compounds or reactants for chemical reactions to interact becomes a meeting of chance. Therefore, in small, confined spaces, random interactions or collisions can occur more frequently than they can in large spaces.
The ability of a compound to diffuse depends on the viscosity of the solvent. For example, it is a lot easier for you to move around in air than in water (think about moving around underwater in a pool). Likewise, it is easier for you to swim in a pool of water than in a pool filled with peanut butter. If you put a drop of food coloring into a glass of water, it quickly diffuses until the entire glass has changed color. Now what do you think would happen if you put that same drop of food coloring into a glass of corn syrup (very viscous and sticky)? It will take a lot longer for the glass of corn syrup to change color.
The relevance of these examples is to note that the cytoplasm tends to be very viscous. It contains many proteins, metabolites, small molecules, etc. and has a viscosity more like corn syrup than water. So, diffusion in cells is slower and more limited than you might have originally expected. Therefore, if cells rely solely on diffusion to move compounds around, what do you think happens to the efficiency of these processes as cells increase in size and their internal volumes get bigger? Is there a potential problem to getting big that is related to the process of diffusion?
So how do cells get bigger?
As you've likely concluded from the discussion above, with cells that rely on diffusion to move things around the cell—like bacteria and archaea—size does matter. So how do you suppose Epulopiscium fishelsoni and Thiomargarita namibiensis got so big? Take a look at these links, and see what these bacteria look like morphologically and structurally: Epulopiscium fishelsoni and Thiomargarita namibiensis.
Based on what we have just discussed, in order for cells to get bigger, that is, for their volume to increase, intracellular transport must somehow become independent of diffusion. One of the great evolutionary leaps was the ability of cells (eukaryotic cells) to transport compounds and materials intracellularly, independent of diffusion. Compartmentalization also provided a way to localize processes to smaller organelles, which overcame another problem caused by the large size. Compartmentalization and the complex intracellular transport systems have allowed eukaryotic cells to become very large in comparison to the diffusion-limited bacterial and archaeal cells. We'll discuss specific solutions to these challenges in the following sections.
Membranes
Plasma membranes enclose and define the borders between the inside and the outside of cells. They are typically composed of dynamic bilayers of phospholipids into which various other lipid soluble molecules and proteins have also been embedded. These bilayers are asymmetric—the outer leaf being different than the inner leaf in lipid composition and in the proteins and carbohydrates that are displayed to either the inside or outside of the cell. Various factors influence the fluidity, permeability, and various other physical properties of the membrane. These include the temperature, the configuration of the fatty acid tails (some kinked by double bonds), the presence of sterols (i.e., cholesterol) embedded in the membrane, and the mosaic nature of the proteins embedded within it. The cell membrane has selectivity; it allows only some substances through while excluding others. In addition, the plasma membrane must, in some cases, be flexible enough to allow certain cells, such as amoebae, to change shape and direction as they move through the environment, hunting smaller, single-celled organisms.
Amoebae Hunting Video
Cellular membranes
A subgoal in our "build-a-cell" design challenge is to create a boundary that separates the "inside" of the cell from the environment "outside". This boundary needs to serve multiple functions that include:
- Act as a barrier by blocking some compounds from moving in and out of the cell.
- Be selectively permeable in order to transport specific compounds into and out of the cell.
- Receive, sense, and transmit signals from the environment to inside of the cell.
- Project "self" to others by communicating identity to other nearby cells.

Figure 1. The diameter of a typical balloon is 25cm and the thickness of the plastic of the balloon of around 0.25mm. This is a 1000X difference. A typical eukaryotic cell will have a cell diameter of about 50µm and a cell membrane thickness of 5nm. This is a 10,000X difference.
Note: possible discussion
The ratio of membrane thickness compared to the size of an average eukaryotic cell is much greater compared to that of a balloon stretched with air. To think that the boundary between life and nonlife is so small, and seemingly fragile, more so than a balloon, suggests that structurally the membrane must be relatively stable. Discuss why cellular membranes are stable. You will need to pull from information we have already covered in this class.
Fluid mosaic model
The existence of the plasma membrane was identified in the 1890s, and its chemical components were identified in 1915. The principal components identified at that time were lipids and proteins. The first widely accepted model of the plasma membrane’s structure was proposed in 1935 by Hugh Davson and James Danielli; it was based on the “railroad track” appearance of the plasma membrane in early electron micrographs. They theorized that the structure of the plasma membrane resembles a sandwich, with protein being analogous to the bread, and lipids being analogous to the filling. In the 1950s, advances in microscopy, notably transmission electron microscopy (TEM), allowed researchers to see that the core of the plasma membrane consisted of a double, rather than a single, layer. A new model that better explains both the microscopic observations and the function of that plasma membrane was proposed by S.J. Singer and Garth L. Nicolson in 1972.
The explanation proposed by Singer and Nicolson is called the fluid mosaic model. The model has evolved somewhat over time, but it still best accounts for the structure and functions of the plasma membrane as we now understand them. The fluid mosaic model describes the structure of the plasma membrane as a mosaic of components—including phospholipids, cholesterol, proteins, and carbohydrates—that gives the membrane a fluid character. Plasma membranes range from 5 to 10 nm in thickness. For comparison, human red blood cells, visible via light microscopy, are approximately 8 µm wide, or approximately 1,000 times wider than a plasma membrane.

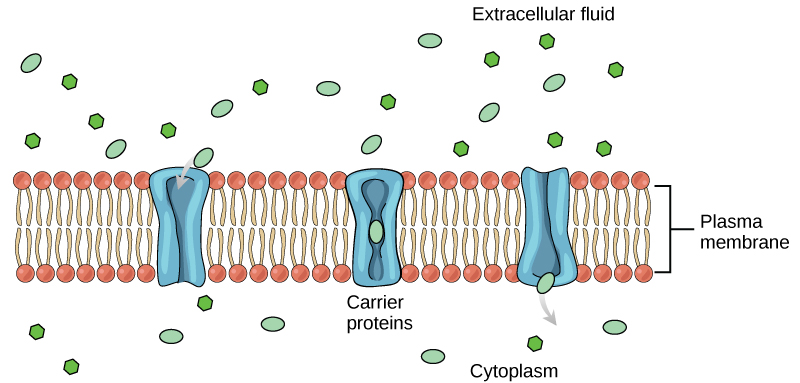
Figure 2. The fluid mosaic model of the plasma membrane describes the plasma membrane as a fluid combination of phospholipids, cholesterol, and proteins. Carbohydrates attached to lipids (glycolipids) and to proteins (glycoproteins) extend from the outward-facing surface of the membrane.
The principal components of a plasma membrane are lipids (phospholipids and cholesterol), proteins, and carbohydrates. The proportions of proteins, lipids, and carbohydrates in the plasma membrane vary with organism and cell type, but for a typical human cell, proteins account for about 50 percent of the composition by mass, lipids (of all types) account for about 40 percent of the composition by mass, and carbohydrates account for the remaining 10 percent of the composition by mass. However, the concentration of proteins and lipids varies with different cell membranes. For example, myelin, an outgrowth of the membrane of specialized cells, insulates the axons of the peripheral nerves, contains only 18 percent protein and 76 percent lipid. The mitochondrial inner membrane contains 76 percent protein and only 24 percent lipid. The plasma membrane of human red blood cells is 30 percent lipid. Carbohydrates are present only on the exterior surface of the plasma membrane and are attached to proteins, forming glycoproteins, or to lipids, forming glycolipids.
Phospholipids
Phospholipids are major constituents of the cell membrane, the outermost layer of cells. Like fats, they are composed of fatty acid chains attached to a polar head group. Specifically, there are two fatty acid tails and a phosphate group as the polar head group. The phospholipid is an amphipathic molecule, meaning it has a hydrophobic part and a hydrophilic part. The fatty acid chains are hydrophobic and cannot interact with water, whereas the phosphate-containing head group is hydrophilic and interacts with water.
Note
Make sure to note in Figure 3 that the phosphate group has an R group linked to one of the oxygen atoms. R is a variable commonly used in these types of diagrams to indicate that some other atom or molecule is bound at that position. That part of the molecule can be different in different phospholipids—and will impart some different chemistry to the whole molecule. At the moment, however, you are responsible for being able to recognize this type of molecule (no matter what the R group is) because of the common core elements—the glycerol backbone, the phosphate group, and the two hydrocarbon tails.

Figure 3. A phospholipid is a molecule with two fatty acids and a modified phosphate group attached to a glycerol backbone. The phosphate may be modified by the addition of charged or polar chemical groups. Several chemical R groups may modify the phosphate. Choline, serine, and ethanolamine are shown here. These attach to the phosphate group at the position labeled R via their hydroxyl groups.
Attribution: Marc T. Facciotti (own work)
A phospholipid bilayer forms as the basic structure of the cell membrane. The fatty acid tails of phospholipids face inside, away from water, whereas the phosphate group faces outside, hydrogen bonding with water. Phospholipids are responsible for the dynamic nature of the plasma membrane.

Figure 4. In the presence of water, some phospholipids will spontaneously arrange themselves into a micelle. The lipids will be arranged such that their polar groups will be on the outside of the micelle, and the nonpolar tails will be on the inside. A lipid bilayer can also form, a two layered sheet only a few nanometers thick. The lipid bilayer consists of two layers of phospholipids organized in a way that all the hydrophobic tails align side by side in the center of the bilayer and are surrounded by the hydrophilic head groups.
Source: Created by Erin Easlon (own work)
Note: possible discussion
Above it says that if you were to take some pure phospholipids and drop them into water that some if it would spontaneously (on its own) form into micelles. This sounds a lot like something that could be described by an energy story. Go back to the energy story rubric and try to start creating an energy story for this process—I expect that the steps involving the description of energy might be difficult at this point (we'll come back to that later) but you should be able to do at least the first three steps. You can constructively critique (politely) each other's work to create an optimized story.
Note: possible discussion
Note that the phospholipid depicted above has an R group linked to the phosphate group. Recall that this designation is generic—these can be different than the R groups on amino acids. What might be a benefit/purpose of "functionalizing" or "decorating" different lipids with different R groups? Think of the functional requirements for membranes stipulated above.
Membrane proteins
Proteins make up the second major component of plasma membranes. Integral membrane proteins are, as their name suggests, integrated completely into the membrane structure, and their hydrophobic membrane-spanning regions interact with the hydrophobic region of the the phospholipid bilayer. Single-pass integral membrane proteins usually have a hydrophobic transmembrane segment that consists of 20–25 amino acids. Some span only part of the membrane—associating with a single layer—while others stretch from one side of the membrane to the other, and are exposed on either side. This type of protein has a hydrophilic region or regions, and one or several mildly hydrophobic regions. This arrangement of regions of the protein tends to orient the protein alongside the phospholipids, with the hydrophobic region of the protein adjacent to the tails of the phospholipids and the hydrophilic region or regions of the protein protruding from the membrane and in contact with the cytosol or extracellular fluid.
Peripheral proteins are found on either the exterior or interior surfaces of membranes; and weakly or temporarily associated with the membranes. They can interact with either integral membrane proteins or simply interact weakly with the phospholipids within the membrane.

Figure 5. Integral membranes proteins may have one or more α-helices (pink cylinders) that span the membrane (examples 1 and 2), or they may have β-sheets (blue rectangles) that span the membrane (example 3). (credit: “Foobar”/Wikimedia Commons)
Carbohydrates
Carbohydrates are the third major component of plasma membranes. They are always found on the exterior surface of cells and are bound either to proteins (forming glycoproteins) or to lipids (forming glycolipids). These carbohydrate chains may consist of 2–60 monosaccharide units and can be either straight or branched. Along with peripheral proteins, carbohydrates form specialized sites on the cell surface that allow cells to recognize each other (one of the core functional requirements noted above in "cellular membranes").
Membrane fluidity
The mosaic characteristic of the membrane, described in the fluid mosaic model, helps to illustrate its nature. The integral proteins and lipids exist in the membrane as separate molecules and they "float" in the membrane, moving somewhat with respect to one another. The membrane is not like a balloon, however, in that can expand and contract dramatically; rather, it is fairly rigid and can burst if penetrated or if a cell takes in too much water. However, because of its mosaic nature, a very fine needle can easily penetrate a plasma membrane without causing it to burst, and the membrane will flow and self-seal when the needle is extracted.
The mosaic characteristics of the membrane explain some but not all of its fluidity. There are two other factors that help maintain this fluid characteristic. One factor is the nature of the phospholipids themselves. In their saturated form, the fatty acids in phospholipid tails are saturated with hydrogen atoms. There are no double bonds between adjacent carbon atoms. This results in tails that are relatively straight. By contrast, unsaturated fatty acids do not have a full complement of hydrogen atoms on their fatty acid tails, and therefore contain some double bonds between adjacent carbon atoms; a double bond results in a bend in the string of carbons of approximately 30 degrees.

Figure 6. Any given cell membrane will be composed of a combination of saturated and unsaturated phospholipids. The ratio of the two will influence the permeability and fluidity of the membrane. A membrane composed of completely saturated lipids will be dense and less fluid, and a membrane composed of completely unsaturated lipids will be very loose and very fluid.
Note: possible discussion
Organisms can be found living in extreme temperature conditions. Both in extreme cold or extreme heat. What types of differences would you expect to see in the lipid composition of organisms that live at these extremes?
Saturated fatty acids, with straight tails, are compressed by decreasing temperatures, and they will press in on each other, making a dense and fairly rigid membrane. When unsaturated fatty acids are compressed, the “kinked” tails elbow adjacent phospholipid molecules away, maintaining some space between the phospholipid molecules. This “elbow room” helps to maintain fluidity in the membrane at temperatures at which membranes with high concentrations of saturated fatty acid tails would “freeze” or solidify. The relative fluidity of the membrane is particularly important in a cold environment. Many organisms (fish are one example) are capable of adapting to cold environments by changing the proportion of unsaturated fatty acids in their membranes in response to the lowering of the temperature.
Cholesterol
Animals have an additional membrane constituent that assists in maintaining fluidity. Cholesterol, which lies alongside the phospholipids in the membrane, tends to dampen the effects of temperature on the membrane. Thus, this lipid functions as a "fluidity buffer", preventing lower temperatures from inhibiting fluidity and preventing increased temperatures from increasing fluidity too much. Thus, cholesterol extends, in both directions, the range of temperature in which the membrane is appropriately fluid and consequently functional. Cholesterol also serves other functions, such as organizing clusters of transmembrane proteins into lipid rafts.

Figure 7. Cholesterol fits between the phospholipid groups within the membrane.
Review of the components of the membrane
| The components and functions of the plasma membrane | |
|---|---|
| Component | Location |
| Phospholipid | Main fabric of the membrane |
| Cholesterol | Between phospholipids and between the two phospholipid layers of animal cells |
| Integral proteins (e.g., integrins) | Embedded within the phospholipid layer(s); may or may not penetrate through both layers |
| Peripheral proteins | On the inner or outer surface of the phospholipid bilayer; not embedded within the phospholipids |
| Carbohydrates (components of glycoproteins and glycolipids) | Generally attached to proteins on the outside membrane layer |
Transport across the membrane
Design challenge problem and subproblems
General Problem: The cell membrane must simultaneously act as a barrier between "IN" and "OUT" and control specifically which substances enter and leave the cell and how quickly and efficiently they do so.
Subproblems: The chemical properties of molecules that must enter and leave the cell are highly variable. Some subproblems associated with this are: (a) Large and small molecules or collections of molecules must be able to pass across the membrane. (b) Both hydrophobic and hydrophilic substances must have access to transport. (c) Substances must be able to cross the membrane with and against concentration gradients. (d) Some molecules look very similar (e.g. Na+ and K+) but transport mechanisms must still be able to distinguish between them.
Energy story perspective
Transport across a membrane can be considered from an energy story perspective; it is a process after all. For instance, at the beginning of the process a generic substance X may be either on the inside or outside of the cell. At the end of the process, the substance will be on the opposite side from which it started.
e.g. X(in) ---> X(out),
where in and out refer to inside the cell and outside the cell, respectively.
At the beginning the matter in the system might be a very complicated collection of molecules inside and outside of the cell but with one molecule of X more inside the cell than out. At the end, there is one more molecule of X on the outside of the cell and one less on the inside. The energy in the system at the beginning is stored largely in the molecular structures and their motions and in electrical and chemical concentration imbalances across the cell membrane. The transport of X out of the cell will not change the energies of the molecular structures significantly but it will change the energy associated with the imbalance of concentration and or charge across the membrane. That is the transport will, like all other reactions, be either exergonic or endergonic. Finally, some mechanism or sets of mechanisms of transport will need to be described.
Selective permeability
One of the great wonders of the cell membrane is its ability to regulate the concentration of substances inside the cell. These substances include: ions such as Ca2+, Na+, K+, and Cl–; nutrients including sugars, fatty acids, and amino acids; and waste products, particularly carbon dioxide (CO2), which must leave the cell.
The membrane’s lipid bilayer structure provides the first level of control. The phospholipids are tightly packed, and the membrane has a hydrophobic interior. This structure alone creates what is known as a selectively permeable barrier, one that only allows substances meeting certain physical criteria to pass through it. In the case of the cell membrane, only relatively small, nonpolar materials can move through the lipid bilayer at biologically relevant rates (remember, the lipid tails of the membrane are nonpolar).
Selective permeability of the cell membrane refers to its ability to differentiate between different types of molecules, only allowing some molecules through while blocking others. Some of this selective property stems from the intrinsic diffusion rates for different molecules across a membrane. A second factor affecting the relative rates of movement of various substances across a biological membrane is activity of various protein-based membrane transporters, both passive and active, that will be discussed in more detail in subsequent sections. First, we take on the notion of intrinsic rates of diffusion across the membrane.
Relative permeability
The fact that different substances might cross a biological membrane at different rates should be relatively intuitive. There are differences in the mosaic composition of membranes in biology and differences in the sizes, flexibility, and chemical properties of molecules so it stands to reason that the permeability rates vary. It is a complicated landscape. The permeability of a substance across a biological membrane can be measured experimentally and the rate of movement across a membrane can be reported in what are known as membrane permeability coefficients.
Membrane permeability coefficients
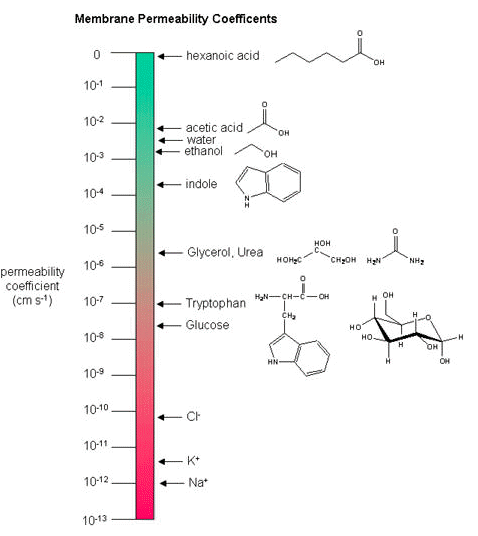
Below, a variety of compounds are plotted with respect to their membrane permeability coefficients (MPC) as measured against a simple biochemical approximation of a real biological membrane. The reported permeability coefficient for this system is the rate at which simple diffusion through a membrane occurs and is reported in units of centimeters per second (cm/s). The permeability coefficient is proportional to the partition coefficient and is inversely proportional to the membrane thickness.
It is important that you are able to read and interpret the diagram below. The larger the coefficient, the more permeable the membrane is to the solute. For example, hexanoic acid is very permeable, a MPC of 0.9; acetic acid, water, and ethanol have MPCs between 0.01 and 0.001, and they are less permeable than hexanoic acid. Where as ions, such as sodium (Na+), have an MPC of 10-12, and cross the membrane at a comparatively slow rate.
While there are certain trends or chemical properties that can be roughly associated with different compound permeability (small thing go through "fast", big things "slowly", charged things not at all etc.), we caution against over-generalizing. The molecular determinants of membrane permeability are complicated and involve numerous factors including: the specific composition of the membrane, temperature, ionic composition, hydration; the chemical properties of the solute; the potential chemical interactions between the solute in solution and in the membrane; the dielectric properties of materials; and the energy trade-offs associated with moving substances into and out of various environments. So, in this class, rather than try to apply "rules" and try to develop too many arbitrary "cut-offs", we will strive to develop a general sense of some properties that can influence permeability and leave the assignment of absolute permeability to experimentally reported rates. In addition, we will also try to minimize the use of vocabulary that depends on a frame of reference. For instance, saying that compound A diffuses "quickly" or "slowly" across a bilayer only means something if the terms "quickly" or "slowly" are numerically defined or the biological context is understood.
Energetics of transport
All substances that move through the membrane do so by one of two general methods, which are categorized based on whether or not the transport process is exergonic or endergonic. Passive transport is the exergonic movement of substances across the membrane. In contrast, active transport is the endergonic movement of substances across the membrane that is coupled to an exergonic reaction.
Passive transport
Passive transport does not require the cell to expend energy. In passive transport, substances move from an area of higher concentration to an area of lower concentration, down their concentration gradient . Depending on the chemical nature of the substance, different processes may be associated with passive transport.
Diffusion
Diffusion is a passive process of transport. A single substance tends to move from an area of high concentration to an area of low concentration until the concentration is equal across a space. You are familiar with diffusion of substances through the air. For example, think about someone opening a bottle of ammonia in a room filled with people. The ammonia gas is at its highest concentration in the bottle; its lowest concentration is at the edges of the room. The ammonia vapor will diffuse, or spread away, from the bottle; gradually, more and more people will smell the ammonia as it spreads. Materials move within the cell’s cytosol by diffusion, and certain materials move through the plasma membrane by diffusion.
Factors that affect diffusion
If unconstrained, molecules will move through and explore space randomly at a rate that depends on their size, their shape, their environment, and their thermal energy. This type of movement underlies the diffusive movement of molecules through whatever medium they are in. The absence of a concentration gradient does not mean that this movement will stop, just that there may be no net movement of the number of molecules from one area to another, a condition known as dynamic equilibrium.
Factors influencing diffusion include:
- Extent of the concentration gradient: The greater the difference in concentration, the more rapid the diffusion. The closer the distribution of the material gets to equilibrium, the slower the rate of diffusion becomes.
- Shape, size and mass of the molecules diffusing: Large and heavier molecules move more slowly; therefore, they diffuse more slowly. The reverse is typically true for smaller, lighter molecules.
- Temperature: Higher temperatures increase the energy and therefore the movement of the molecules, increasing the rate of diffusion. Lower temperatures decrease the energy of the molecules, thus decreasing the rate of diffusion.
- Solvent density: As the density of a solvent increases, the rate of diffusion decreases. The molecules slow down because they have a more difficult time getting through the denser medium. If the medium is less dense, rates of diffusion increase. Since cells primarily use diffusion to move materials within the cytoplasm, any increase in the cytoplasm’s density will decrease the rate at which materials move in the cytoplasm.
- Solubility: As discussed earlier, nonpolar or lipid-soluble materials pass through plasma membranes more easily than polar materials, allowing a faster rate of diffusion.
- Surface area and thickness of the plasma membrane: Increased surface area increases the rate of diffusion, whereas a thicker membrane reduces it.
- Distance traveled: The greater the distance that a substance must travel, the slower the rate of diffusion. This places an upper limitation on cell size. A large, spherical cell will die because nutrients or waste cannot reach or leave the center of the cell, respectively. Therefore, cells must either be small in size, as in the case of many prokaryotes, or be flattened, as with many single-celled eukaryotes.
Facilitated transport
In facilitated transport, also called facilitated diffusion, materials diffuse across the plasma membrane with the help of membrane proteins. A concentration gradient exists that allows these materials to diffuse into or out of the cell without expending cellular energy. In the case that the materials are ions or polar molecules (compounds that are repelled by the hydrophobic parts of the cell membrane), facilitated transport proteins help shield these materials from the repulsive force of the membrane, allowing them to diffuse into the cell.
Note: possible discussion
Compare and contrast passive diffusion and facilitated diffusion.
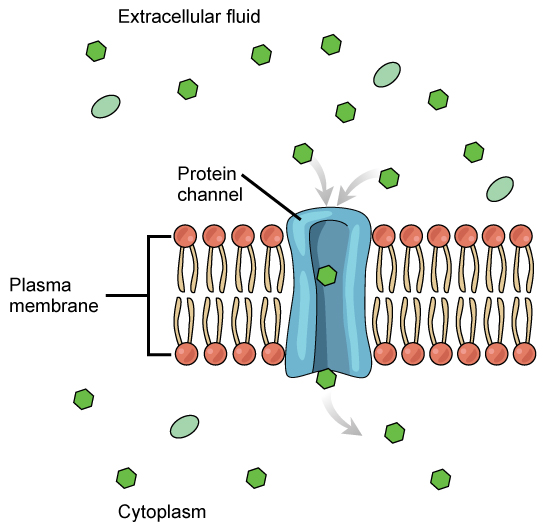
Channels
The integral proteins involved in facilitated transport are collectively referred to as transport proteins, and they function as either channels for the material or carriers. In both cases, they are transmembrane proteins. Different channel proteins have different transport properties. Some have evolved to have very high specificity for the substance that is being transported while others transport a variety of molecules sharing some common characteristic(s). The interior "passageway" of channel proteins have evolved to provide a low energetic barrier for transport of substances across the membrane through the complementary arrangement of amino acid functional groups (of both backbone and side-chains). Passage through the channel allows polar compounds to avoid the nonpolar central layer of the plasma membrane that would otherwise slow or prevent their entry into the cell. While at any one time significant amounts of water crosses the membrane both in and out, the rate of individual water molecule transport may not be fast enough to adapt to changing environmental conditions. For such cases Nature has evolved a special class of membrane proteins called aquaporins that allow water to pass through the membrane at a very high rate.
Channel proteins are either open at all times or they are “gated.” The latter controls the opening of the channel. Various mechanisms may be involved in the gating mechanism. For instance, the attachment of a specific ion or small molecule to the channel protein may trigger opening. Changes in local membrane "stress" or changes in voltage across the membrane may also be triggers to open or close a channel.
Different organisms and tissues in multicellular species express different sets of channel proteins in their membranes depending on the environments they live in or specialized function they play in an organisms. This provides each type of cell with a unique membrane permeability profile that is evolved to complement its "needs" (note the anthropomorphism). For example, in some tissues, sodium and chloride ions pass freely through open channels, whereas in other tissues a gate must be opened to allow passage. This occurs in the kidney, where both forms of channels are found in different parts of the renal tubules. Cells involved in the transmission of electrical impulses, such as nerve and muscle cells, have gated channels for sodium, potassium, and calcium in their membranes. Opening and closing of these channels changes the relative concentrations on opposing sides of the membrane of these ions, resulting a change in electrical potential across the membrane that lead to message propagation in the case of nerve cells or in muscle contraction in the case of muscle cells.
Carrier proteins
Another type of protein embedded in the plasma membrane is a carrier protein. This aptly named protein binds a substance and, in doing so, triggers a change of its own shape, moving the bound molecule from the outside of the cell to its interior; depending on the gradient, the material may move in the opposite direction. Carrier proteins are typically specific for a single substance. This selectivity adds to the overall selectivity of the plasma membrane. The molecular-scale mechanism of function for these proteins remains poorly understood.
Carrier protein play an important role in the function of kidneys. Glucose, water, salts, ions, and amino acids needed by the body are filtered in one part of the kidney. This filtrate, which includes glucose, is then reabsorbed in another part of the kidney with the help of carrier proteins. Because there are only a finite number of carrier proteins for glucose, if more glucose is present in the filtrate than the proteins can handle, the excess is not reabsorbed and it is excreted from the body in the urine. In a diabetic individual, this is described as “spilling glucose into the urine.” A different group of carrier proteins called glucose transport proteins, or GLUTs, are involved in transporting glucose and other hexose sugars through plasma membranes within the body.
Channel and carrier proteins transport material at different rates. Channel proteins transport much more quickly than do carrier proteins. Channel proteins facilitate diffusion at a rate of tens of millions of molecules per second, whereas carrier proteins work at a rate of a thousand to a million molecules per second.
Active transport
Active transport mechanisms require the use of the cell’s energy, usually in the form of adenosine triphosphate (ATP). If a substance must move into the cell against its concentration gradient—that is, if the concentration of the substance inside the cell is greater than its concentration in the extracellular fluid (and vice versa)—the cell must use energy to move the substance. Some active transport mechanisms move small-molecular weight materials, such as ions, through the membrane. Other mechanisms transport much larger molecules.
Moving against a gradient
To move substances against a concentration or electrochemical gradient, the cell must use energy. This energy is harvested from ATP generated through the cell’s metabolism. Active transport mechanisms, collectively called pumps, work against electrochemical gradients. Small substances constantly pass through plasma membranes. Active transport maintains concentrations of ions and other substances needed by living cells in the face of these passive movements. Much of a cell’s supply of metabolic energy may be spent maintaining these processes. (Most of a red blood cell’s metabolic energy is used to maintain the imbalance between exterior and interior sodium and potassium levels required by the cell.) Because active transport mechanisms depend on a cell’s metabolism for energy, they are sensitive to many metabolic poisons that interfere with the supply of ATP.
Two mechanisms exist for the transport of small-molecular weight material and small molecules. Primary active transport moves ions across a membrane and creates a difference in charge across that membrane, which is directly dependent on ATP. Secondary active transport describes the movement of material that is due to the electrochemical gradient established by primary active transport that does not directly require ATP.
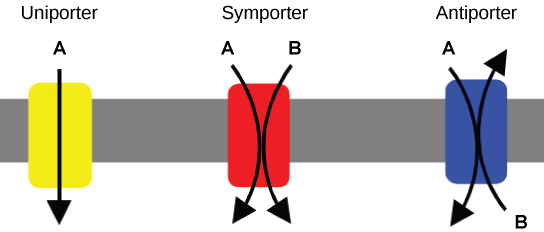
Carrier proteins for active transport
An important membrane adaption for active transport is the presence of specific carrier proteins or pumps to facilitate movement: there are three types of these proteins or transporters. A uniporter carries one specific ion or molecule. A symporter carries two different ions or molecules, both in the same direction. An antiporter also carries two different ions or molecules, but in different directions. All of these transporters can also transport small, uncharged organic molecules like glucose. These three types of carrier proteins are also found in facilitated diffusion, but they do not require ATP to work in that process. Some examples of pumps for active transport are Na+-K+ ATPase, which carries sodium and potassium ions, and H+-K+ ATPase, which carries hydrogen and potassium ions. Both of these are antiporter carrier proteins. Two other carrier proteins are Ca2+ATPase and H+ ATPase, which carry only calcium and only hydrogen ions, respectively. Both are pumps.
Primary active transport
In primary active transport, the energy is often - though not exclusively - derived directly from the hydrolysis of ATP. Often, primary active transport, such as that shown below, which functions to transport sodium and potassium ions allows secondary active transport to occur (discussed in the section below). The second transport method is still considered active because it depends on the use of energy from the primary transport.
One of the most important pumps in animal cells is the sodium-potassium pump (Na+-K+ ATPase), which maintains the electrochemical gradient (and the correct concentrations of Na+and K+) in living cells. The sodium-potassium pump moves K+ into the cell while moving Na+ out at the same time, at a ratio of three Na+ for every two K+ ions moved in. The Na+-K+ATPase exists in two forms depending on its orientation to the interior or exterior of the cell and its affinity for either sodium or potassium ions. The process consists of the following six steps.
- With the enzyme oriented towards the interior of the cell, the carrier has a high affinity for sodium ions. Three ions bind to the protein.
- ATP is hydrolyzed by the protein carrier and a low-energy phosphate group attaches to it.
- As a result, the carrier changes shape and re-orients itself towards the exterior of the membrane. The protein’s affinity for sodium decreases and the three sodium ions leave the carrier.
- The shape change increases the carrier’s affinity for potassium ions, and two such ions attach to the protein. Subsequently, the low-energy phosphate group detaches from the carrier.
- With the phosphate group removed and potassium ions attached, the carrier protein repositions itself towards the interior of the cell.
- The carrier protein, in its new configuration, has a decreased affinity for potassium, and the two ions are released into the cytoplasm. The protein now has a higher affinity for sodium ions, and the process starts again.
Several things have happened as a result of this process. At this point, there are more sodium ions outside of the cell than inside and more potassium ions inside than out. For every three ions of sodium that move out, two ions of potassium move in. This results in the interior being slightly more negative relative to the exterior. This difference in charge is important in creating the conditions necessary for the secondary process. The sodium-potassium pump is, therefore, an electrogenic pump (a pump that creates a charge imbalance), creating an electrical imbalance across the membrane and contributing to the membrane potential.
Link to learning
Visit the site to see a simulation of active transport in a sodium-potassium ATPase.
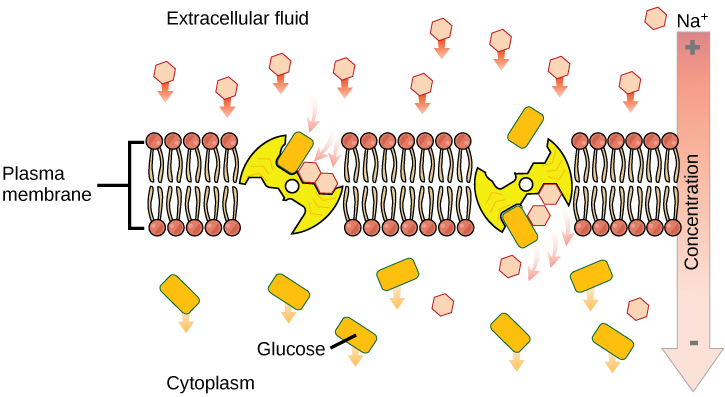
Secondary active transport (co-transport)
Secondary active transport brings sodium ions, and possibly other compounds, into the cell. As sodium ion concentrations build outside of the plasma membrane because of the action of the primary active transport process, an electrochemical gradient is created. If a channel protein exists and is open, the sodium ions will be pulled through the membrane. This movement is used to transport other substances that can attach themselves to the transport protein through the membrane. Many amino acids, as well as glucose, enter a cell this way. This secondary process is also used to store high energy hydrogen ions in the mitochondria of plant and animal cells for the production of ATP. The potential energy that accumulates in the stored hydrogen ions is translated into kinetic energy as the ions surge through the channel protein ATP synthase, and that energy is used to convert ADP into ATP.
Osmosis
Osmosis is the movement of water through a semipermeable membrane according to the concentration gradient of water across the membrane, which is inversely proportional to the concentration of solutes. While diffusion transports material across membranes and within cells, osmosis transports only water across a membrane and the membrane limits the diffusion of solutes in the water. Not surprisingly, the aquaporins that facilitate water movement play a large role in osmosis, most prominently in red blood cells and the membranes of kidney tubules.
Mechanism
Osmosis is a special case of diffusion. Water, like other substances, moves from an area of high concentration to one of low concentration. An obvious question is what makes water move at all? Imagine a beaker with a semipermeable membrane separating the two sides or halves. On both sides of the membrane the water level is the same, but there are different concentrations of a dissolved substance, or solute, that cannot cross the membrane (otherwise the concentrations on each side would be balanced by the solute crossing the membrane). If the volume of the solution on both sides of the membrane is the same, but the concentrations of solute are different, then there are different amounts of water, the solvent, on either side of the membrane.
To illustrate this, imagine two full glasses of water. One has a single teaspoon of sugar in it, whereas the second one contains one-quarter cup of sugar. If the total volume of the solutions in both cups is the same, which cup contains more water? Because the large amount of sugar in the second cup takes up much more space than the teaspoon of sugar in the first cup, the first cup has more water in it.
Returning to the beaker example, recall that it has a mixture of solutes on either side of the membrane. A principle of diffusion is that the molecules move around and will spread evenly throughout the medium if they can. However, only the material capable of getting through the membrane will diffuse through it. In this example, the solute cannot diffuse through the membrane, but the water can. Water has a concentration gradient in this system. Thus, water will diffuse down its concentration gradient, crossing the membrane to the side where it is less concentrated. This diffusion of water through the membrane—osmosis—will continue until the concentration gradient of water goes to zero or until the hydrostatic pressure of the water balances the osmotic pressure. Osmosis proceeds constantly in living systems.
Tonicity
Tonicity describes how an extracellular solution can change the volume of a cell by affecting osmosis. A solution's tonicity often directly correlates with the osmolarity of the solution. Osmolarity describes the total solute concentration of the solution. A solution with low osmolarity has a greater number of water molecules relative to the number of solute particles; a solution with high osmolarity has fewer water molecules with respect to solute particles. In a situation in which solutions of two different osmolarities are separated by a membrane permeable to water, though not to the solute, water will move from the side of the membrane with lower osmolarity (and more water) to the side with higher osmolarity (and less water). This effect makes sense if you remember that the solute cannot move across the membrane, and thus the only component in the system that can move—the water—moves along its own concentration gradient. An important distinction that concerns living systems is that osmolarity measures the number of particles (which may be molecules) in a solution. Therefore, a solution that is cloudy with cells may have a lower osmolarity than a solution that is clear if the second solution contains more dissolved molecules than there are cells.
Hypotonic solutions
Three terms—hypotonic, isotonic, and hypertonic—are used to relate the osmolarity of a cell to the osmolarity of the extracellular fluid that contains the cells. In a hypotonic situation, the extracellular fluid has lower osmolarity than the fluid inside the cell, and water enters the cell (in living systems, the point of reference is always the cytoplasm, so the prefix hypo- means that the extracellular fluid has a lower concentration of solutes, or a lower osmolarity, than the cell cytoplasm). It also means that the extracellular fluid has a higher concentration of water in the solution than does the cell. In this situation, water will follow its concentration gradient and enter the cell.
Hypertonic solutions
As for a hypertonic solution, the prefix hyper- refers to the extracellular fluid having a higher osmolarity than the cell’s cytoplasm; therefore, the fluid contains less water than the cell does. Because the cell has a relatively higher concentration of water, water will leave the cell.
Isotonic solutions
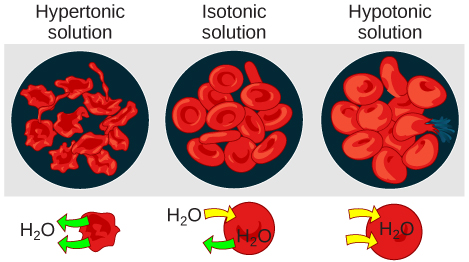
In an isotonic solution, the extracellular fluid has the same osmolarity as the cell. If the osmolarity of the cell matches that of the extracellular fluid, there will be no net movement of water into or out of the cell, although water will still move in and out. Blood cells and plant cells in hypertonic, isotonic, and hypotonic solutions take on characteristic appearances.
Connection
A doctor injects a patient with what the doctor thinks is an isotonic saline solution. The patient dies, and an autopsy reveals that many red blood cells have been destroyed. Do you think the solution the doctor injected was really isotonic?
Link to learning
For a video illustrating the process of diffusion in solutions, visit this site.
Tonicity in living systems
In a hypotonic environment, water enters a cell, and the cell swells. In an isotonic condition, the relative concentrations of solute and solvent are equal on both sides of the membrane. There is no net water movement; therefore, there is no change in the size of the cell. In a hypertonic solution, water leaves a cell and the cell shrinks. If either the hypo- or hyper- condition goes to excess, the cell’s functions become compromised, and the cell may be destroyed.
A red blood cell will burst, or lyse, when it swells beyond the plasma membrane’s capability to expand. Remember, the membrane resembles a mosaic, with discrete spaces between the molecules composing it. If the cell swells, and the spaces between the lipids and proteins become too large, and the cell will break apart.
In contrast, when excessive amounts of water leave a red blood cell, the cell shrinks, or crenates. This has the effect of concentrating the solutes left in the cell, making the cytosol denser and interfering with diffusion within the cell. The cell’s ability to function will be compromised and may also result in the death of the cell.
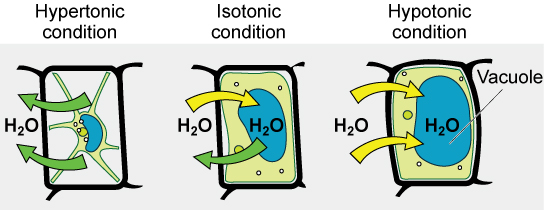
Various living things have ways of controlling the effects of osmosis—a mechanism called osmoregulation. Some organisms, such as plants, fungi, bacteria, and some protists, have cell walls that surround the plasma membrane and prevent cell lysis in a hypotonic solution. The plasma membrane can only expand to the limit of the cell wall, so the cell will not lyse. In fact, the cytoplasm in plants is always slightly hypertonic to the cellular environment, and water will always enter a cell if water is available. This inflow of water produces turgor pressure, which stiffens the cell walls of the plant. In nonwoody plants, turgor pressure supports the plant. Conversely, if the plant is not watered, the extracellular fluid will become hypertonic, causing water to leave the cell. In this condition, the cell does not shrink because the cell wall is not flexible. However, the cell membrane detaches from the wall and constricts the cytoplasm. This is called plasmolysis. Plants lose turgor pressure in this condition and wilt.
Tonicity is a concern for all living things. For example, paramecia and amoebas, which are protists that lack cell walls, have contractile vacuoles. This vesicle collects excess water from the cell and pumps it out, keeping the cell from bursting as it takes on water from its environment.
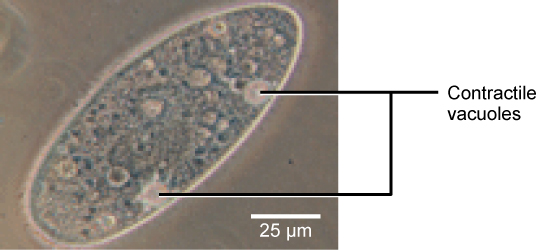
Figure 12. A paramecium’s contractile vacuole, here visualized using bright field light microscopy at 480x magnification, continuously pumps water out of the organism’s body to keep it from bursting in a hypotonic medium. (credit: modification of work by NIH; scale-bar data from Matt Russell)
Many marine invertebrates have internal salt levels matched to their environments, making them isotonic with the water in which they live. Fish, however, must spend approximately five percent of their metabolic energy maintaining osmotic homeostasis. Freshwater fish live in an environment that is hypotonic to their cells. These fish actively take in salt through their gills and excrete diluted urine to rid themselves of excess water. Saltwater fish live in the reverse environment, which is hypertonic to their cells, and they secrete salt through their gills and excrete highly concentrated urine.
In vertebrates, the kidneys regulate the amount of water in the body. Osmoreceptors are specialized cells in the brain that monitor the concentration of solutes in the blood. If the levels of solutes increase beyond a certain range, a hormone is released that retards water loss through the kidney and dilutes the blood to safer levels. Animals also have high concentrations of albumin, which is produced by the liver, in their blood. This protein is too large to pass easily through plasma membranes and is a major factor in controlling the osmotic pressures applied to tissues.