2.3: Meiosis
- Page ID
- 8618
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Sexual Reproduction
Sexual reproduction was an early evolutionary innovation after the appearance of eukaryotic cells. The fact that most eukaryotes reproduce sexually is evidence of its evolutionary success. In many animals, it is the only mode of reproduction. And yet, scientists recognize some real disadvantages to sexual reproduction. On the surface, offspring that are genetically identical to the parent may appear to be more advantageous. If the parent organism is successfully occupying a habitat, offspring with the same traits would be similarly successful. There is also the obvious benefit to an organism that can produce offspring by asexual budding, fragmentation, or asexual eggs. These methods of reproduction do not require another organism of the opposite sex. There is no need to expend energy finding or attracting a mate. That energy can be spent on producing more offspring. Indeed, some organisms that lead a solitary lifestyle have retained the ability to reproduce asexually. In addition, in asexual populations every individual is capable of reproduction. In contrast, the males in sexual populations (half the population) are not producing offspring themselves. Because of this, an asexual population can in theory grow twice as fast as a sexual population. This means that in competition, the asexual population would have the advantage. All of these advantages to asexual reproduction, which are also disadvantages to sexual reproduction, should mean that the number of species with asexual reproduction should be more common.
However, multicellular organisms that exclusively depend on asexual reproduction are rare.
So why is sexual reproduction so common?
This is one of the important questions in evolutionary biology and has been the focus of much research from the latter half of the twentieth century until now. A likely explanation is that the variation that sexual reproduction creates among offspring is very important to the survival and reproduction of the species. The only source of genetic variation in asexual organisms is mutation. In sexually reproducing organisms, mutations are continually reshuffled between generations when parents combine their unique genomes, and the alleles are mixed into different combinations by the process of meiosis and fertilization. This allows natural selection to act on a wide variety of combinations of alleles- not just as heterozygous or homozygous versions of alleles at a single gene, but of various combinations of alleles at different genes.
Muller's Ratchet
To better understand the benefits of sexual reproduction vs. asexual reproduction, Herman Joseph Muller proposed the following model:
- Mutations are inevitable.
- We know that random mutations are more frequently deleterious (have a negative influence on survival and reproduction rate) than beneficial (have a positive influence on survival and reproduction rate). That's just the nature of things- it's easier to accidentally break something than to accidentally improve it.
- Therefore, as the generations proceed, deleterious mutations will occur more frequently than beneficial ones. Even chromosomes that acquire rare beneficial mutations will be "dragged down", in terms of adaptive advantage, by rapidly accumulating dysfunctional alleles that are also occurring on that same chromosome.
- Meiotic recombination can separate (through fortuitous crossing over) rare beneficial alleles from deleterious alleles on the same chromosome, random assortment of chromosomes can separate beneficial alleles from poor ones in the same parental genome, and fertilization can allow (again through luck) combinations of weak alleles to be selected against, and combinations of beneficial alleles to be selected for.
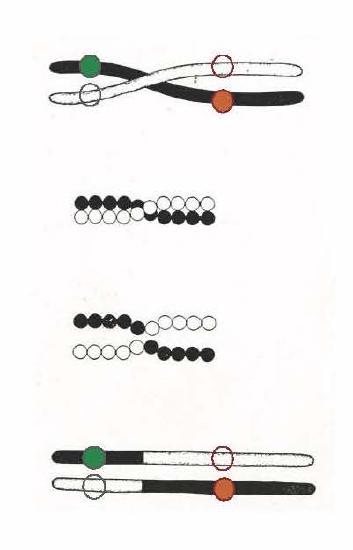
credit: "Scheme to illustrate a method of crossing-over", Thomas Hunt Morgan, 1916 "A critique of the Theory of Evolution" Morgan was the first to present convincing evidence that chromosomes carried genes.
An evolutionary benefit of recombination. In the illustration above, we see two homologous chromosomes (black vs. white) from a diploid organism. Homologous chromosomes carry the same genes, but perhaps carry different alleles for many of those genes. The white chromosome is "wild-type" with traditional, commonly found alleles at two different genes (open circles indicate their location on the chromosome). The black chromosome carries two new but recessive alleles. The solid red allele is deleterious (= bad for reproductive fitness). The solid green allele is beneficial- it promotes reproductive success. In individuals carrying the black chromosome the positive aspects of the good alleles are canceled out by the negative allele, and vice versa. Thus natural selection cannot act to decrease the frequency (in the population's gene pool) of the bad allele or to increase the frequency of the good allele. Can you see how meiotic recombination allows natural selection to influence the frequency of helpful vs. deleterious alleles?
The Red Queen Hypothesis
There is no question that sexual reproduction provides evolutionary advantages to organisms that employ this mechanism to produce offspring. The problematic question is why, even in the face of seemingly stable conditions, sexual reproduction persists when it is more difficult and produces fewer offspring for individual organisms? Variation is the outcome of sexual reproduction, but why is ongoing variation necessary? Enter the Red Queen hypothesis, first proposed by Leigh Van Valen in 1973.1 The concept was named in reference to the Red Queen's race in Lewis Carroll's book, Through the Looking-Glass, in which the Red Queen says one must run at full speed just to stay where one is.
All species coevolve with other organisms. For example, predators coevolve with their prey, and parasites coevolve with their hosts. A remarkable example of coevolution between predators and their prey is the unique coadaptation of night flying bats and their moth prey. Bats find their prey by emitting high-pitched clicks, but moths have evolved simple ears to hear these clicks so they can avoid the bats. The moths have also adapted behaviors, such as flying away from the bat when they first hear it, or dropping suddenly to the ground when the bat is upon them. Bats have evolved “quiet” clicks in an attempt to evade the moth’s hearing. Some moths have evolved the ability to respond to the bats’ clicks with their own clicks as a strategy to confuse the bats echolocation abilities.
Each tiny advantage gained by favorable variation gives a species an edge over close competitors, predators, parasites, or even prey. The only method that will allow a coevolving species to keep its own share of the resources is also to continually improve its ability to survive and produce offspring. As one species gains an advantage, other species must also develop an advantage or they will be outcompeted. Species whose individuals cannot keep up become extinct. The Red Queen’s catchphrase was, “It takes all the running you can do to stay in the same place.” This is an apt description of coevolution between competing species.
Meiosis
Sexual reproduction requires fertilization, the union of two cells from two individual organisms. If those two cells each contain one set of chromosomes, then the resulting cell contains two sets of chromosomes. Haploid cells contain one set of chromosomes, diploid cells contain two sets of chromosomes. The number of sets of chromosomes in a cell is called its ploidy level. If the reproductive cycle is to continue, then the diploid cell must somehow reduce its number of chromosome sets before fertilization can occur again, or there will be a continual doubling in the number of chromosome sets in every generation. So, in addition to fertilization, sexual reproduction includes a nuclear division that reduces the number of chromosome sets.
The nuclear division that forms haploid cells, which is called meiosis, is related to mitosis. In mitosis, both the parent and the daughter nuclei are at the same ploidy level—diploid for most plants and animals. Meiosis employs many of the same mechanisms as mitosis. However, the starting nucleus is always diploid and the nuclei that result at the end of a meiotic cell division are haploid. To achieve this reduction in chromosome number, a meiotic cell cycle consists of one round of chromosome duplication (S phase, just like mitosis) and two rounds of nuclear division. Because the events that occur during each of the division stages are analogous to the events of mitosis, the same stage names are assigned. However, because there are two rounds of division, the major process and the stages are designated with a “I” or a “II.” Thus, meiosis I is the first round of meiotic division and consists of prophase I, prometaphase I, and so on. Meiosis II, in which the second round of meiotic division takes place, includes prophase II, prometaphase II, and so on.
Meiosis I
Meiosis is preceded by an interphase consisting of the G1, S, and G2 phases, which are nearly identical to the phases preceding mitosis.
Prophase I
Early in prophase I, before the chromosomes can be seen clearly microscopically, homologous chromosomes are attached at their tips (their telomeres) to the nuclear envelope by proteins. Homologous chromosomes are similar but not identical chromosomes. For example, chromosome 12 from your mother and chromosome 12 from your father will both be present inside each of your cells. Each chromosome 12 contains the same genes, usually in the same locations, however, each gene can be a different allele. Gene A on chromosome 12 from your mother may be allele A and gene A on chromosome 12 from your father may be allele a. In species such as humans, even though the X and Y sex chromosomes are not homologous (most of their genes differ), they have a small region of homology that allows the X and Y chromosomes to pair up during prophase I. It will be very important to understand what homologous chromosomes are when following the process of meiosis.
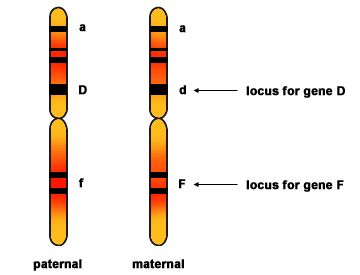
Two homologous chromosomes are shown prior to DNA replication. Each chromosome has three genes with their loci (locations on the chromosome) marked. Homologous chromosomes contain the same genes but are not identical. They each can contain different alleles of each gene.
Source: http://mrphome.net/mrp/Homologous_Chromosome.html
Remember that the meiotic M phase, like mitotic M phase, begins with a replicated, diploid genome. Early in meiotic prophase, DNA double strand breaks (DSBs) - hundreds of them- are intentionally induced by the cell, at somewhat random positions on all chromosomes. We will discuss repair of DSBs in more detail in a later module, but suffice it to say that one mode of repair for DSBs is to search for an unbroken copy that sequence, essentially taking the DSB on a hunt through the genome, in search of the lost information (which can then be copied). Normally, in a G2 cell, this information is easily located on the sister chromatid, which is very near to its recently replicated sister (thanks to the slow diffusion of DNA, and the actions of cohesins). In meiosis, however, the sister chromatid is excluded as a template for repair, forcing the break to go on a hunt for its homolog, which of course carries the identical, or near-identical, sequence. As the breaks find their homologous sequences, the homologous chromosomes are thereby aligned. Thus meiosis takes advantage of an existing repair pathway to force homologs to find each other and align- by intentionally inducing DNA damage! A protein complex called the synaptonemal complex than zips the homologs together (see figure below). At the same time, virtually all the breaks are repaired by copying information from the unbroken homolog, and then using this information to seal both ends of the break together without any loss of information at the break site.
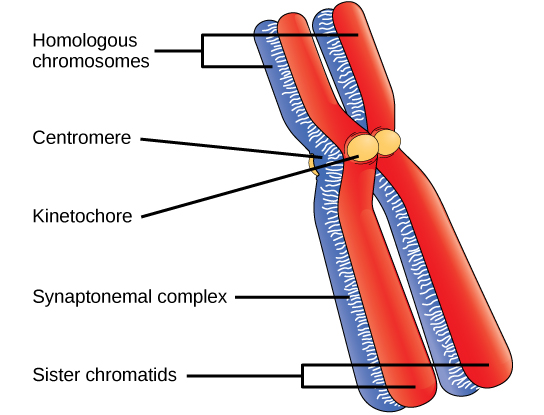
In prophase I, homologous chromosomes synapse. The homologs (one red, one blue) are bound tightly together and in alignment by a protein lattice called a synaptonemal complex, while the sisters are held together by cohesins along their length (not shown!). Please note that the chromatin is still very condensed, and forms large loops emanating from the synaptonemal complex. Also note that the synaptonemal complex, having helped to align the homologs will disappear soon- its job is done.
A question about the illustration above: note that the kinetochores of the sister chromatids are both pointing in the same direction. You might recall that in mitosis we discussed how important it is that they face in opposite directions. Why was this important? Is this an error in the diagram? Defend your answer.
Many are called, but few are chosen
A very small minority of the breaks- about 1 per chromosome arm- are repaired by a different mechanism, in which the two ends of the DSB are repaired in a way that causes them to join with the homolog (at exactly the right position), rather than rejoin with each other. This "crossing over" can be observed visually after the exchange as chiasmata (singular = chiasma) (see figure below).
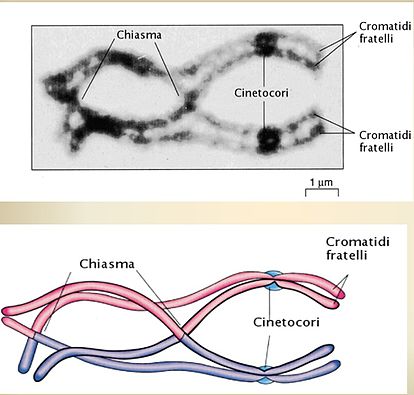
A very few DSBs are chosen to become chiasmata. I'm guessing you can read the Italian for "kinetochore" and maybe even "brother (!) chromatids". The synaptonemal complex is gone now. Not shown here, but very important to the mechanism of separation of homologs, is the fact that cohesins are holding the sister chromatids together along their entire length. Think of the two sisters as being glued together. What would happen when you pull the homologous centromeres (the centromere for pink vs. the centromere for blue) away from each other?
At the end of prophase I, the homologs are held together only at the chiasmata (figure below) and are called tetrads because the four sister chromatids of each pair of homologous chromosomes are now visible. The sisters are bound to each other throughout their length by cohesins.
The crossover events are the first source of genetic variation in the nuclei produced by meiosis. A single crossover event between homologous non-sister chromatids leads to a reciprocal exchange of equivalent DNA between a maternal chromosome and a paternal chromosome. Now, that chromatid carries some DNA from one parent of the individual and some DNA from the other parent (and its sister carries the reciprocal recombinant event) . The recombinant chromatid has a combination of maternal and paternal genes that did not exist before the crossover.
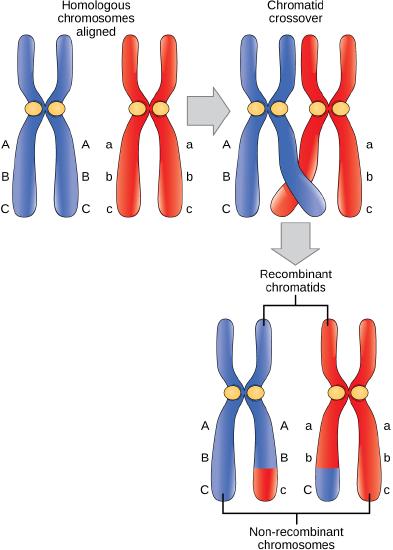
Crossover occurs between non-sister chromatids of homologous chromosomes. The result is an exchange of genetic material between homologous chromosomes. This diagram clarifies the novel genetic content found at the end of meiosis I after homologs separate. It is not a good illustration of the process of crossing over.
Possible discussion
What are the major differences between Prophase I of Meiosis and Prophase of Mitosis?
Prometaphase I
The key event in prometaphase I is the attachment of the spindle fiber microtubules to the kinetochore proteins at the centromeres. Kinetochore proteins are multiprotein complexes that bind the centromeres of a chromosome to the microtubules of the mitotic spindle. Microtubules grow from centrosomes placed at opposite poles of the cell. The microtubules move toward the middle of the cell and attach to one of the two fused homologous chromosomes. The microtubules attach at each chromosomes' kinetochores. A spindle fiber that has attached to a kinetochore is called a kinetochore microtubule. At the end of prometaphase I, each tetrad (pair of homologs, each with 2 sister chromatids) is attached to microtubules from both poles, with one homologous chromosome facing each pole. The homologous chromosomes are still held together at chiasmata, and the sisters are held together by cohesins all along their length. In addition, the nuclear membrane has broken down entirely.
Metaphase I
During metaphase I, the homologous chromosomes are arranged in the center of the cell (the metaphase plate) with the kinetochores facing opposite poles. The homologous pairs orient themselves randomly. For example, if the two homologous members of chromosome 1 are labeled a and b, then the chromosomes could line up a-b, or b-a. This is important in determining the genes carried by a gamete, as each will only receive one of the two homologous chromosomes. This is called Independent Assortment. Recall that homologous chromosomes are not identical, they contain slight differences in their genetic information, causing each gamete to have a unique genetic makeup.
This randomness is the physical basis for the creation of the second form of genetic variation in offspring. Consider that the homologous chromosomes of a sexually reproducing organism are originally inherited as two separate sets, one from each parent. Using humans as an example, one set of 23 chromosomes is present in the egg donated by the mother. The father provides the other set of 23 chromosomes in the sperm that fertilizes the egg. Every cell of the multicellular offspring has copies of the original two sets of homologous chromosomes. In prophase I of meiosis, the homologous chromosomes form the tetrads. In metaphase I, these pairs line up at the midway point between the two poles of the cell to form the metaphase plate. Because there is an equal chance that a microtubule fiber will encounter a maternally or paternally inherited chromosome, the arrangement of the tetrads at the metaphase plate is random. Any maternally inherited chromosome may face either pole. Any paternally inherited chromosome may also face either pole. The orientation of each tetrad is independent of the orientation of the other 22 tetrads. There are two possibilities for orientation at the metaphase plate; the possible number of alignments therefore equals 2n, where n is the number of chromosomes per set. Humans have 23 chromosome pairs, which results in over eight million (223) possible genetically-distinct gametes. This number does not include the variability that was previously created in the sister chromatids by crossover. Given these two mechanisms, it is highly unlikely that any two haploid cells resulting from meiosis will have the same genetic composition (see figure below). And this is without even considering the first type of variation generated by crossing over!
How does the cell ensure that the homologs are oriented in opposite directions?
You may recall from mitosis that kinetochores were oriented to opposite poles by holding the sisters together with cohesins, and then tugging on the kinetochores. If each sister's kinetochore is pointed a different way, and this is true for every chromosome, the cell will be able to detect that tension is present at every kinetochore- none are "relaxed". The absence of this "relaxed" signal will induce cleavage of the cohesins, and the chromatids can now separate and head to opposite poles.
In meiosis, the cell has not only handcuffed the sisters together via cohesins, it has also covalently bound the homologs together, at just a few spots, via the formation of chiasma. When the centromeres of each homolog are oriented in opposite directions, and the motor proteins of the kinetochores attempt to creep along the microtubules, tension will be established. Once all kinetochores are experiencing tension, the cohesins present on the chromosome arms will be cleaved, allowing the homologous centromeres to separate. The cohesins very near the centromere will remain, for use in the establishment of tension between sister kinetochores, in metaphase of meiosis II.
To summarize the genetic consequences of meiosis I, the maternal and paternal genes are recombined by crossover events that occur between each homologous pair during prophase I. In addition, the random assortment of tetrads on the metaphase plate produces a unique combination of maternal and paternal chromosomes that will make their way into the gametes.
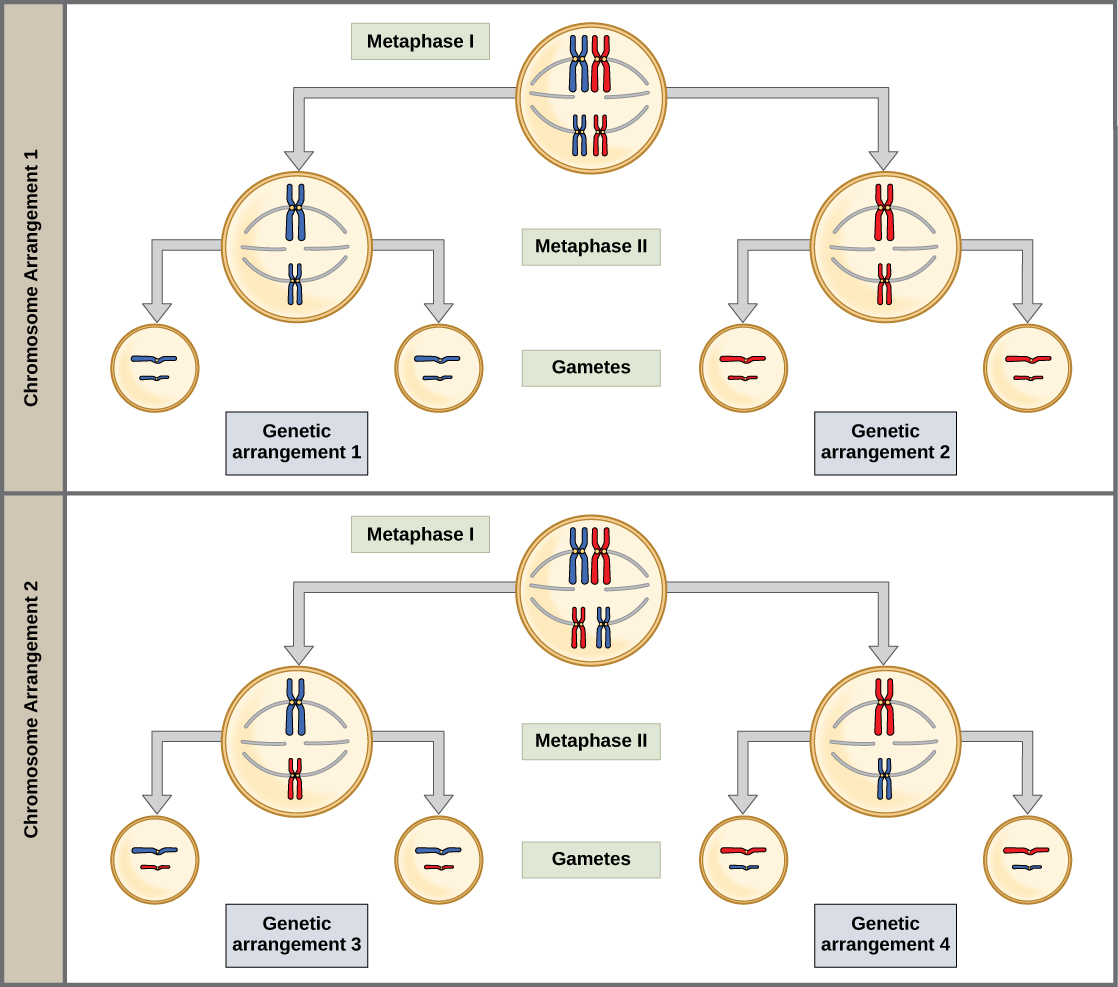
Random, independent assortment during metaphase I can be demonstrated by considering a simple cell with a set of only two chromosomes (n = 2). In this case, there are two possible arrangements at the equatorial plane in metaphase I. The total possible number of different gametes is 2n, where n equals the number of chromosomes in a set. In this example, there are four possible genetic combinations for the gametes. With n = 23 in human cells, there are over 8 million possible combinations of paternal and maternal chromosomes. Note that this figure- and this discussion- doesn't including the diversity generated by crossing over.
Anaphase I
Anaphase I is triggered by the cleavage of cohesins along the length of the chromosome arms. The kinetochore motor proteins are now free walk along the spindle and pull the homologous chromosomes apart. The spindle itself falls apart behind the progressing kinetochores. The sister centromeres remain tightly bound together via cohesins at the centromere.
Possible discussion
What major difference occurs in Anaphase I of Meiosis compared to Anaphase of Mitosis?
Telophase I and Cytokinesis
In telophase, the separated chromosomes arrive at opposite poles. The remainder of the typical telophase events may or may not occur, depending on the species. In some organisms, the chromosomes decondense and nuclear envelopes form around the chromosomes in telophase I. In other organisms, cytokinesis—the physical separation of the cytoplasmic components into two daughter cells—occurs without reformation of the nuclei (considering that another round of chromosome migration is about to occur (Mitosis II) this failure to reform a nuclear membrane makes a certain amount of sense. In nearly all species of animals and some fungi, cytokinesis separates the cell contents via a cleavage furrow (constriction of the actin ring that leads to cytoplasmic division) after Meiosis I. In plants, cytokinesis might or might not occur at this stage, depending on the species. In Arabidopsis, a popular plant model system, cytokinesis does not occur until Meiosis II is complete.
Two haploid cells are the end result of the first meiotic division. The cells are haploid because at each pole, there is just one homolog for each chromosome. Each homolog still consists of two sister chromatids. Recall that sister chromatids are almost duplicates- don't forget that they might carry information from one of the homologs somewhere along their arms due to exchanges that occurred during crossing over, and these crossover positions will be different for each sister.
Meiosis II
In meiosis II, these two sister chromatids will separate. Thus the net result of meiosis I and II will be 4 haploid cells, with chromosomes that only have 1 chromatid.
The two cells produced in meiosis I go through the events of meiosis II in synchrony. During meiosis II, the sister chromatids within the two daughter cells separate, forming a total of four new haploid gametes. The mechanics of meiosis II is similar to mitosis.
Prophase II
If the chromosomes decondensed in telophase I, they condense again. If nuclear envelopes were formed, they fragment into vesicles. The centrosomes that were duplicated between Meiosis I and II move away from each other toward opposite poles, and new spindles are formed.
Prometaphase II
The nuclear envelopes are now completely broken down, and the spindle is fully formed. Each sister chromatid's kinetochore attaches to microtubules.
Metaphase II
The sister chromatids are maximally condensed and aligned at the equator of the cell. Cohesins prevent the kinetochores from pulling apart, establishing tension and telling the cell that the kinetochores are properly aligned for every chromosome.
Anaphase II
The sister chromatids are pulled apart by the kinetochore microtubules and move toward opposite poles. Non-kinetochore microtubules elongate the cell.
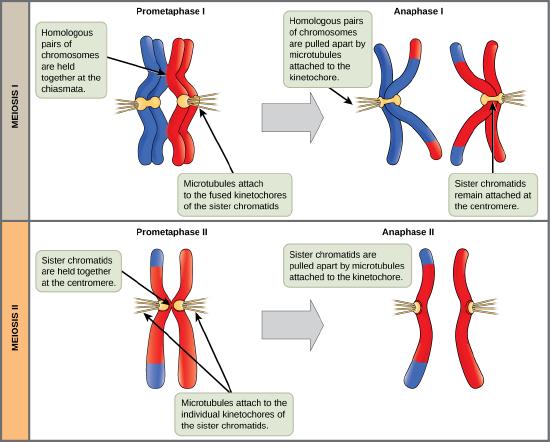
The process of chromosome alignment differs between meiosis I and meiosis II. In prometaphase I, microtubules attach to the fused kinetochores of homologous chromosomes, and the homologous chromosomes are arranged at the midpoint of the cell in metaphase I. In anaphase I, the homologous chromosomes are separated. In prometaphase II, microtubules attach to the kinetochores of sister chromatids, and the sister chromatids are arranged at the midpoint of the cells in metaphase II. In anaphase II, the sister chromatids are separated.
Telophase II and Cytokinesis
The chromosomes arrive at opposite poles and begin to decondense. Nuclear envelopes form around the chromosomes. Cytokinesis separates the two cells into four unique haploid cells. At this point, the newly formed nuclei are both haploid. The cells produced are genetically unique because of the random assortment of paternal and maternal homologs and because of the recombination of maternal and paternal segments of chromosomes (with their sets of genes) that occurs during crossing over. The entire process of meiosis is outlined in the figure below.
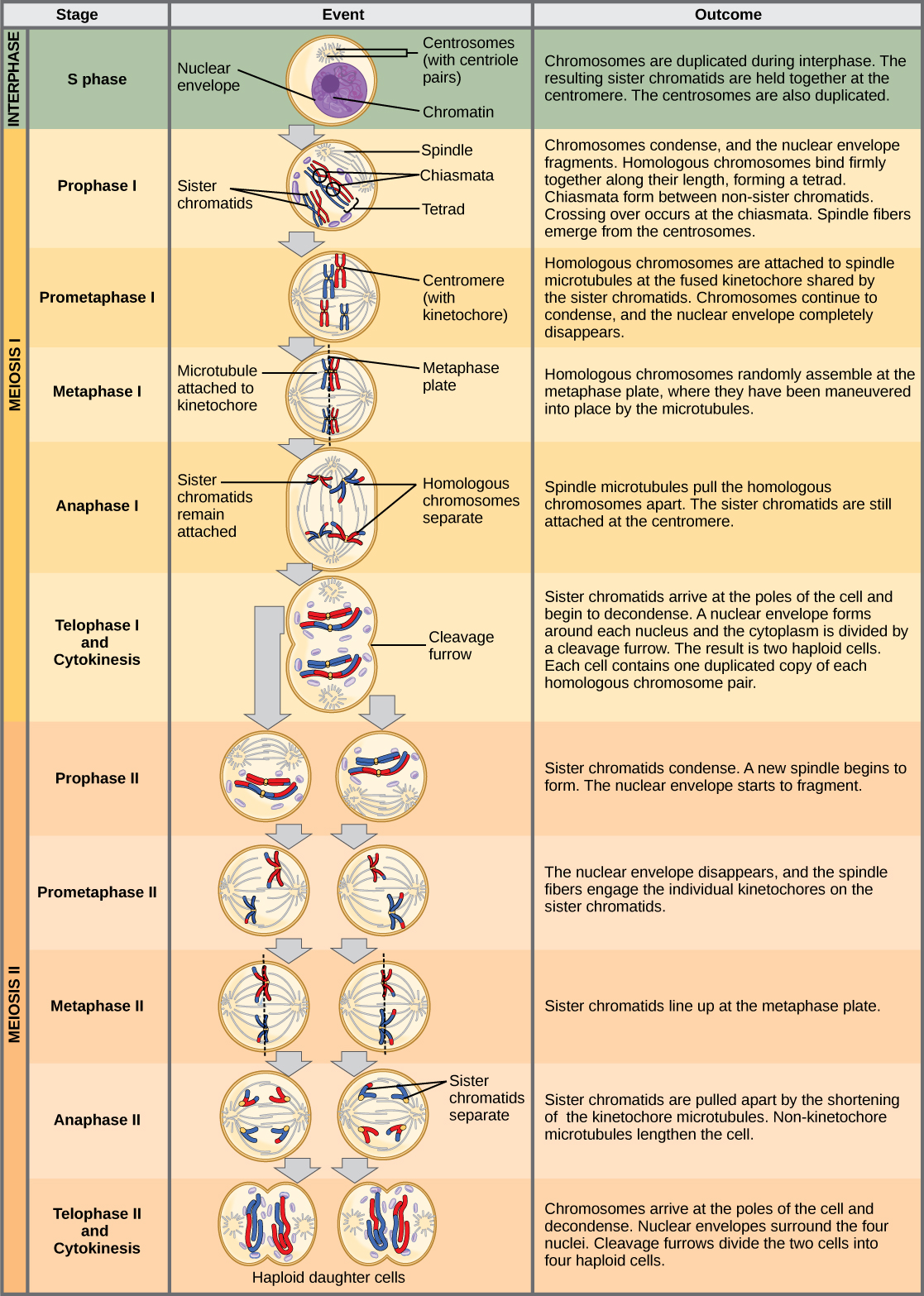
An animal cell with a diploid number of four (2n = 4) proceeds through the stages of meiosis to form four haploid daughter cells.
Comparing Mitosis and Meiosis
Mitosis and meiosis are both forms of division of the nucleus in eukaryotic cells. They share some similarities, but also exhibit distinct differences that lead to very different outcomes. Mitosis is a single nuclear division that results in two nuclei that are usually partitioned into two new cells. The nuclei resulting from a mitotic division are genetically identical to the original nucleus. They have the same number of sets of chromosomes, one set in the case of haploid cells and two sets in the case of diploid cells. In most plants and all animal species, it is typically diploid cells that undergo mitosis to form new diploid cells. In contrast, meiosis consists of two nuclear divisions resulting in four nuclei that are usually partitioned into four new cells. The nuclei resulting from meiosis are not genetically identical and they contain one chromosome set only. This is half the number of chromosome sets in the original cell, which is diploid.
The main differences between mitosis and meiosis occur in meiosis I, which is a very different nuclear division than mitosis. In meiosis I, the homologous chromosome pairs become associated with each other, are bound together with the synaptonemal complex, develop chiasmata and undergo crossover between sister chromatids, and line up along the metaphase plate in tetrads with kinetochore fibers from opposite spindle poles attached to each kinetochore of a homolog in a tetrad. All of these events occur only in meiosis I.
When the chiasmata resolve and the tetrad is broken up with the homologs moving to one pole or another, the ploidy level—the number of sets of chromosomes in each future nucleus—has been reduced from two to one. For this reason, meiosis I is referred to as a reduction division. There is no such reduction in ploidy level during mitosis.
Meiosis II is much more analogous to a mitotic division. In this case, the duplicated chromosomes (only one set of them) line up on the metaphase plate with divided kinetochores attached to kinetochore fibers from opposite poles. During anaphase II, as in mitotic anaphase, the kinetochores divide and one sister chromatid—now referred to as a chromosome—is pulled to one pole while the other sister chromatid is pulled to the other pole. If it were not for the fact that there had been crossover, the two products of each individual meiosis II division would be identical (like in mitosis). Instead, they are different because there has always been at least one crossover per chromosome. Meiosis II is not a reduction division because although there are fewer copies of the genome in the resulting cells, there is still one set of chromosomes, as there was at the end of meiosis I.
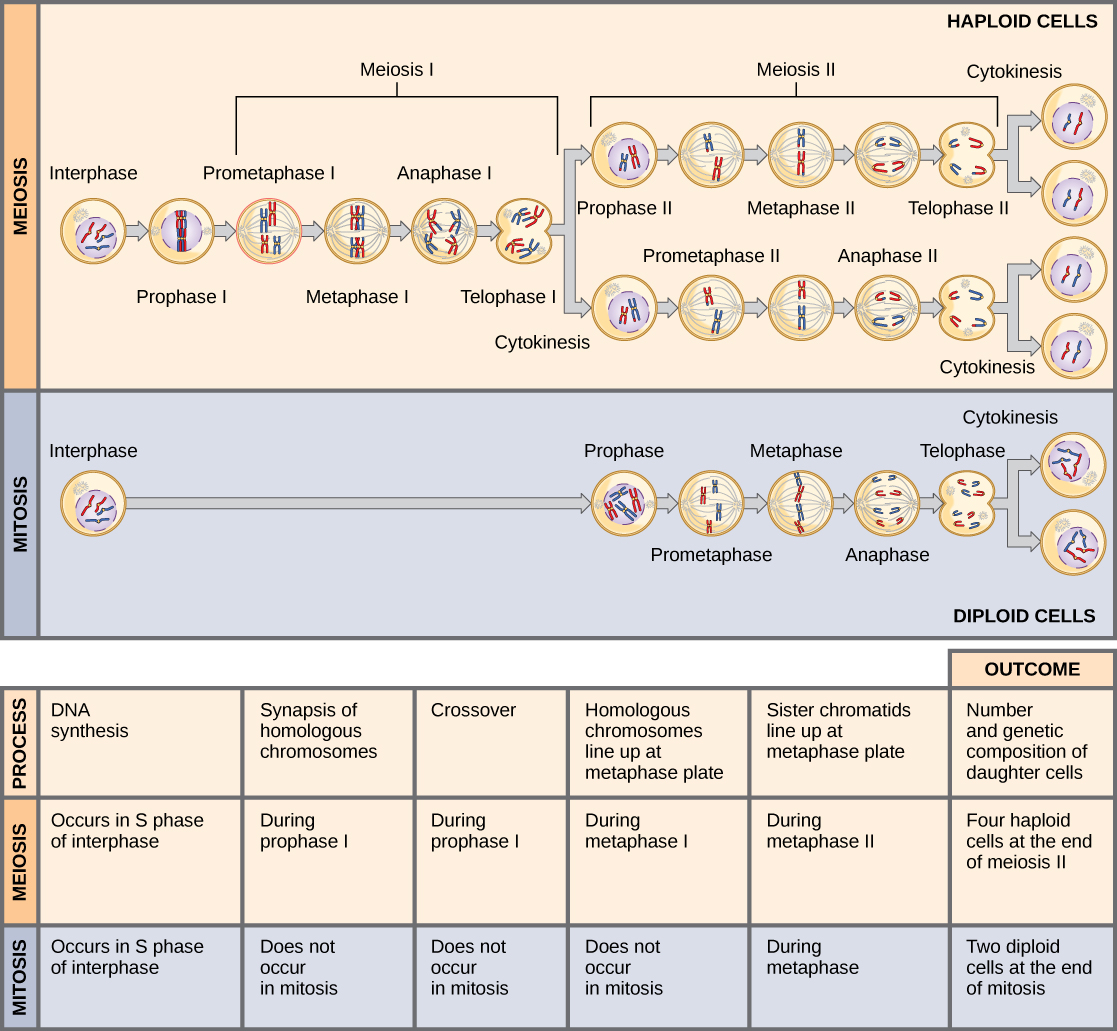
Meiosis and mitosis are both preceded by one round of DNA replication; however, meiosis includes two nuclear divisions. The four daughter cells resulting from meiosis are haploid and genetically distinct. The daughter cells resulting from mitosis are diploid and identical to the parent cell.
The Mystery of the Evolution of Meiosis
Meiosis is such an extraordinarily complex series of cellular events that biologists have had trouble hypothesizing and testing how it may have evolved. Although meiosis is inextricably entwined with sexual reproduction and its advantages and disadvantages, it is important to separate the questions of the evolution of meiosis and the evolution of sex, because early meiosis may have been advantageous for different reasons than it is now. Thinking outside the box and imagining what the early benefits from meiosis might have been is one approach to uncovering how it may have evolved.
Meiosis and mitosis share obvious cellular processes and it makes sense that meiosis evolved from mitosis. The difficulty lies in the clear differences between meiosis I and mitosis. Adam Wilkins and Robin Holliday2 summarized the unique events that needed to occur for the evolution of meiosis from mitosis. These steps are homologous chromosome pairing, crossover exchanges, sister chromatids remaining attached during anaphase, and suppression of DNA replication in interphase. They argue that the first step is the hardest and most important, and that understanding how it evolved would make the evolutionary process clearer. They suggest genetic experiments that might shed light on the evolution of synapsis.
There are other approaches to understanding the evolution of meiosis in progress. Different forms of meiosis exist in single-celled protists. Some appear to be simpler or more “primitive” forms of meiosis. Comparing the meiotic divisions of different protists may shed light on the evolution of meiosis. Marilee Ramesh and colleagues 3 compared the genes involved in meiosis in protists to understand when and where meiosis might have evolved. Although research is still ongoing, recent scholarship into meiosis in protists suggests that some aspects of meiosis may have evolved later than others. This kind of genetic comparison can tell us what aspects of meiosis are the oldest and what cellular processes they may have borrowed from in earlier cells.
Link to Learning
Click through the steps of this interactive animation to compare the meiotic process of cell division to that of mitosis: How Cells Divide.
FOOTNOTES
- Leigh Van Valen, “A new evolutionary law,” Evolutionary Theory 1 (1973): 1–30.
- Adam S. Wilkins and Robin Holliday, “The Evolution of Meiosis from Mitosis,” Genetics 181 (2009): 3–12.
- Marilee A. Ramesh, Shehre-Banoo Malik and John M. Logsdon, Jr, “A Phylogenetic Inventory of Meiotic Genes: Evidence for Sex in Giardia and an Early Eukaryotic Origin of Meiosis,” Current Biology 15 (2005):185–91.

