2.2: The Mitotic Cell Cycle
- Page ID
- 8430
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Two cells from one
The evolutionary goal of all living systems is to survive and reproduce. Since the basic unit of life is a cell, and we know that life begets new life, this means that there must be a process by which to create new cells from preexisting cells. We also know that multicellular organisms must somehow increase their number of cells during their growth by creating copies of existing cells. The process by which one cell creates another cell, for both single and multi-celled organisms, requires a parental cell to divide and is called cell division.
From the standpoint of the Design Challenge framework we can stipulate that the goal of cell division is to make a copy of a cell. If a condition for success requires that the daughter cells be viable then a number of subproblems can be defined:
1. The cell must replicate its DNA so that each daughter cell has a functional copy after cell division is complete.
2. The cell must have sufficient copies of the rest of the cellular content so that daughter cells are viable or it must find a way to ensure that the copied DNA (even without a full replica of cellular content) is viable.
3. The cell must divide the replicated cell content and DNA between at least two independent compartments.
The important thing to recognize about cell division is that the two copies of the genome (the instructions for building the cell), must be divided perfectly between the daughter cells. In contrast there are many copies of almost everything else (glucose, proteins, ribosomes, membranes). Although all of these things are essential for life, because they are present in large numbers the cell can simply trust to luck that when it divides that each daughter cell will get enough of all of these "goodies" to easily go on with growth and perhaps another round of cell division.
When we observe Nature we find that it has evolved two major modes of reproduction: sexual and asexual. Within each of these modes of reproduction we find several major modes of cell division that occur frequently across all domains of life. We consider three of these modes: binary fission (used primarily by single celled bacteria and archaea), mitosis (used often by eukaryotes in processes of cell division NOT associated with sexual reproduction) and meiosis (a process of cell division required for sexual reproduction). The major distinction between each of these modes is the mechanism for dealing with the segregation of the genome between daughter cells. We discuss these processes in the sections that follow.
Cell division in prokaryotes
Bacteria and Archaea
Typically, bacterial and archaeal cells grow, duplicate all major cellular constituents (DNA, ribosomes, enzymes, etc.), and then divide into two nearly identical daughter cells. This process is called binary fission (two cells of roughly equal size) and is shown mid-process in the figure below. While some bacterial species are known to use several alternative reproductive strategies, including making multiple offspring or budding, binary fission is the most common laboratory-observed mechanisms for cell division in prokaryotes so we limit our discussion to this mechanism alone.
(Aside: Those who want to read more about alternatives to binary fission in bacteria should check this link out.)
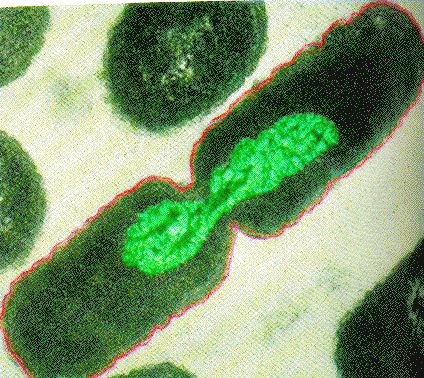
Binary Fission in bacteria starts with DNA replication at the replication origin (a special DNA sequence) near the midpoint of the cell. New replication forks can form even before the first cell division ends; this phenomenon allows an extremely rapid rate of reproduction.
Source: http://biology.kenyon.edu/courses/bi...01/week01.html
Binary Fission
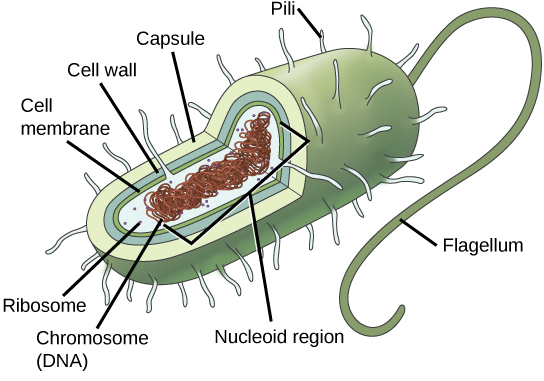
Prokaryotes, including bacteria and archaea, have a single, circular chromosome located in a central region called the nucleoid.
The process of binary fission is the most commonly observed mechanism for cell division in bacteria and archaea (at least the culturable ones studied in the laboratory). The actual division of the cell- the pinching off of the membrane to create two separate compartments, should be distinguished from the division of the daughter chromosomes.
One structural feature of relevance to DNA replication and segregation in the bacteria and archaea is that their genetic material is not enclosed in a membrane-bound nucleus, but instead exists in a semi-condensed form in a location, termed the nucleoid, within the cell. Another is that there is a single, circular chromosome- unlike eukaryotes that carry many linear chromosomes, each with a unique set of genes, which complicates chromosome segregation enormously. In contrast, because prokaryotes carry only one chromosome, the bacterium simply needs to somehow guarantee the one copy of the chromosome goes one way, and the other the opposite way. The mechanisms through which the two identical circular daughter chromosomes separate differ widely between species, and it is not a well understood process. Many models have been proposed and again, different species clearly achieve this goal in different ways. One of the most recent, and (surprisingly) simplest, models is that DNA replication itself inevitably results in the separation of the two daughters- as the "mother" genome is absorbed the two new genomes are extruded and pushed in opposite directions from a single replication site. In a cylindrical space they will spontaneously extrude towards each pole (see figure below).
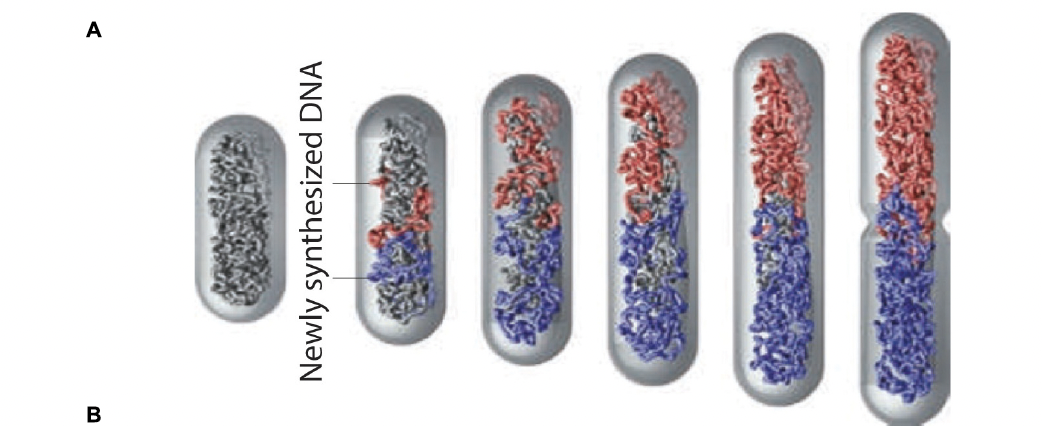
Another organizational feature to note is that the bacterial chromosome in some species is attached to the plasma membrane at about the midpoint of the cell (again, there are significant exceptions). This inspired another model for segregation. The starting point of replication, the chromosome's single origin or replication, is close to this attachment site. As the new daughter strands are formed, each origin point (now duplicated) moves away from the membrane attachment toward the opposite ends of the cell. It's is not clear how this occurs- are the origins pushed or pulled apart? What are the motors and filaments that achieve this? Are motors even required?
It is important to note that both models for the separation of the recently replicated origins solve the key problem of cell division- how to get one copy of the genome into each daughter cell. The bacterial genome has only one (circular) chromosome, which has only one origin of replication. Separating the newly duplicated origins therefore separates the genomes perfectly- each daughter cell gets a complete set of genes. In contrast, the eukaryotic genome has thousands of origins of replication, located on several chromosomes, each of which carries different parts of the genome. Can you see why this same strategy would not work in eukaryotes?
The division of the cytoplasm and all its many constituents is another issue. The formation of a ring composed of repeating units of a protein called FtsZ (a cytoskeletal protein) directs the formation of a partition between the two new nucleoids. Formation of the FtsZ ring triggers the accumulation of other proteins that work together to recruit new membrane and cell wall materials to the site. Gradually, a septum is formed between the nucleoids, extending from the periphery toward the center of the cell. When the new cell walls are in place, the daughter cells separate.

These images show the steps of binary fission in prokaryotes. (credit: modification of work by “Mcstrother”/Wikimedia Commons)
Possible discussion
How does attaching the replicating chromosome to the cell membrane aid in dividing the two chromosomes after replication is complete?
Eukaryotic Cell Cycle and Mitosis
The cell cycle is an orderly sequence of events used by biological systems to coordinate cell division. In eukaryotes, cell division proceeds via a cell cycle that includes multiple spatially and temporally coordinated events. These include a long preparatory/growth period, called interphase, and a mitotic phase called M phase. Interphase is often further divided into distinguishable subphases called G1, S, and G2 phases. Mitosis is the process by which replicated DNA is distributed to daughter cells and is itself often subdivided into five distinguishable stages: prophase, metaphase, anaphase, and telophase. Mitosis is generally followed by a process called cytokinesis, during which the cytoplasmic components of the daughter cells are separated either by an actin ring (animal cells) or by cell plate formation (plant cells). Passage through these phases is controlled by checkpoints. There are three major checkpoints in the cell cycle: one near the end of G1, a second at the G2–M transition, and the third during metaphase. These regulatory checks serve to ensure that the processes required to successfully move on to the next phase of the cell cycle have been fully completed, that the genome is undamaged, and that sufficient resources exist to move on to the next phase of cell division.
Cell Cycle
In asexually reproducing eukaryotic cells, one “turn” of the cell cycle consists of two general phases: interphase, followed by mitosis and then cytokinesis. Most of the cells in a fully-developed multicellular organisms are typically found in interphase. Mitosis is the point in the cell cycle associated with division or distribution of the chromosomes to two daughter cells. During mitosis the nuclear membrane breaks down and, after the chromosomes segregate to each pole, two new, fully functional, nuclei are formed. After mitosis is finished, Cytokinesis divides the cytoplasm into two distinct cells.
Interphase
G1 Phase
The first stage of interphase is called the G1 phase, or first "gap". During G1, the cell is quite active at the biochemical level. The cell is accumulating the building blocks of all cellular components, as well as accumulating enough energy reserves to complete the task of replicating each chromosome in the nucleus. Throughout interphase, nuclear DNA remains in a semi-condensed chromatin configuration- some regions of the genome are tightly wound and not expressed, while other parts of the chromatin more open, and actively transcribed.
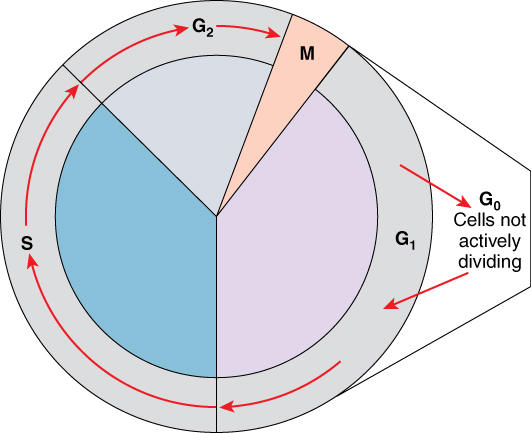
A cell moves through a series of phases in an orderly manner. During interphase, G1 involves cell growth and protein synthesis, the S phase involves DNA replication and the replication of the centrosome, and G2 involves further growth and protein synthesis. Mitosis follows interphase, and cytokinesis follows mitosis. Mitosis and cytokinesis, together, are termed "M phase".
S Phase
In S phase (synthesis phase), DNA replication results in the formation of two identical copies of each chromosome—sister chromatids—that are connected to each other along their length by proteins called cohesins. At the end of this stage, each chromosome has been replicated.
Eukaryotic cells include various microtubule organizing centers (MTOCs); in animal cells these are the centrosomes. The centrosomes are often duplicated during S phase (they will move to opposite poles of the cell during M phase). The two resulting centrosomes will give rise to the mitotic spindle, the apparatus that orchestrates the movement of chromosomes later during mitosis. MTOCs perform other functions in the organization of the microtubules, including the growth and localization of flagella. In some organisms (notably animals) centrosomes include of a pair of rod-like centrioles composed of tubulin and other proteins that sit at right angles to one another other. These structures are also often found at the sites of organization of cilia or flagella. However, centrioles are not observed in many fungi or higher plants.
G2 Phase
G2 phase, or second gap, is defined as the stage in which cells appear to have twice the amount of DNA observed in G1, while the chromosomes still appear to be diffuse (as they are throughout interphase). However, G2 cells are often dealing with issues encountered during replication (including replication forks stalled by DNA damage) and these issues must be resolved prior to entry into M phase. Some cell organelles are duplicated, and the cytoskeleton is dismantled to provide resources for the mitotic spindle. The final preparations for the mitotic phase must be completed before the cell is able to enter the first stage of mitosis. As discussed below, this a critical juncture for the cell, and the site of a cell cycle "checkpoint". Problems observed by the cell at this point will lead to an arrest of the cell cycle in G2 until these problems are resolved.
G0 Phase
Cells in the G0 phase are not actively preparing to divide. The cell is in a quiescent (mitotically inactive) stage, having exited the cell cycle. Some cells enter G0 temporarily until an external signal triggers the onset of G1. Other cells that never or rarely divide, such as mature cardiac muscle and nerve cells, remain in G0 permanently. These cells generally exit the cell cycle immediately after cytokinesis.
A Quick Aside: Structure of Chromosomes During the Cell Cycle
If the DNA from all 46 chromosomes in a human cell nucleus was laid out end to end, it would measure approximately two meters; however, its diameter would be only 2 nm. Considering that the size of a typical human cell is about 10 µm, DNA must be tightly packaged to fit in the cell’s nucleus. At the same time, it must also be readily accessible for the genes to be expressed. During some stages of the cell cycle, the long strands of DNA are condensed into compact chromosomes. There are a number of ways that chromosomes are compacted.
Suggested discussion
When should we expect to see highly condensed DNA in the cell (which phases of the cell cycle)? When would the DNA remain relatively un-compacted (during which phases of the cell cycle)?
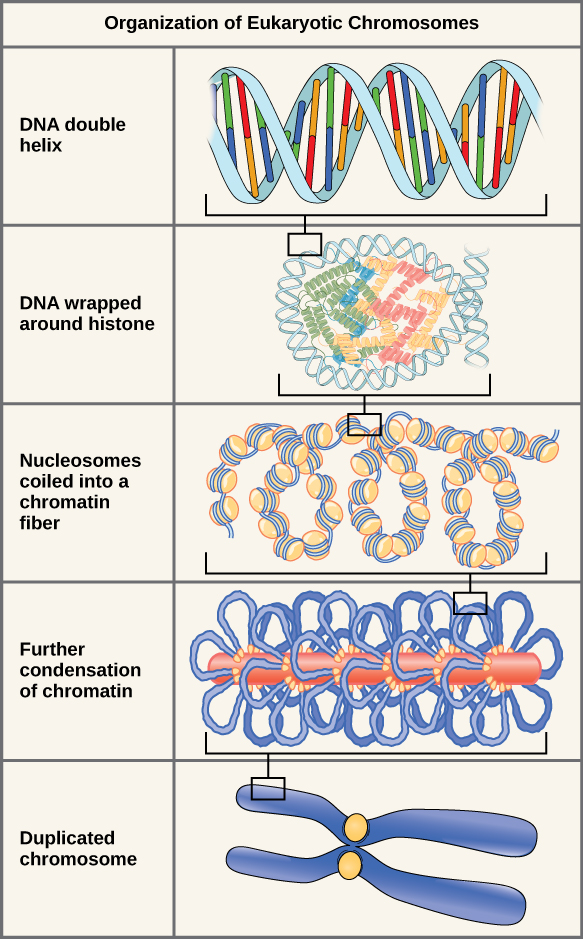
Double-stranded DNA wraps around histone proteins to form nucleosomes that have the appearance of “beads on a string.” The nucleosomes are coiled into a 30-nm chromatin fiber. When a cell undergoes mitosis, the chromosomes condense even further.
Mitosis and Cytokinesis
During cell division a cell undergoes two major processes. First, it completes mitosis, during which the duplicated information enclosed in the nucleus is distributed between two daughter nuclei. Cytokinesis then occurs, dividing the cytoplasm and cell body into two new cells.
Note:
The major phases of Mitosis are visually distinct from one another and were originally characterized by what could be seen by viewing dividing cells under a microscope. Some instructors may ask you be able to distinguish each phase be looking at images of cells or more commonly by inspection of cartoon depiction of mitosis. Dr. Britt will employ these vocabulary words, so you should learn them.
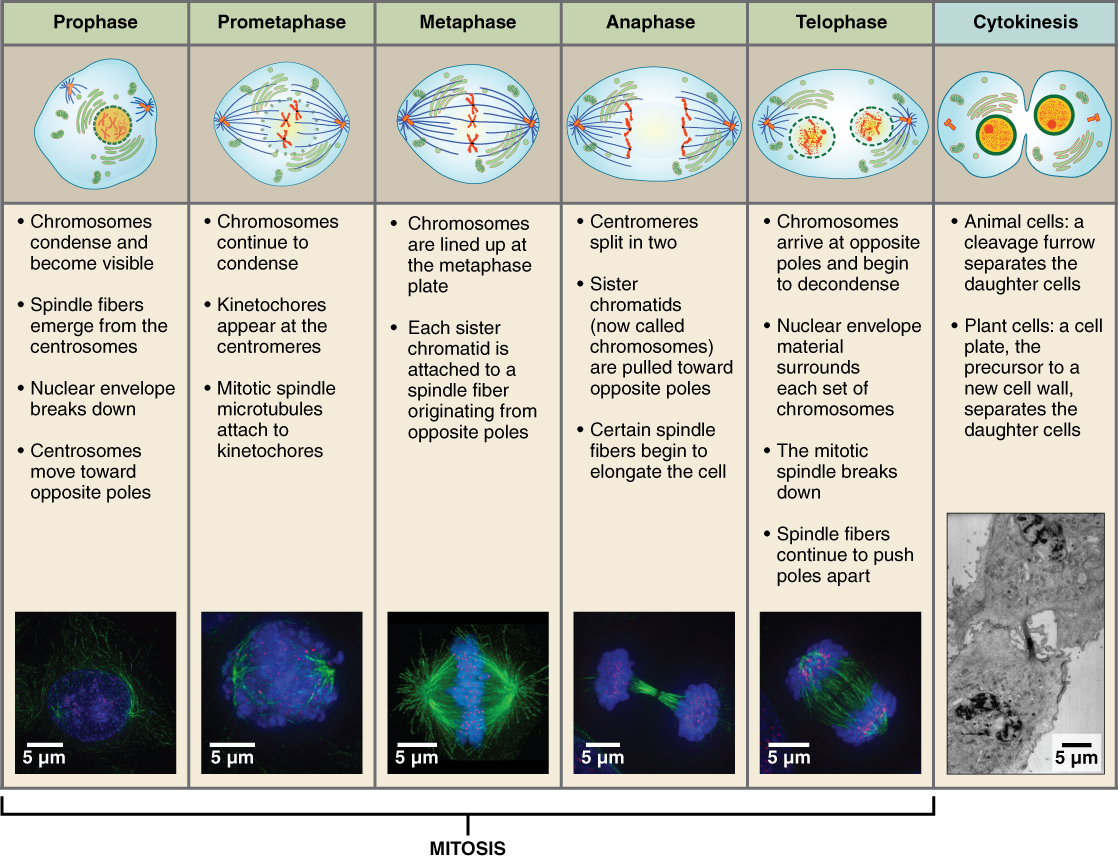
DNA is stained blue, microtubules are green. The stages of cell division oversee the separation of identical genetic material into two new nuclei, followed by the division of the cytoplasm. Mitosis is divided into five stages—prophase, prometaphase, metaphase, anaphase, and telophase—visualized here by light microscopy with fluorescence. Mitosis is usually followed by cytokinesis, shown here by a transmission electron microscope. (credit "diagrams": modification of work by Mariana Ruiz Villareal; credit "mitosis micrographs": modification of work by Roy van Heesbeen; credit "cytokinesis micrograph": modification of work by the Wadsworth Center, NY State Department of Health; donated to the Wikimedia foundation; scale-bar data from Matt Russell)
Prophase is the first phase of mitosis, during which the loosely packed chromatin coils and condenses into visible chromosomes. During prophase, each chromosome becomes visible with its identical partner (sister chromatid) attached, forming the familiar X-shape of sister chromatids. The nucleolus disappears early during this phase, and the nuclear envelope also disintegrates. The centrosome begins to migrate to each pole of the cell, and microtubules begin to emanate from this region. This marks the beginning of formation of the mitotic spindle.
Near the end of prophase there is an invasion of the nuclear area by microtubules from the mitotic spindle. The nuclear membrane has disintegrated. The kinetochore is a protein structure on the centromere that is the point of attachment between the mitotic spindle and the sister chromatids. This attachment results from the massive accumulation of motor proteins on each kinetochore- their cargo is the kinetochore itself, and they travel along the microtubules. This stage is referred to as late prophase or alternatively “prometaphase” to indicate the transition between prophase and metaphase.
What is the kinetochore?
You may recall that we have discussed that larger objects diffuse very slowly in the cell, and are assisted in their diffusion, and provided directionality, by motor proteins (dyneins and kinesins) that propel their cargo along microtubules. A chromosome is certainly the most massive structure that has to be moved by motor proteins. For most eukaryotes (again, there are always interesting exceptions), each sister chromatid carries a centromere at a single, unique position. (That position is replicated at each round of DNA replication.) Some proteins bind at the centromere throughout the cell cycle. Other additional proteins only load onto this position in prophase, and the entire assemblage (proteins new and old, at this position) is now called the kinetochore. The kinetochore includes several hundred regulatory, structural, and motor proteins. This massive structure has two major roles- to act as a tow truck for chromosomes, and to determine whetehr each sister chromatid is headed to a different pole. The kinetochore has several sequential jobs: to pull chromosomes to the metaphase plate, to establish tension by pulling each sister in opposite directions, to detect the presence or absence of tension, to trigger the release of the sister chromatids from one another once tension is established at all kinetochores simultaneously, and finally to pull the sisters to opposite poles.
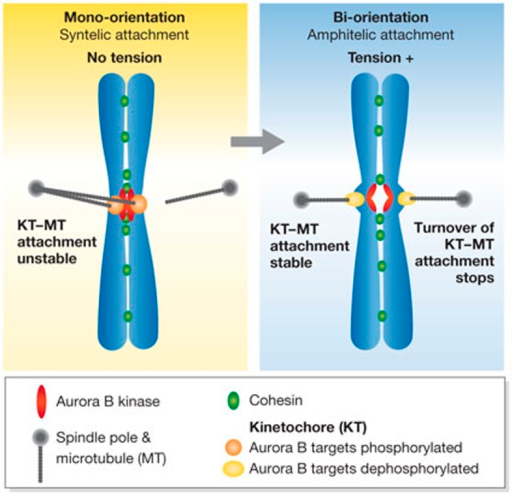
Why is tension important? From the discussion and diagram shown above, can you determine why the establishment of tension is critical for ensuring that sister chromatids move to opposite poles?
Metaphase is the second stage of mitosis. During this stage, the sister chromatids, with their attached microtubules, line up along a linear plane in the middle of the cell. The metaphase plate is the name for the plane through the center of the spindle on which the sister chromatids are positioned (it is a location, not a physical object). The microtubules are now poised to pull apart the sister chromatids and bring one from each pair to each side of the cell. Establishment of tension at every sister chromatid will result in the severing of cohesin proteins that bind the two sisters together, allowing the now properly aligned kinetochores to haul each sister to opposite poles. This migration is known as...
...Anaphase. Anaphase takes place over a few minutes, when the pairs of sister chromatids are separated from one another. These chromosomes are pulled to opposite ends of the cell by their kinetochores, as the microtubules shorten. Each end of the cell receives one partner from each pair of sister chromatids, ensuring that the two new daughter cells will contain identical genetic material. The genome has now been successfully replicated (in S phase) and evenly distributed!
Telophase is the final stage of mitosis. Telophase is characterized by the formation of two new daughter nuclei at either end of the dividing cell. These newly formed nuclear membranes surround the genetic material, which uncoils such that the chromosomes return to loosely packed chromatin. Nucleoli also reappear within the new nuclei, and the mitotic spindle breaks apart, each new cell receiving its own complement of DNA, organelles, membranes, and centrioles (if it's a species that has this feature). At this point, the cell is already beginning to split in half as cytokinesis begins.
Note
A cell accidentally produces 4 MTOCs, instead of 2. What might be the consequences for mitosis and the genetic content of the daughter cells?
Cytokinesis
Cytokinesis is the second part of the mitotic phase during which cell division is completed by the physical separation of the cytoplasmic components into two daughter cells. Although the stages of mitosis are similar for most eukaryotes, the process of cytokinesis is quite different for eukaryotes that have cell walls, such as plant cells.
In cells such as animal cells that lack cell walls, cytokinesis begins following the onset of anaphase. A contractile ring composed of actin filaments forms just inside the plasma membrane at the former metaphase plate. The actin filaments pull the equator of the cell inward, forming a fissure. This fissure, or “crack,” is called the cleavage furrow. The furrow deepens as the actin ring contracts, and eventually the membrane and cell are cleaved in two (see the figure below).
In plant cells, a cleavage furrow is not possible because of the rigid cell walls surrounding the plasma membrane. A new cell wall must form between the daughter cells. During interphase, the Golgi apparatus accumulates enzymes, structural proteins, and glucose molecules prior to breaking up into vesicles and dispersing throughout the dividing cell. During telophase, these Golgi vesicles move on microtubules to collect at the metaphase plate. There, the vesicles fuse from the center toward the cell walls; this structure is called a cell plate. As more vesicles fuse, the cell plate enlarges until it merges with the cell wall at the periphery of the cell. Enzymes use the glucose that has accumulated between the membrane layers to build a new cell wall of cellulose. The Golgi membranes become the plasma membrane on either side of the new cell wall (see panel b in the figure below).
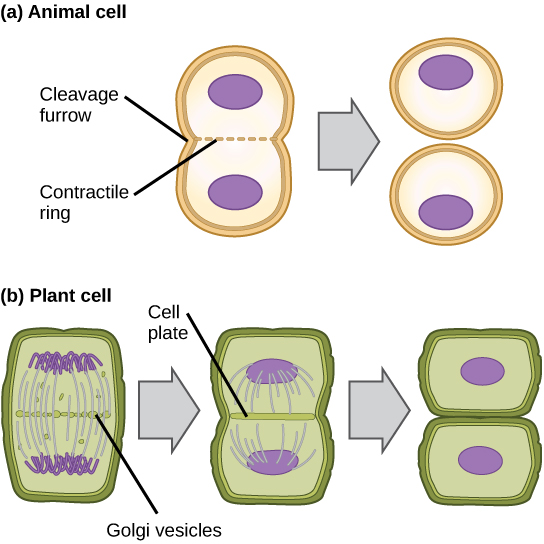
In part (a), a cleavage furrow forms at the former metaphase plate in the animal cell. The plasma membrane is drawn in by a ring of actin fibers contracting just inside the membrane. The cleavage furrow deepens until the cells are pinched in two. In part (b), vesicles coalesce at the former metaphase plate in a plant cell. The vesicles fuse and form the cell plate. The cell plate grows from the center toward the cell walls. New cell walls are made from the vesicle contents.
Cell Cycle Checkpoints
It is essential that daughter cells be nearly exact duplicates of the parent cell. Mistakes in the duplication or distribution of the chromosomes lead to mutations that may be passed forward to every new cell produced from the abnormal cell. To prevent a compromised cell from continuing to divide, there are internal control mechanisms that operate at three main cell cycle checkpoints at which the cell cycle can be stopped until conditions are favorable. These checkpoints occur near the end of G1, within S phase, at the G2–M transition, and during metaphase (see figure below).
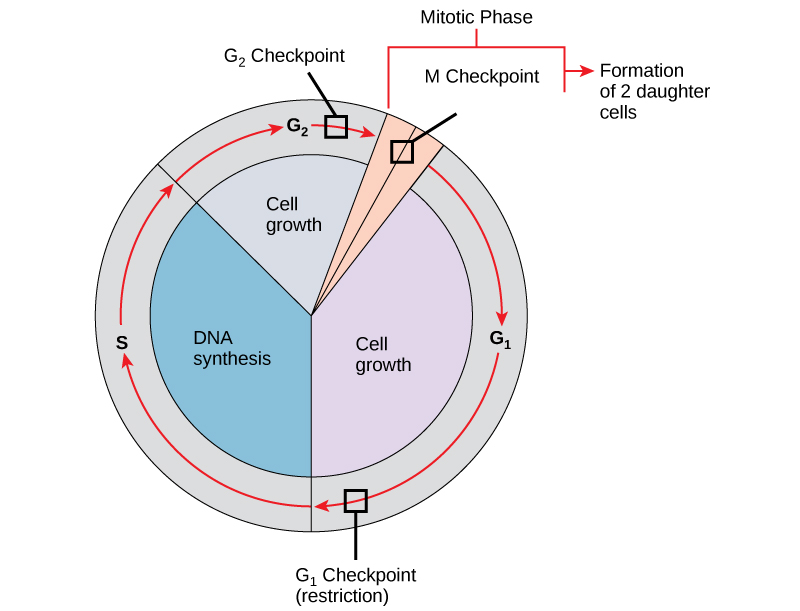
The cell cycle is controlled at three checkpoints. Integrity of the DNA is assessed at multiple checkpoints. Proper chromosome duplication is assessed at the G2 checkpoint. Attachment of each kinetochore to a spindle fiber is assessed at the M checkpoint.
G1 Checkpoint
The G1 checkpoint determines whether all conditions are favorable for cell division to proceed into S phase where DNA replication occurs. The G1 checkpoint, also called the restriction point, is the point at which the cell irreversibly commits to the cell-division process. In addition to adequate reserves and cell size, there is a check for damage to the genomic DNA- particularly breaks in the chromosome- at the G1 checkpoint. A cell that does not meet all the requirements will not be released into the S phase.
G2 Checkpoint
The G2 checkpoint bars the entry to the mitotic phase if certain conditions are not met. As in the G1 checkpoint, cell size and protein reserves are assessed. However, the most important role of the G2 checkpoint is to ensure that all of the chromosomes have been completely replicated and that the replicated DNA is not damaged.
M Checkpoint
The M checkpoint occurs near the end of the metaphase stage of mitosis. The M checkpoint is also known as the spindle checkpoint because it determines if all the sister chromatids are correctly attached to the spindle microtubules. Because the separation of the sister chromatids during anaphase is a critical step, the cycle will not proceed until the kinetochores of each pair of sister chromatids are firmly anchored to spindle fibers arising from opposite poles of the cell.
When the Cell Cycle gets out of Control
Most people understand that cancer or tumors are caused by abnormal cells that multiply continuously. If the abnormal cells continue to divide unstopped, they can damage the tissues around them, spread to other parts of the body, and eventually result in death. In healthy cells, the tight regulation mechanisms of the cell cycle prevent this from happening, while failures of cell cycle control can cause unwanted and excessive cell division. Failures of control may be caused by inherited genetic abnormalities that compromise the function of certain “stop” and “go” signals. Environmental insult that damages DNA can also cause dysfunction in those signals. Often, a combination of both genetic predisposition and environmental factors lead to cancer.
The process of a cell escaping its normal control system and becoming cancerous may actually happen throughout the body quite frequently. Fortunately, certain cells of the immune system are capable of recognizing cells that have become cancerous and destroying them. However, in certain cases the cancerous cells remain undetected and continue to proliferate. If the resulting tumor does not pose a threat to surrounding tissues, it is said to be benign and can usually be easily removed. If capable of damage, the tumor is considered malignant and the patient is diagnosed with cancer.
Homeostatic Imbalances:
Cancer Arises from Homeostatic Imbalances
Cancer is an extremely complex condition, capable of arising from a wide variety of genetic and environmental causes. Typically, mutations or aberrations in a cell’s DNA that compromise normal cell cycle control systems lead to cancerous tumors. Cell cycle control is an example of a homeostatic mechanism that maintains proper cell function and health. While progressing through the phases of the cell cycle, a large variety of intracellular molecules provide stop and go signals to regulate movement forward to the next phase. These signals are maintained in an intricate balance so that the cell only proceeds to the next phase when it is ready. This homeostatic control of the cell cycle can be thought of like a car’s cruise control. Cruise control will continually apply just the right amount of acceleration to maintain a desired speed, unless the driver hits the brakes, in which case the car will slow down. Similarly, the cell includes molecular messengers, such as cyclins, that push the cell forward in its cycle.
In addition to cyclins, a class of proteins that are encoded by genes called proto-oncogenes provide important signals that regulate the cell cycle and move it forward. Examples of proto-oncogene products include cell-surface receptors for growth factors, or cell-signaling molecules, two classes of molecules that can promote DNA replication and cell division. In contrast, a second class of genes known as tumor suppressor genes sends stop signals during a cell cycle. For example, certain protein products of tumor suppressor genes signal potential problems with the DNA and thus stop the cell from dividing, while other proteins signal the cell to die if it is damaged beyond repair. Some tumor suppressor proteins also signal a sufficient surrounding cellular density, which indicates that the cell need not presently divide. The latter function is uniquely important in preventing tumor growth: normal cells exhibit a phenomenon called “contact inhibition;” thus, extensive cellular contact with neighboring cells causes a signal that stops further cell division.
These two contrasting classes of genes, proto-oncogenes and tumor suppressor genes, are like the accelerator and brake pedal of the cell’s own “cruise control system,” respectively. Under normal conditions, these stop and go signals are maintained in a homeostatic balance. Generally speaking, there are two ways that the cell’s cruise control can lose control: a malfunctioning (overactive) accelerator, or a malfunctioning (underactive) brake. When compromised through a mutation, or otherwise altered, proto-oncogenes can be converted to oncogenes, which produce oncoproteins that push a cell forward in its cycle and stimulate cell division even when it is undesirable to do so. For example, a cell that should be programmed to self-destruct (a process called apoptosis) due to extensive DNA damage might instead be triggered to proliferate by an oncoprotein. On the other hand, a dysfunctional tumor suppressor gene may fail to provide the cell with a necessary stop signal, also resulting in unwanted cell division and proliferation.
A delicate homeostatic balance between the many proto-oncogenes and tumor suppressor genes delicately controls the cell cycle and ensures that only healthy cells replicate. Therefore, a disruption of this homeostatic balance can cause aberrant cell division and cancerous growths.

