1.4: In vitro applications of DNA replication
- Page ID
- 25725
Learning Objectives
- Compare cellular DNA replication with the human applications of PCR and sequencing methods.
- Explain the role of primers, Taq polymerase, and the temperature cycles in PCR.
- Explain the relationship between the structure of dideoxynucleotides and their function in sequencing reactions.
- Describe the process of traditional (Sanger) sequencing.
- Give an example of a NextGen sequencing method and briefly explain how it differs from traditional sequencing.
Understanding and adapting cellular DNA replication has led to innovations that have had an immeasurable impact on research, biotechnology, and medicine. Both polymerase chain reaction (PCR) and DNA sequencing are in vitro applications of cellular DNA replication.
PCR
Molecular cloning was the first method available to isolate a gene of interest and make many copies of it to obtain sufficient amounts of the DNA to study. Today, there is a faster and easier way to obtain large amounts of a DNA sequence of interest: the polymerase chain reaction (PCR). PCR allows one to use the power of DNA replication to amplify DNA enormously in a short period of time. As you know, cells replicate their DNA before they divide, and in doing so, double the amount of the cell’s DNA. PCR essentially mimics cellular DNA replication in the test tube, repeatedly copying the target DNA over and over, to produce large quantities of the desired DNA.
Selective Replication
In contrast to cellular DNA replication, which amplifies all of a cell’s DNA during a replication cycle, PCR does targeted amplification to replicate only a segment of DNA bounded by the two primers that determine where DNA polymerase begins replication. Each cycle of PCR involves three steps, denaturing, annealing and extension, each of which occurs at a different temperature.

The Starting Materials
Since PCR is, basically, replication of DNA in a test-tube, many of the usual ingredients needed for DNA replication are required:
- A template (the DNA containing the target sequence that is being copied)
- Primers (to initiate the synthesis of the new DNA strands)
- Thermostable DNA polymerase (to carry out the synthesis). The polymerase needs to be heat stable, because heat is used to separate the template DNA strands in each cycle.
- dNTPs (DNA nucleotides to build the new DNA strands).
- The template is the DNA that contains the target you want to amplify (the "target" is the specific region of the DNA you want to amplify).
The primers are short synthetic single-stranded DNA molecules whose sequence is complementary to a region flanking the target sequence. These DNA molecules can be chemically-synthesized to produce any given base sequence to use as primers. Primers are present in millions of fold excess over the template, because each new DNA strand will begin from a primer. Therefore, it is necessary to know a little bit of the template sequence on either side of the region of DNA to be amplified. DNA polymerases and dNTPs are commercially available from biotechnology supply companies.
- The first step of the process involves separating the strands of the target DNA by heating to near boiling.
- Next, the solution is cooled to a temperature that favors complementary DNA sequences finding each other and making base pairs, a process called annealing. Since the primers are present in great excess, the complementary sequences they target are readily found and base-paired to the primers. These primers direct the synthesis of DNA. Only where a primer anneals to a DNA strand will replication occur, since DNA polymerases require a primer to begin synthesis of a new strand.
- In the third step in the process, the DNA polymerase replicates DNA by extension from the 3’ end of the primer, making a new DNA strand. At the end of the first cycle, there are twice as many DNA molecules, just as in cellular replication. But in PCR, steps 1-3 are repeated, usually for between 25 and 30 cycles. At the end of the process, there is a theoretical yield of 230 (over 1 billion times) more DNA than there was to start. (This enormous amplification power is the reason that PCR is so useful for forensic investigations, where very tiny amounts of DNA may be available at a crime scene.)
The temperature cycles are controlled in a thermocycler, which repeatedly raises and lowers temperatures according to the set program. Since each cycle can be completed in a couple of minutes, the entire amplification can be completed very rapidly. The resulting DNA is analyzed on a gel to ensure that it is of the expected size, and depending on what it is to be used for, may also be sequenced, to be certain that it is the desired fragment.
Example \(\PageIndex{1}\)
PCR is used for one type of test for SARS-CoV-2, the virus associated with COVID-19. Why would this tool be helpful to detect the virus? What materials would be needed to perform this test?
Solution
Because PCR exponentially amplifies the number of copies of genetic material, even when only a small amount of genetic material is present, many copies can be produced and detected. In this case, a small amount of virus would be extremely hard to detect without PCR. However, this virus has an RNA (not DNA) genome, so an extra step must convert RNA to DNA, then copies can be made!
You would need: a sample of cells from an individual to isolate genetic material from, Taq polymerase and appropriate buffer, dNTPs, primers specific to the virus genetic sequence
Query \(\PageIndex{1}\)
DNA sequencing
DNA sequencing determines the order of bases in a piece of DNA, a whole chromosome, or even a whole genome. Remember that DNA obeys the rules of base pairing, so determining the sequence of one strand means that we know the other strand as well. DNA sequencing is an application of DNA replication, because DNA strands are built using nucleotides or methods that allow the scientist to visualize which base was incorporated at each position.
Dideoxy sequencing (also known as Sanger sequencing)
The original method developed by Sanger and others relied on variant nucleotides (dNTPs), dideoxynucleotides (ddNTPs), which are also known as chain terminators. Remember, that DNA synthesis always proceeds from 5' to 3' by attaching the phosphate of the incoming nucleotide to the 3' OH of the previous nucleotide. When that hydroxyl group is absent as in a dideoxynucleotide, synthesis ends.
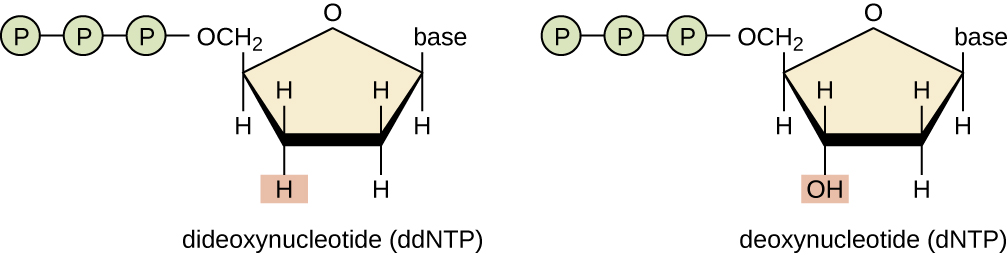
Figure \(\PageIndex{4}\): A dideoxynucleotide has a hydrogen on the 3' carbon, whereas a deoxynucletide has a hydroxyl group. (https://cnx.org/contents/5CvTdmJL@4.4, CC BY 4.0, https://commons.wikimedia.org/w/inde...curid=53713264)
When a combination of dNTPs and ddNTPs are included, a mixture of DNA sequences of different lengths is produced. Each fragment ends with a labeled didexoynucledotide. Note that this process first utilized radioactively-labeled nucleotides; only one type of radioactively-labeled dideoxynucleotide (ddATP, ddGTP, ddTTP, or ddCTP) could be included in each reaction, meaning four individual reactions had to be performed. The use of fluorescent labels for the dideoxynucleotides simplified this process, allowing all four bases to be sequenced in a single reaction. Ordering the mixture of fragments by size using electrophoresis and reading the wavelength of each label using a laser will reveal the order of bases.
The reagents needed for a sequencing reaction include:
- Template DNA (the DNA you want to know the sequence of)
- A mixture of all four deoxynucleotides (dATP, dCTP, dGTP, AND dTTP)
- Fluorescently-labeled dideoxynucleotide (ddATP, ddCTP, ddGTP, OR ddTTP)
- DNA polymerase (and buffer for efficient enzyme function)
- DNA oligonucleotide primer (complementary to the sequence 5' of the unknown sequence)
Video \(\PageIndex{2}\): See how the mixture of fragments produced by this reaction can be used to determine a sequence of DNA in this video. Note that whether a dNTP or ddNTP is incorporated at each position occurs at random. By chance, all possible fragments lengths will be generated in a successful sequencing reaction. (Copyright SWLeacock own work)
Visualizing the sequence
After the sequencing reaction is completed, the randomly-terminated fragments are separated by size using electrophoresis. If the reaction proceeded optimally, the products are a mixture of fragments with each base represented at the end of a fragment. Each fragment ends with a fluorescently-labeled dideoxyribonucleotide. The smallest fragments will migrate the fastest and pass by a laser that will detect the intensity of the fluorescence and record the corresponding base. The results can be visualized in a plot that represents the peaks of fluorescence intensity at each size.
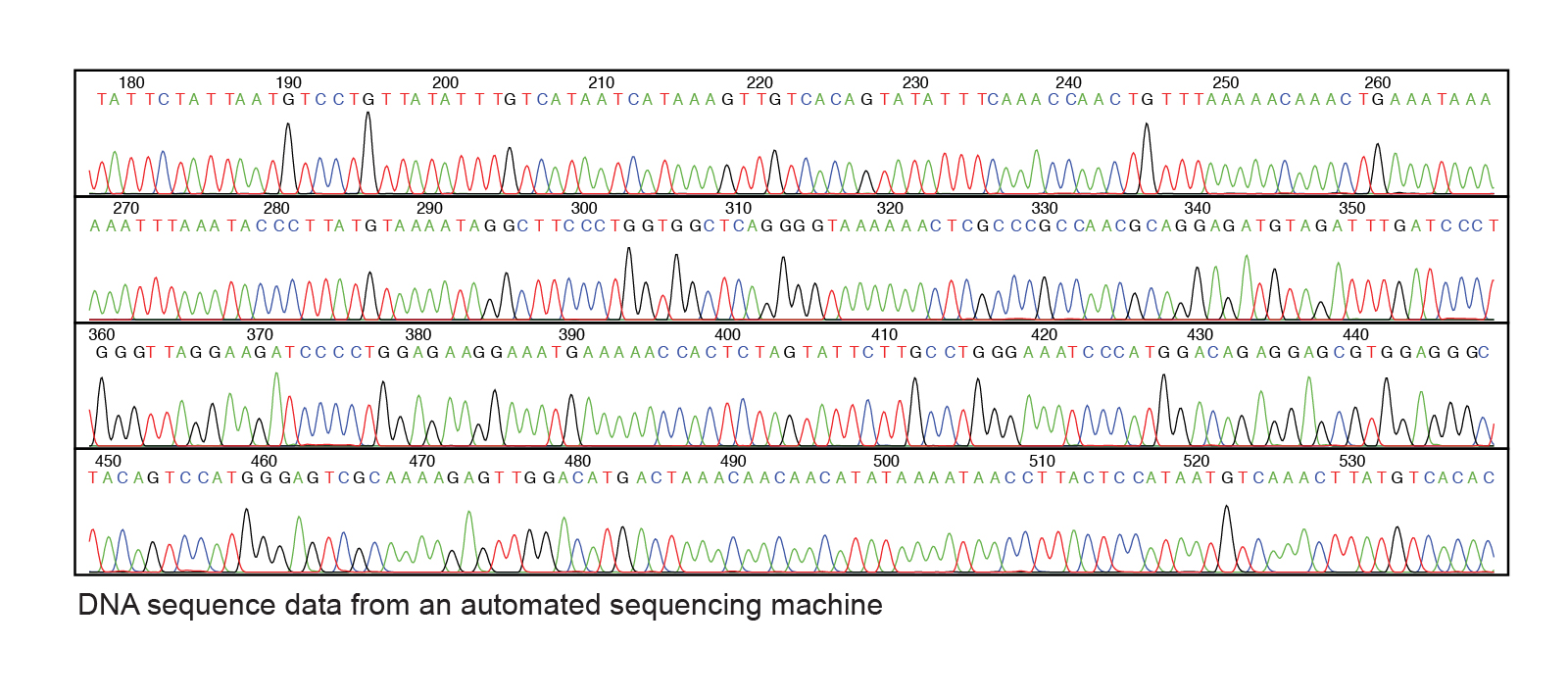
Exercise \(\PageIndex{1}\)
1. Starting at the A at nucleotide 450 in the figure above, give 20 nucleotides of sequence that recorded? (Show the 5' and 3' ends.)
2. Based on your answer above, what was the sequence of the template DNA strand? (Show the 5' and 3' ends.)
- Answer
-
1. 5' ACAGTCCATGGGAGTCGCAA 3'
2. 3' TGTCAGGTACCCTCAGCGTT 5'
Next-generation sequencing
Traditional Sanger sequencing is still used for some sequencing applications, particularly for small pieces of DNA. However, newer technologies have improved the speed and lowered the cost of DNA sequencing.
Three of these next-generation technologies (named for the companies that developed them) are highlighted here, but others exist and new ones are likely.
Illumina Sequencing by Synthesis
Pacific Biosciences
Nanopore
Compare and contrast
- Compare and contrast cellular DNA replication with DNA sequencing, addressing how DNA sequencing depends on the rules of base pairing, DNA structure, and DNA synthesis.
- Compare and contrast traditional sequencing and next-generation methods.
Exercise \(\PageIndex{1}\)
Which platform of next generation sequencing does not involve sequencing the DNA as it is being synthesized by a DNA polymerase?
A. Sanger sequencing
B. Illumina sequencing
C. Pacific Biosciences sequencing
D. Nanopore sequencing
- Answer
-
D. In Oxford Nanopore sequencing a double-stranded DNA is separated and one strand is processed through the "pore" and changes in current are measured as nucleotides pass through the channel. DNA polymerase is not synthesizing the strand while it is being sequenced.


