1.3.1: DNA replication
- Page ID
- 31680
Learning Objectives
- Summarize the process of DNA replication, including bidirectional synthesis and explain the Meselson-Stahl experiment that demonstrated semiconservative replication.
- Recall that sister chromatids are identical copies of each other produced during S phase.
- Understand how DNA polymerase proofreads using exonuclease activity.
- Explain how telomerase replicates ends of chromosomes and the cellular significance of chromosome shortening, addressing the specific function of telomerase RNA.
- Compare prokaryotic and eukaryotic DNA replication.
DNA Replication
When the cell enters S (synthesis) phase in the cell cycle, all the chromosomal DNA must be replicated. DNA polymerases synthesize new strands by adding nucleotides to the 3'-OH group present on the previous nucleotide using the separated single strands of DNA as templates. This process generates two new double-stranded molecules, called sister chromatids, from one double helix. But how are the new and old strands distributed? The answer to this question was elucidated by classic experiments by Meselson and Stahl.
Videos about the experiment that demonstrated semiconservative DNA replication
The Mechanisms of Replication
Like many molecular events we will study, replication can be divided into three stages: initiation, elongation, and termination.
Initiation
DNA polymerase does not begin at random locations on chromosomes. DNA replication in both prokaryotes and eukaryotes begins at an origin of replication (Ori), which are specific sequences at specific positions on chromosomes. In E. coli, the OriC origin is ~245 bp in size. Chromosome replication begins with the binding of an initiator protein (DnaA) to an AT-rich sequence (for example TTATCCACA in E. coli) in the OriC and melts (disrupts the hydrogen bonding between) the two strands. Helicase enzymes will be added to unwind additional DNA from this point.
In prokaryotes, with a small, simple, circular chromosome, only one origin of replication is needed to replicate the whole genome. For example, E. coli has a ~4.5 Mb genome (chromosome) that can be duplicated in ~40 minutes assuming a single origin, bi-directional replication, and a speed of ~1000 bases/second/fork for the polymerase. At least five prokaryotic DNA polymerases have been discovered to date with DNA Polymerase III (Pol III) being the primary polymerase for replication. Pol I is used primarily to fill in gaps created during lagging strand synthesis (described below) or through error-correcting mechanisms. DNA polymerase II, IV and V are used to synthesize DNA when certain types of repair is needed at other times in the cellular life cycle.
Thinking about sequences
Why would an initiator protein binding site in the OriC be an AT-rich region?
Hint: Think about the number of hydrogen bonds between AT and GC base pairs.
In eukaryotes, with multiple linear chromosomes, more than one origin of replication is required per chromosome to duplicate the whole chromosome set in the 8-hours of S-phase of the cell cycle. For example, the human diploid genome has 46 chromosomes (6 x 109 base pairs). Even the shortest chromosomes are ~50 Mbp long and so could not possibly be replicated from one origin. Additionally, the rate of replication fork movement is slower than in prokaryotes, only ~100 base/second. Thus, eukaryotes contain multiple origins of replication distributed over the length of each chromosome to enable the duplication of each chromosome within S-phase.
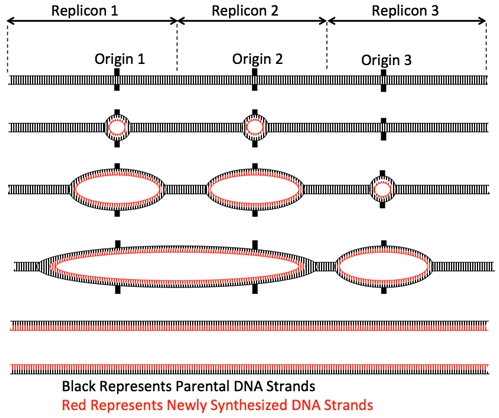
Elongation
As DNA polymerase proceeds along the template, the nucleotide that base pairs with each base on the template is covalently bonded to the 3' end of the growing strand. Note that the energy is provided by the nucleotide triphosphate itself; two phosphates are released and one phosphate remains as a part of the phosphodiester bond.
Termination
In prokaryotes, elongation proceeds bidirectionally until the replication forks meet. RNA primers are removed by a specialized DNA polymerase and then DNA is synthesized in their place. The resulting DNA fragments are then "sealed" together with DNA ligase, an enzyme that forms covalent phosphodiester bonds between two nucleotides.
In eukaryotes, replication also proceeds bidirectionally until adjacent forks meet, or the fork encounters the end of the chromosome.
Query \(\PageIndex{1}\)
Implications and limitations of the 5'-to-3' activity of DNA polymerase
Recall that enzymes are specific to their substrates. In the case of DNA polymerase, the structure only allows it to add nucleotides to the hydroxyl (-OH) group on the 3' carbon of an existing DNA strand, which presents some questions:
1. If the enzyme can only add nucleotides to an existing DNA strand (meaning that it cannot take two free nucleotides and attach them), how will it get started?
2. Because DNA is double stranded, each strand needs to be used as a template, but these strands are antiparallel and DNA polymerase replicates both strands via one complex of proteins. How can one complex make new DNA in opposite directions?
3. How will the 3' end be replicated when there is no longer a place for a primer on the complementary strand?
Obstacle #1: How to begin
Once oriC has been opened and the helicases have attached to the two sides of the replication fork, the replication machine, aka the replisome can begin to form. However, before the DNA polymerases take positions, they need to be primed. DNA polymerases are unable to join two individual free nucleotides together to begin forming a nucleic acid; they can only add onto a pre-existing strand of at least two nucleotides. Therefore, a specialized RNA polymerase (RNAP’s do not have this limitation) known as primase is a part of the replisome, and reads creates a short RNA strand termed the primer for the DNA polymerase to add onto. Although only a few nucleotides are needed, the prokaryotic primers may be as long as 60 nt depending on the species. The helicase will continue to travel in front of the fork to unwind new DNA and allow primase to add new primers as needed.
Obstacle #2: Make two strands in opposite directions at the same time
Because DNA is being unwound in the direction of fork movement, both strands need to be synthesized in the unwound region at the same time.The two subunits of DNA polymerase that are adding nucleotides are actually tethered together, so they cannot travel in opposite directions.
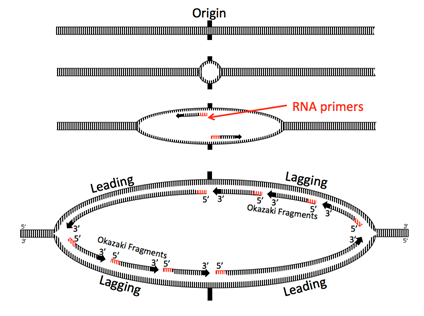
Helicase opens up the double stranded DNA and leads the rest of the replication machine along. Because nucleic acids are polymerized by adding the 5’ phosphate of a new nucleotide to the 3’ hydroxyl of the previous nucleotide, one of the strands, called the leading strand, is being synthesized in the same direction that the replication machine moves. No problem there.
The other strand is problematic: looked at linearly, the newly synthesized strand would be going 3’ to 5’ in the direction of helicase movement, but DNA polymerases cannot add nucleotides to the 5’ end (remember that it has the phosphate, not a a hydroxyl). How do cells resolve this problem? In the current model, the replication machine consists of the helicase, primases, and two DNA polymerase III holoenzymes moving in the same physical direction (following the helicase). In fact, the Pol III complexes are physically linked.

For the other strand to be replicated, the strand must be fed into the polymerase backwards, by looping the DNA around. Moving behind DNA helicase, primase quickly synthesizes a short primer before having to move forward with the replisome and synthesizing again, leaving intermittent primers in its wake. Thus, Pol III is forced to synthesize only short fragments of the chromosome at a time, called Okazaki fragments after their discoverers, Reiji and Tsuneko Okazaki. Pol III begins synthesizing by adding nucleotides onto the 3’ end of a primer and continues until it hits the 5’ end of the next primer. Pol III does not (and can not) connect the strand it is synthesizing with the 5’ end of the RNA primer.
DNA replication is called a semi-discontinuous process because while the leading strand is being synthesized continuously, the lagging strand is synthesized in fragments. This leads to two major problems: first, there are little bits of RNA left behind in the newly made strands (just at the 5’ end for the leading strand, in many places for the lagging); and second, Pol III can only add free nucleotides to a fragment of single stranded DNA; it cannot connect another fragment. Therefore, the new “strand” is not whole, but riddled with missing phosphodiester bonds.
The first problem is resolved by DNA polymerase I. Unlike Pol III, Pol I is a monomeric protein and acts alone, without additional proteins. There are also 10-20 times as many Pol I molecules as there are Pol III molecules, because they are needed for so many Okazaki fragments. DNA Polymerase I has three activities.
- Like Pol III, it can synthesize a DNA strand based on a DNA template,
- Also like Pol III, it is a 3’-5’ proofreading exonuclease.
- Unlike Pol III, it is also a 5’-3’ exonuclease. The 5’-3’ exonuclease activity is crucial in removing the RNA primer. The 5’-3’ exonuclease binds to double- stranded DNA that has a single-stranded break in the phosphodiester backbone such as what happens after Okazaki fragments have been synthesized from one primer to the next, but cannot be connected. This 5’-3’ exonuclease then removes the RNA primer. The polymerase activity then adds new DNA nucleotides to the upstream Okazaki fragment, filling in the gap created by the removal of the RNA primer. The proofreading exonuclease acts just like it does for Pol III, immediately removing a newly incorporated incorrect nucleotide. After proofreading, the overall error rate of nucleotide incorporation is approximately 1 in 107.
Even though the RNA has been replaced with DNA, this still leaves a fragmented strand. The last major player in the DNA replication story finally appears: DNA ligase. This enzyme has one simple but crucial task: it catalyzes the attack of the 3’-OH from one fragment on the 5’ phosphate of the next fragment, generating a phosphodiester bond.
Obstacle #3: How to copy the end?
The ends of linear chromosomes present a problem – at each end one strand cannot be completely replicated because there is no primer to extend. Although the loss of such a small sequence would not be a problem, the continued rounds of replication would result in the continued loss of sequence from the chromosome end to a point where it would begin to lose essential gene sequences. Thus, this DNA must be replicated. Most eukaryotes solve this problem with a specialized DNA polymerase called telomerase, in combination with a regular polymerase. Telomerases are RNA-directed DNA polymerases that are riboproteins, as they are composed of both protein and RNA. These enzymes contain a small piece of RNA that serves as a portable and reusable template from which the complementary DNA is synthesized. The RNA in human telomerases uses the sequence 3'-AAUCCC-5' as the template, and thus human telomeric DNA has the complementary sequence 5'-TTAGGG-3' repeated thousands of times. After telomerase has made the first strand, a primase synthesizes an RNA primer and a DNA polymerase then makes a complement of the extended sequence.
Telomerase = non-coding RNA + reverse transcriptase protein
In humans, the gene for the telomerase RNA is named TERC (telomerase RNA component) and is found on chromosome 3. This gene therefore encodes a product (an RNA) but not a protein!
The gene for the protein is named TERT (telomerase reverse transcriptase) and is found on chromosome 5.
These independently-encoded and independently-produced gene products must assemble to perform their function.
Note that the number of repeats, and thus the size of the telomere, is not set, but may fluctuate after each round of the cell cycle. Because there are many repeats at the end, this fluctuation maintains a length buffer – sometimes it’s longer, sometimes it’s shorter – but the average length will be maintained over the generations of cell replication.
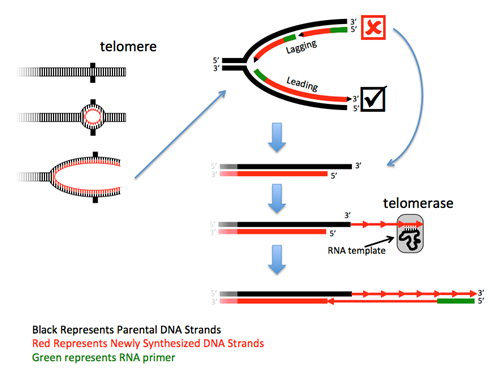
Query \(\PageIndex{1}\)
In the absence of telomerase, as is the case in human somatic cells, repeated cell division leads to telomere shortening. If a critical limit is reached, the cells enter a senescence phase of non-growth. The activation of telomerase expression, which often occurs in cancer cells, permits a cell and its descendants to become immortal because their telomeres will not reach the critical limit. Cells that can be grown in culture, such as HeLa cells that overexpress telomerase can be propagated essentially indefinitely. HeLa cells, which were originally isolated from an African-American cancer patient, Henrietta Lacks, without her consent, have been kept in culture since 1951.
DNA polymerase proofreading
DNA polymerase incorporates the correct nucleotides into each new strand based on the favorable hydrogen bonding that occurs between bases that that bind using the rules of base pairing (A-T and G-C). If an incorrect base is incorporated, the structure of the double helix will be distorted and will often be detected by the enzyme if it has not proceeded too far down the strand. DNA polymerase has 3' to 5' exonuclease activity to remove nucleotides through the mismatch and then repeat synthesis of that region of the strand with the correct nucleotides. As a result of this proofreading ability, DNA polymerases have very low error rates, allowing organisms to successfully pass genetic information from cell to cell and from generation to generation.
If a nucleotide that was missing the OH group at the 3’ end was added to the growing strand by DNA polymerase, what do you think would happen to replication?
(a) Replication would continue as normal
(b) Replication would stop as another nucleotide cannot be added to this nucleotide
- Answer
-
(b) Replication would stop as another nucleotide cannot be added to this nucleotide. Deoxyribose nucleotides that are missing the 3' OH are called dideoxyribose nucleotides and are utilized in Sanger DNA sequencing. The lack of a free OH prevents DNA polymerase from adding on another nucleotide, causing DNA synthesis to terminate.
Contributors and Attributions
Todd Nickle and Isabelle Barrette-Ng Online Open Genetics
E.V. Wong, Cells - Molecules and Mechanisms (https://bio.libretexts.org/Bookshelves/Cell_and_Molecular_Biology/Book%3A_Cells_-_Molecules_and_Mechanisms_(Wong)/7%3A_DNA/7.3%3A_Prokaryotic_Replication)


