20.2: Cloning
- Page ID
- 70760
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)In general, cloning means the creation of a perfect replica. Typically, the word is used to describe the creation of a genetically identical copy. In biology, the re-creation of a whole organism is referred to as “reproductive cloning.” Long before attempts were made to clone an entire organism, researchers learned how to copy short stretches of DNA—a process that is referred to as molecular cloning.
Molecular Cloning
Cloning allows for the creation of multiple copies of genes, expression of genes, and study of specific genes. To get the DNA fragment into a bacterial cell in a form that will be copied or expressed, the fragment is first inserted into a plasmid. A plasmid (also called a vector in this context) is a small circular DNA molecule that replicates independently of the chromosomal DNA in bacteria. In cloning, the plasmid molecules can be used to provide a “vehicle” in which to insert a desired DNA fragment. Modified plasmids are usually reintroduced into a bacterial host for replication. As the bacteria divide, they copy their own DNA (including the plasmids). The inserted DNA fragment is copied along with the rest of the bacterial DNA. In a bacterial cell, the fragment of DNA from the human genome (or another organism that is being studied) is referred to as foreign DNA to differentiate it from the DNA of the bacterium (the host DNA).
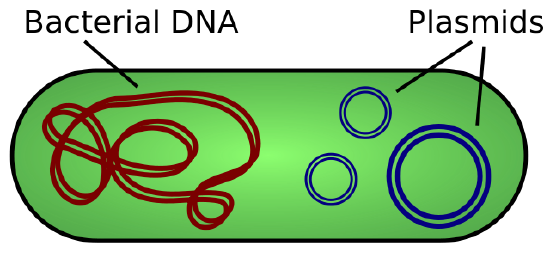
Plasmids occur naturally in bacterial populations (such as Escherichia coli) and have genes that can contribute favorable traits to the organism, such as antibiotic resistance (the ability to be unaffected by antibiotics). Plasmids have been highly engineered as vectors for molecular cloning and for the subsequent large-scale production of important molecules, such as insulin. A valuable characteristic of plasmid vectors is the ease with which a foreign DNA fragment can be introduced. These plasmid vectors contain many short DNA sequences that can be cut with different commonly available restriction enzymes. Restriction enzymes (also called restriction endonucleases) recognize specific DNA sequences and cut them in a predictable manner; they are naturally produced by bacteria as a defense mechanism against foreign DNA. Many restriction enzymes make staggered cuts in the two strands of DNA, such that the cut ends have a 2- to 4-nucleotide single-stranded overhang. The sequence that is recognized by the restriction enzyme is a four- to eight-nucleotide sequence that is a palindrome. Like with a word palindrome, this means the sequence reads the same forward and backward. In most cases, the sequence reads the same forward on one strand and backward on the complementary strand. When a staggered cut is made in a sequence like this, the overhangs are complementary (Figure \(\PageIndex{2}\)).
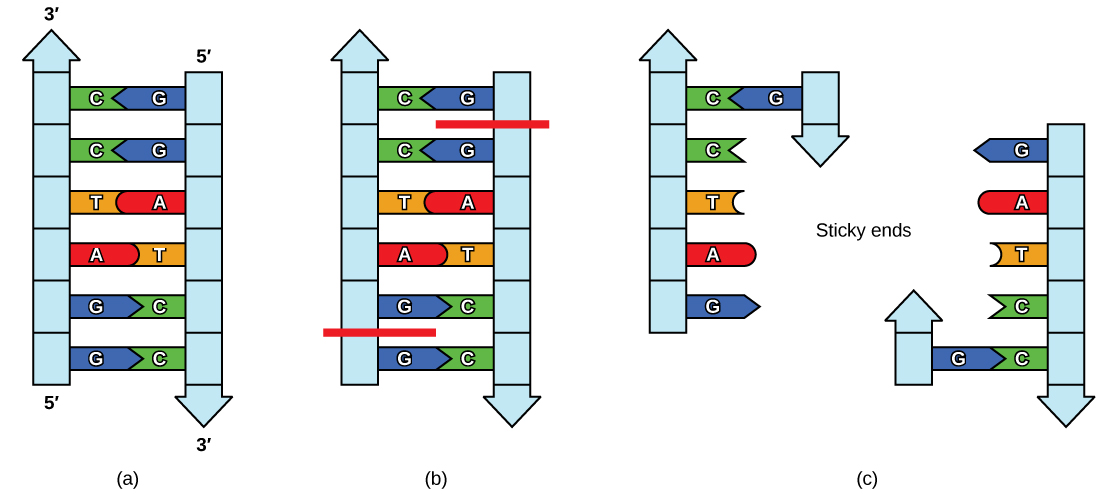
Because these overhangs are capable of coming back together by hydrogen bonding with complementary overhangs on a piece of DNA cut with the same restriction enzyme, these are called “sticky ends.” The process of forming hydrogen bonds between complementary sequences on single strands to form doublestranded DNA is called annealing. Addition of an enzyme called DNA ligase, which takes part in DNA replication in cells, permanently joins the DNA fragments when the sticky ends come together. In this way, any DNA fragment can be spliced between the two ends of a plasmid DNA that has been cut with the same restriction enzyme (Figure \(\PageIndex{3}\)).
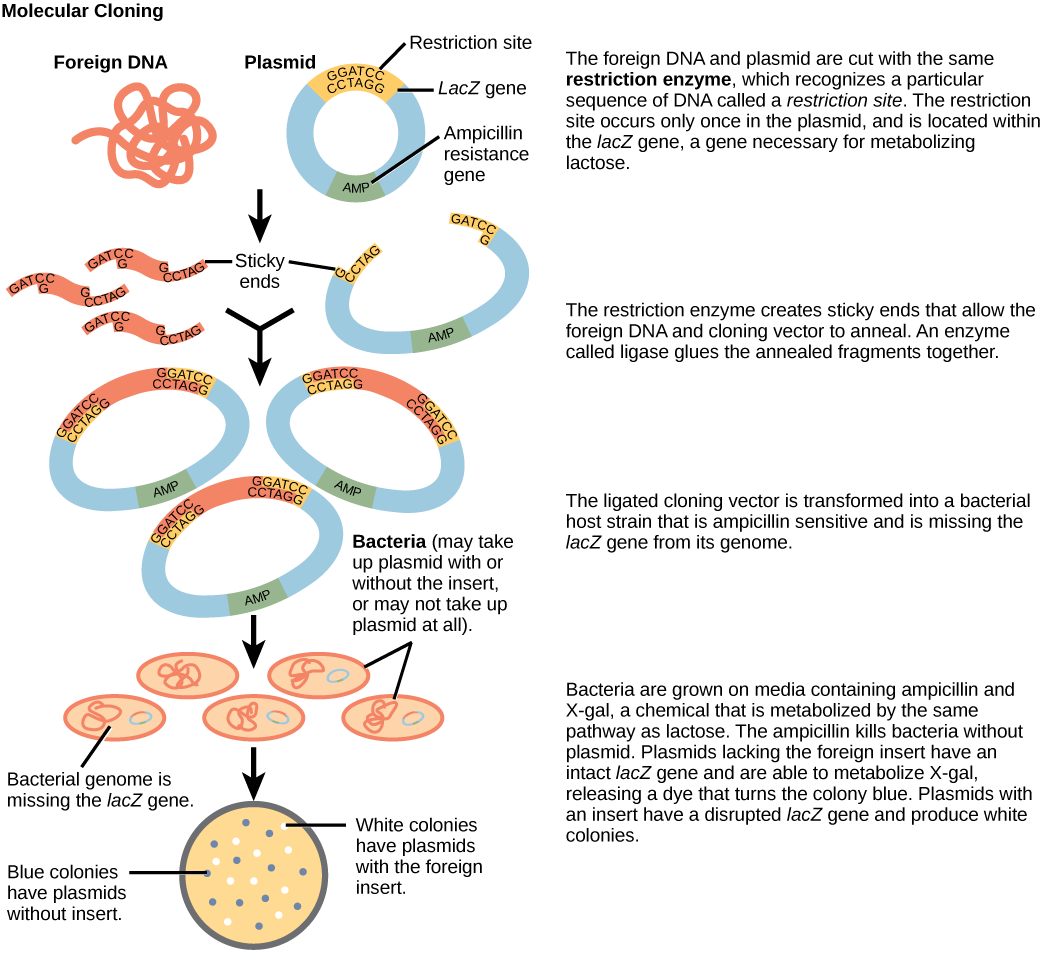
Plasmids with foreign DNA inserted into them are called recombinant DNA molecules because they contain new combinations of genetic material. Proteins that are produced from recombinant DNA molecules are called recombinant proteins. Not all recombinant plasmids are capable of expressing genes. Plasmids may also be engineered to express proteins only when stimulated by certain environmental factors, so that scientists can control the expression of the recombinant proteins.
Cellular Cloning
Unicellular organisms, such as bacteria and yeast, naturally produce clones of themselves when they replicate asexually by binary fission; this is known as cellular cloning. The nuclear DNA duplicates by the process of mitosis, which creates an exact replica of the genetic material.
Reproductive Cloning
Reproductive cloning is a method used to make a clone or an identical copy of an entire multicellular organism. Most multicellular organisms undergo reproduction by sexual means, which involves the contribution of DNA from two individuals (parents), making it impossible to generate an identical copy or a clone of either parent. Recent advances in biotechnology have made it possible to reproductively clone mammals in the laboratory.
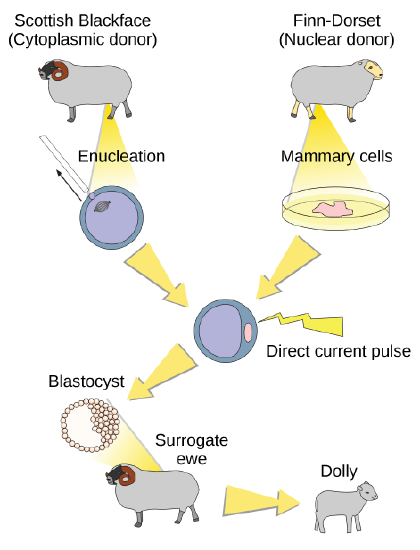
Natural sexual reproduction involves the union, during fertilization, of a sperm and an egg. Each of these gametes is haploid, meaning they contain one set of chromosomes in their nuclei. The resulting cell, or zygote, is then diploid and contains two sets of chromosomes. This cell divides mitotically to produce a multicellular organism. However, the union of just any two cells cannot produce a viable zygote; there are components in the cytoplasm of the egg cell that are essential for the early development of the embryo during its first few cell divisions. Without these provisions, there would be no subsequent development. Therefore, to produce a new individual, both a diploid genetic complement and an egg cytoplasm are required. The approach to producing an artificially cloned individual is to take the egg cell of one individual and to remove the haploid nucleus (Figure \(\PageIndex{4}\)). Then a diploid nucleus from a body cell of a second individual, the donor, is put into the egg cell. The egg is then stimulated to divide so that development proceeds. This sounds simple, but in fact it takes many attempts before each of the steps is completed successfully.
The first cloned agricultural animal was Dolly, a sheep who was born in 1996. The success rate of reproductive cloning at the time was very low. Dolly lived for six years and died of a lung tumor. There was speculation that because the cell DNA that gave rise to Dolly came from an older individual, the age of the DNA may have affected her life expectancy. Since Dolly, several species of animals (such as horses, bulls, and goats) have been successfully cloned.
There have been attempts at producing cloned human embryos as sources of embryonic stem cells. In the procedure, the DNA from an adult human is introduced into a human egg cell, which is then stimulated to divide. The technology is similar to the technology that was used to produce Dolly, but the embryo is never implanted into a surrogate mother. The cells produced are called embryonic stem cells because they have the capacity to develop into many different kinds of cells, such as muscle or nerve cells. The stem cells could be used to research and ultimately provide therapeutic applications, such as replacing damaged tissues. The benefit of cloning in this instance is that the cells used to regenerate new tissues would be a perfect match to the donor of the original DNA. For example, a leukemia patient would not require a sibling with a tissue match for a bone-marrow transplant.
Query \(\PageIndex{1}\)
References
Unless otherwise noted, images on this page are licensed under CC-BY 4.0 by OpenStax.
OpenStax, Biology. OpenStax CNX. May 27, 2016 http://cnx.org/contents/s8Hh0oOc@9.10:8CA_YwJq@3/Cloning-and-Genetic-Engineerin

