27.3B: Molecular Analyses and Modern Phylogenetic Trees
- Page ID
- 13697
The process of establishing relationships between organisms is increasingly becoming more accurate due to advances in molecular analysis.
- Distinguish between morphological and molecular data in creating phylogenetic trees of animals
Key Points
- The construction of phylogenetic trees is now based on similarities and differences within the molecular sources used for analysis which include DNA, RNA, and proteins.
- The ability to use molecular sources as a basis of phylogenetic tree construction has allowed for determination of previously-unknown evolutionary relationships between organisms.
- In addition to the establishment of new relationships within phylogenetic trees, the ability to use molecular sources for analysis has also created an emergence of new phlyums that were previously classified in larger groups.
- Besides identifying molecular similarities and differences between organisms, by assigning a constant mutation rate to a sequence and performing a sequence alignment, it is possible to determine when two organisms diverged from one another.
Key Terms
- monophyletic: of, pertaining to, or affecting a single phylum (or other taxon) of organisms
Modern Advances in Phylogenetic Understanding Come from Molecular Analyses
The phylogenetic groupings are continually being debated and refined by evolutionary biologists. Each year, new evidence emerges that further alters the relationships described by a phylogenetic tree diagram. Previously, phylogenetic trees were constructed based on homologous and analogous morphology; however, with the advances in molecular biology, construction of phylogenetic trees is increasingly performed using data derived from molecular analyses.
Many evolutionary relationships in the modern tree have only recently been determined due to molecular evidence. Nucleic acid and protein analyses have informed the construction of the modern phylogenetic animal tree. These data come from a variety of molecular sources, such as mitochondrial DNA, nuclear DNA, ribosomal RNA (rRNA), and certain cellular proteins. Evolutionary trees can be made by the determination of sequence information of similar genes in different organisms. Sequences that are similar to each other frequently are considered to have less time to diverge, while less similar sequences have more evolutionary time to diverge. The evolutionary tree is created by aligning sequences and having each branch length proportional to the amino acid differences of the sequences. Furthermore, by assigning a constant mutation rate to a sequence and performing a sequence alignment, it is possible to calculate the approximate time when the sequence of interest diverged into monophyletic groups.
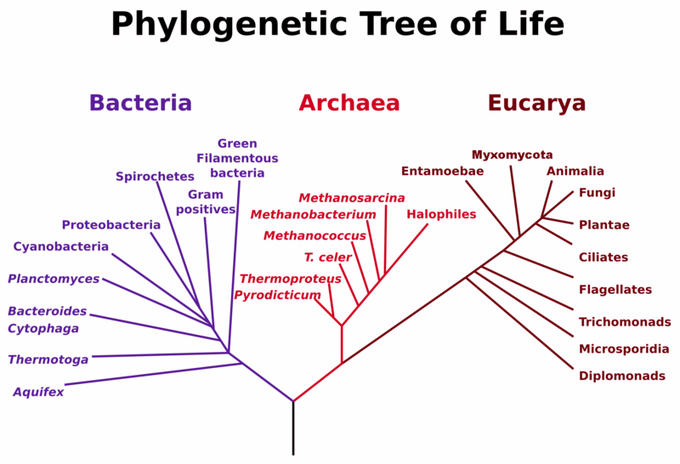
Sequence alignments can be performed on a variety of sequences. For constructing an evolutionary tree from proteins, for example, the sequences are aligned and then compared. rRNA (ribosomal RNA) is typically used to compare organisms since rRNA has a slower mutation rate and is a better source for evolutionary tree construction. This is best supported by research of Dr. Carl Woese that was conducted in the late 1970s. Since the ribosomes are critical to the function of living organisms, they are not easily changed through the process of evolution. Taking advantage of this fact, Dr. Woese compared the minuscule differences in the sequences of ribosomes among a great array of bacteria and showed that they were not all related.
For example, a previously-classified group of animals called lophophorates, which included brachiopods and bryozoans, were long-thought to be primitive deuterostomes. Extensive molecular analysis using rRNA data found these animals to be protostomes, more closely related to annelids and mollusks. This discovery allowed for the distinction of the protostome clade: the lophotrochozoans. Molecular data have also shed light on some differences within the lophotrochozoan group. Some scientists believe that the phyla Platyhelminthes and Rotifera within this group should actually belong to their own group of protostomes termed Platyzoa.
Molecular research similar to the discoveries that brought about the distinction of the lophotrochozoan clade has also revealed a dramatic rearrangement of the relationships between mollusks, annelids, arthropods, and nematodes; a new ecdysozoan clade was formed. Due to morphological similarities in their segmented body types, annelids and arthropods were once thought to be closely related. However, molecular evidence has revealed that arthropods are actually more closely related to nematodes, now comprising the ecdysozoan clade, and annelids are more closely related to mollusks, brachiopods, and other phyla in the lophotrochozoan clade. These two clades now make up the protostomes.
Another change to former phylogenetic groupings because of molecular analyses includes the emergence of an entirely new phylum of worm called Acoelomorpha. These acoel flatworms were long thought to belong to the phylum Platyhelminthes because of their similar “flatworm” morphology. However, molecular analyses revealed this to be a false relationship and originally suggested that acoels represented living species of some of the earliest divergent bilaterians. More recent research into the acoelomorphs has called this hypothesis into question and suggested a closer relationship with deuterostomes. The placement of this new phylum remains disputed, but scientists agree that with sufficient molecular data, their true phylogeny will be determined.
Contributions and Attributions
- OpenStax College, Biology. October 17, 2013. Provided by: OpenStax CNX. Located at: http://cnx.org/content/m44658/latest...ol11448/latest. License: CC BY: Attribution
- Structural Biochemistry/Bioinformatics/Evolution Trees. Provided by: Wikibooks. Located at: en.wikibooks.org/wiki/Structu...volution_Trees. License: CC BY-SA: Attribution-ShareAlike
- orthologous. Provided by: Wiktionary. Located at: en.wiktionary.org/wiki/orthologous. License: CC BY-SA: Attribution-ShareAlike
- homoplasy. Provided by: Wiktionary. Located at: en.wiktionary.org/wiki/homoplasy. License: CC BY-SA: Attribution-ShareAlike
- Tree of life SVG. Provided by: Wikimedia. Located at: commons.wikimedia.org/wiki/Fi...f_life_SVG.svg. License: Public Domain: No Known Copyright
- OpenStax College, Biology. October 17, 2013. Provided by: OpenStax CNX. Located at: http://cnx.org/content/m44658/latest...ol11448/latest. License: CC BY: Attribution
- Structural Biochemistry/Bioinformatics/Evolution Trees. Provided by: Wikibooks. Located at: en.wikibooks.org/wiki/Structu...volution_Trees. License: CC BY-SA: Attribution-ShareAlike
- monophyletic. Provided by: Wiktionary. Located at: en.wiktionary.org/wiki/monophyletic. License: CC BY-SA: Attribution-ShareAlike
- Tree of life SVG. Provided by: Wikimedia. Located at: commons.wikimedia.org/wiki/Fi...f_life_SVG.svg. License: Public Domain: No Known Copyright
- PhylogeneticTree. Provided by: Wikimedia. Located at: commons.wikimedia.org/wiki/Fi...eneticTree.png. License: Public Domain: No Known Copyright


