5.2: DNA Repair
- Page ID
- 1647
Maintaining the Integrity of the Cell's Information: DNA Repair
In the last section we considered the ways in which cells deal with the challenges associated with replicating their DNA, a vital process for all cells. It is evident that if DNA is the master copy of instructions for an organism, then it is important not to make mistakes when copying the DNA to pass on to new cells. Although proofreading by DNA polymerases greatly increases the accuracy of replication, there are additional mechanisms in cells to further ensure that newly replicated DNA is a faithful copy of the original, and also to repair damage to DNA during the normal life of a cell.
All DNA suffers damage over time, from exposure to ultraviolet and other radiation, as well as from various chemicals in the environment. Even chemical reactions naturally occurring within cells can give rise to compounds that can damage DNA. As you already know, even minor changes in DNA sequence, such as point mutations can sometimes have far-reaching consequences. Likewise, unrepaired damage caused by radiation, environmental chemicals or even normal cellular chemistry can interfere with the accurate transmission of information in DNA. Maintaining the integrity of the cell's "blueprint" is of vital importance and this is reflected in the numerous mechanisms that exist to repair mistakes and damage in DNA.
Post-Replicative Mismatch Repair
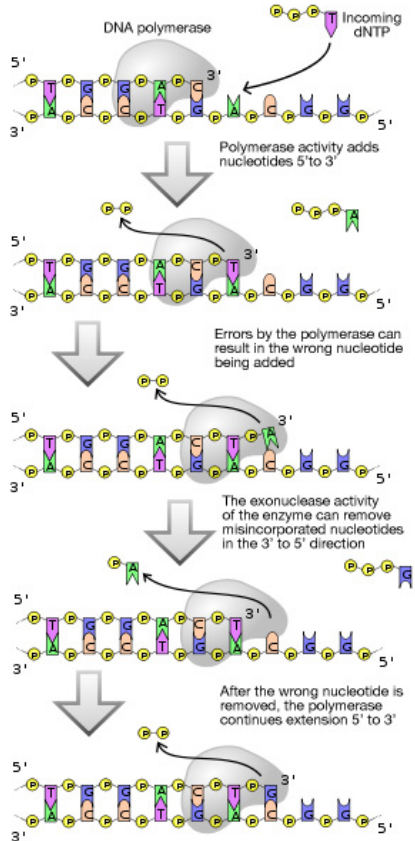
We earlier discussed proof-reading by DNA polymerases during replication. Does proofreading eliminate all errors made during replication. No. While proof-reading significantly reduces the error rate, not all mistakes are fixed on the fly by DNA polymerases. What mechanisms exist to correct the replication errors that are missed by the proof-reading function of DNA polymerases.
Errors that slip by proofreading during replication can be corrected by a mechanism called mismatch repair. While the error rate of DNA replication is about one in \(10^7\) nucleotides in the absence of mismatch repair, this is further reduced a hundred-fold to one in \)10^9\) nucleotides when mismatch repair is functional.
What are the tasks that a mismatch repair system faces. It must:
- Scan newly made DNA to see if there are any mispaired bases (e.g., a G paired to a T)
- Identify and cut out the region of the mismatch.
- Correctly fill in the gap created by the excision of the mismatch region.
Importantly, the mismatch repair system must have a means to distinguish the newly made DNA strand from the template strand, if replication errors are to be fixed correctly. In other words, when the mismatch repair system encounters an A-G mispair, for example, it must know whether the A should be removed and replaced with a C or if the G should be removed and replaced with a T.
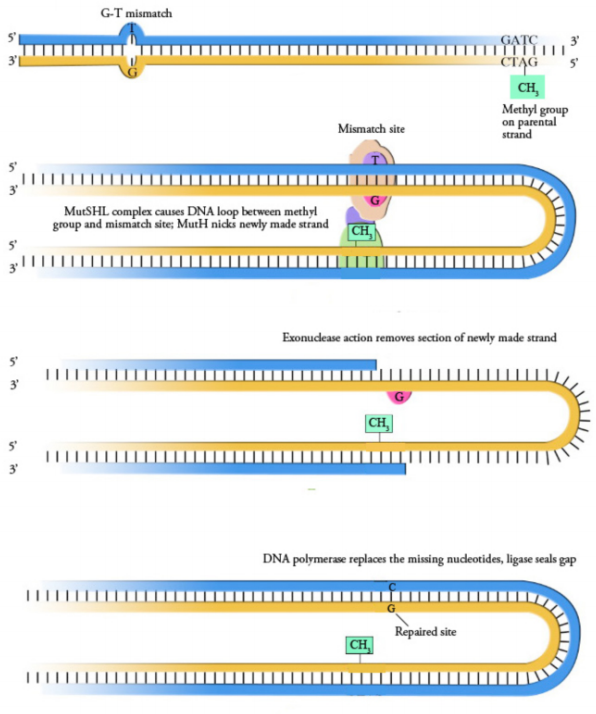
Mismatch repair has been well studied in bacteria, and the proteins involved have been identified. Eukaryotes have a mismatch repair system that repairs not only single base mismatches but also insertions and deletions. In bacteria, mismatch repair proteins are encoded by a group of genes collectively known as the mut genes. Some of the most important components of the mismatch repair machinery are the proteins MutS, L and H. MutS acts to recognize the mismatch, while MutL and MutH are recruited to the mismatch site by the binding of Mut S, to help cut out the region containing the mismatch. A DNA polymerase and ligase fill in the gap and join the ends, respectively.
But how does the mismatch repair system distinguish between the original and the new strands of DNA? In bacteria, the existence of a system that methylates the DNA at GATC sequences is the solution to this problem. E.coli has an enzyme that adds methyl groups on the to adenines in GATC sequences. Newly replicated DNA lacks thismethylation and thus, can be distinguished from the template strand, which is methylated. In Figure \(\PageIndex{2}\), the template strand shown in yellow is methylated at GATC sequences. The mismatch repair proteins selectively replace the strand lacking methylation, shown in blue in the figure, thus ensuring that it is mistakes in the newly made strand that are removed and replaced. Because methylation is the criterion that enables the mismatch repair system to choose the strand that is repaired, the bacterial mismatch repair system is described as being methyl-directed.
Eukaryotic cells do not use this mechanism to distinguish the new strand from the template, and it is not yet understood how the mismatch repair system in eukaryotes "knows" which strand to repair.
Systems to Repair Damage to DNA
In the preceding section we discussed mistakes made when DNA is copied, where the wrong base is inserted during synthesis of the new strand. But even DNA that is not being replicated can get damaged or mutated. These sorts of damage are not associated with DNA replication, rather they can occur at any time.
What causes damage to DNA? Some major causes of DNA damage are:
- Radiation (e.g., UV rays in sunlight, in tanning booths)
- Exposure to damaging chemicals (such as benzopyrene in car exhaust and cigarette smoke)
- Chemical reactions within the cell (such as the deamination of cytosine to give uracil).
This means the DNA in your cells is vulnerable to damage simply from normal sorts of actions, such as walking outdoors, being in traffic, or from the chemical transformations occurring in every cell as part of its everyday activities. (Naturally, the damage is much worse in situations where exposure to radiation or damaging chemicals is greater, such as when people repeatedly use tanning beds or smoke.)
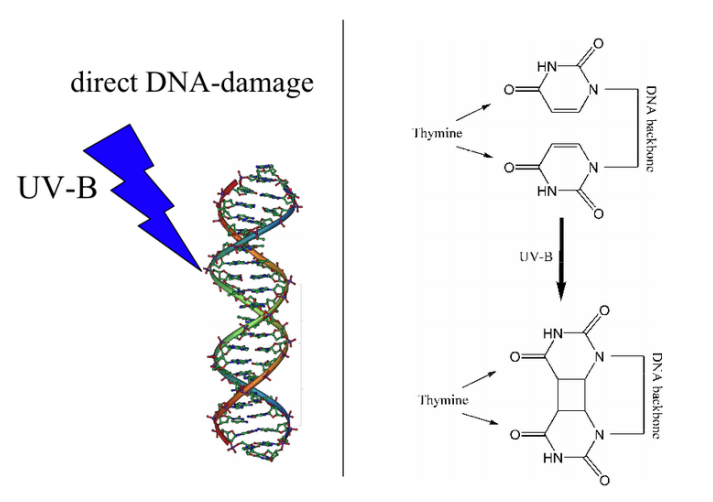
What kinds of damage do these agents cause? Radiation can cause different kinds of damage to DNA. Sometimes, as with much of the damage done by UV rays, two adjacent pyrimidine bases in the DNA will be cross-linked to form pyrimidine dimers (note that we are talking about two neighboring pyrimidine bases on the same strand of DNA). This is illustrated in the figure on the previous page where two adjacent thymines on a single DNA strand are cross-linked to form a thymine dimer. Radiation can also cause breaks in the DNA backbone.
Chemicals like benzopyrene can attach themselves to bases, forming bulky DNA adducts in which large chemical groups are linked to bases in the DNA. The formation of chemical adducts can physically distort the DNA helix, making it hard for DNA and RNA polymerases to copy those regions of DNA.
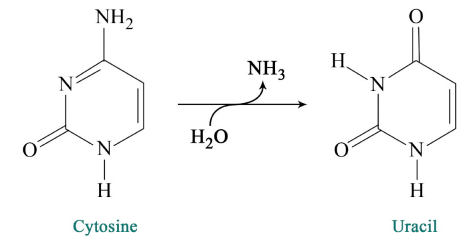
Chemical reactions occurring within cells can cause cytosines in DNA to be deaminated to uracil, as shown in Figure \(\PageIndex{3}\).
Other sorts of damage in this category include the formation of oxidized bases like 8-oxo-guanine. These do not actually change the physical structure of the DNA helix, but they can cause problems because uracil and 8-oxo-guanine pair with different bases than the original cytosine or guanine, leading to mutations on the next round of replication.
How do cells repair such damage? Cells have several ways to remove the sorts of damage described above, with excision repair being a common strategy. Excision repair is a general term for the cutting out and re-synthesis of the damaged region of the DNA. There are a couple of varieties of excision repair:
Nucleotide Excision Repair (NER)
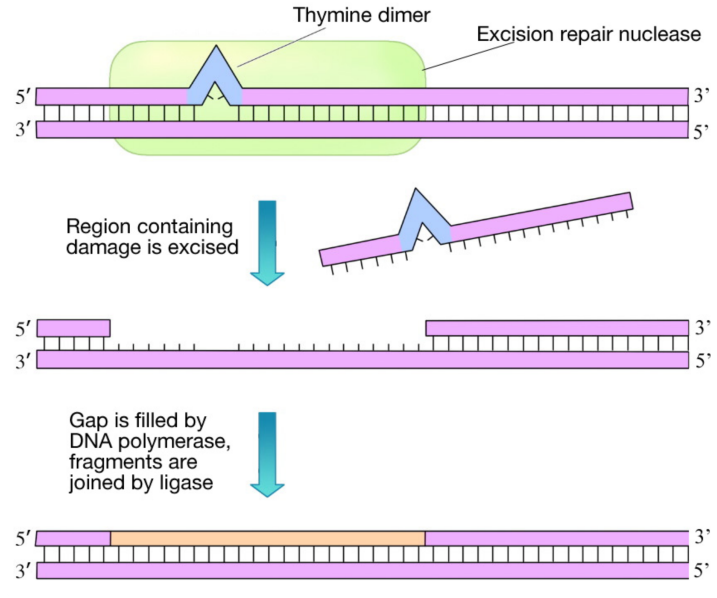
This system fixes damage by chemicals as well as UV damage. As shown in the figure on the previous page, in nucleotide excision repair, the damage is recognized and a cut is made on either side of the damaged region by an enzyme called an excinuclease (shown in green). A short portion of the DNA strand containing the damage is then removed and a DNA polymerase fills in the gap with the appropriate nucleotides. The newly made DNA is joined to the rest of the DNA backbone by the enzyme DNA ligase. In E. coli, NER is carried out by a group of proteins encoded by the uvrABC genes. As you can see, NER is similar, in principle, to mismatch repair. However, in NER, the distortion of the helix, caused by the DNA damage, clearly indicates which strand of the DNA needs to be removed and replaced.
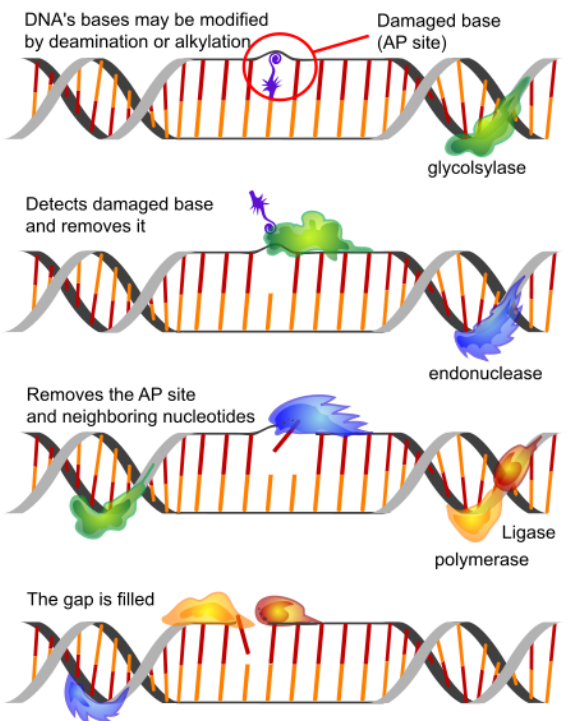
Base Excision Repair (BER)
BER deals with situations like the deamination of cytosine to uracil. As noted earlier, cytosines in DNA sometimes undergo deamination to form the base uracil.
Because cytosines pair with guanines and uracils pair with adenine, the conversion of cytosine to uracil in the DNA would lead to the insertion of an A in the newly replicated strand instead of the G that should have gone in across from a C. To prevent this from happening, uracils are removed from DNA by base excision repair.
In base excision repair, a single base is first removed from the DNA, followed by removal of a region of the DNA surrounding the missing base. The gap is then repaired.
The removal of uracil from DNA is accomplished by the enzyme uracil DNA glycosylase, which breaks the bond between the uracil and the sugar in the nucleotide.
The removal of the uracil base creates a gap called an apyrimidinic site (AP site). The presence of the AP site triggers the activity of an AP endonuclease that cuts the DNA backbone.
A short region of the DNA surrounding the site of the original uracil is then removed and replaced.