2020_SS1_Bis2a_Facciotti_Reading_23
- Page ID
- 34027
Learning objectives associated with 2020_SS1_Bis2a_Facciotti_Reading_23
|
Introduction to gene regulation
Regulation is all about decision making. Gene regulation is, therefore, all about understanding how cells decide about which genes to turn on, turn off or tune up or tune down. In the following section, we discuss some fundamental mechanisms and principles used by cells to regulate gene expression in response to changes in cellular or external factors. This biology is important for understanding how cells adjust changing environments, including how some cells, in multicellular organisms, become specialized for certain functions (e.g. tissues).
Since the subject of regulation is both a deep and broad topic of study in biology, in Bis2a we don't cover every detail - there are way too many. Rather, as we have done for all other topics, we try to focus on (a) outlining some core logical constructs and questions that you must have when you approach ANY scenario involving regulation, (b) learning some common vocabulary and ubiquitous mechanisms and (c) examining a few concrete examples that illustrate the points made in a and
Gene Expression
Introduction
All cells control when and how much each one of its genes is expressed. This simple statement - one that could be derived from observing cellular behavior - brings up many questions that we can decompose using our Design Challenge rubric.
Trying to define "gene expression"
The first thing we need to do, however, is to define what it means when we say that a gene is "expressed". If the gene encodes a protein, one might reasonably propose that "expression" of a gene means how much functional protein the cell makes. But what if the gene does not encode a protein but some functional RNA. Then, in this case, "expressed might mean how much of the functional RNA the cell makes. Yet another person might reasonably suggest that "expression" just refers to the initial step in creating a copy of the genomic information. By that definition, one might want to count how many full-length transcripts are being made. Is it the number of end products encoded by the genomic information or is it the number of reads of the information important to describe "expression" properly. Unfortunately, in practice, we often find that the definition depends on the context of the discussion. Keep that in mind. For the sake of making sure that we are talking about the same thing, in Bis2A we'll try to use the term "expression" primarily to describe the creation of the final functional product(s). Depending on the specific case, the final product may be a protein or RNA species.
The design challenge of regulating gene expression
To drive this discussion from a design challenge perspective, we can formally stipulate that the "big problem" we are interested in understating is that of regulating protein abundance in a cell. Problem: The cell must regulate the abundance of each functional protein. We can then start by posing subproblems:
Let's take a moment, though, first to reload a few ideas. The process of gene expression requires multiple steps depending on what the fate of the final product will be. With structural and regulatory RNAs (i.e. tRNA, rRNA, snRNA, etc.) the process requires that a gene be transcribed and that any needed post-transcriptional processing takes place. With a protein-coding gene, the transcript must also be translated into protein and, if required, modifications to the protein must also be made. Both transcription and translation are multi-step processes, and most of those sub-steps are also potential sites of control.
Some subproblems might therefore be:
- It is reasonable to postulate that there must be some mechanism
( s) to regulate the first step of this multi-step process, the initiation of transcription (just getting things started). So, we could state, "we need a mechanism to regulate the initiation of transcription." We could also turn this into a question and ask, "how can the initiation of transcriptionbe accomplished "? - We can use similar thinking to state, "we need a mechanism for regulating the end of transcription" or to ask "how is transcription terminated?"
- Using this convention we can state, "we need to regulate the initiation of translation and the stop of translation".
- We've talked only about the synthesis of protein and RNA. It is also reasonable to state, "we need a mechanism to regulate the degradation of RNA and protein."
Focusing on transcription
In this course we begin by focusing primarily on examining the first couple of problems/questions, regulating transcription initiation and termination - from genomic information to a functional RNA, either ready as is (e.g. with a functional RNA) or ready for translation. This allows us to examine some fundamental concepts regarding the regulation of gene expression and to examine a few real examples of those concepts in action.
Subproblems for transcription and the activity or RNA polymerase
Let us consider a protein-coding gene and work through some logic. We start by imagining a simple case, where a protein-coding gene is encoded by a single contiguous stretch of DNA. We know that to transcribe this gene an RNA polymerase will need to be recruited to the start of the coding region. The RNA polymerase is not "smart" per se. There needs to be some mechanism, based on chemical logic, to help recruit the RNA polymerase to the start of the protein-coding gene. Likewise, if this process is to be regulated, there needs to be some mechanism, or mechanisms to dictate when an RNA polymerase should be recruited to the start of a gene, when it should not, and/or if it is recruited to the DNA whether it should begin transcription and how many times this process should happen. Note, that the previous sentence, has several distinct subproblems/questions (e.g. when is the polymerase recruited?; if recruited, should it start transcription?; if it starts transcription, how many times should this process repeat?). We can also reasonably infer, that there will need to be some mechanisms to "instruct" (more anthropomorphisms) the polymerase to stop transcription. Finally, since the role of transcription is to create RNA copies of the genome segments, we should also consider problems/questions related to other factors that influence the abundance of RNA, like mechanisms of degradation. There must be some mechanisms, and these mechanisms will probably involve regulating this process.
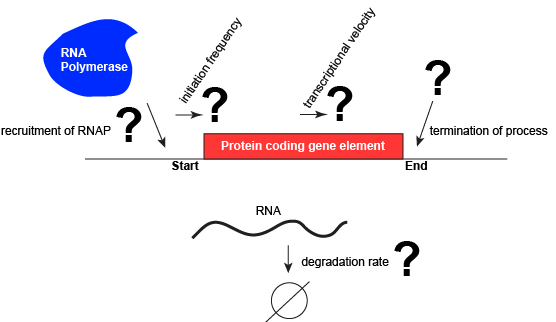
A schematic showing a protein-coding gene and some questions or problems that we need to ask ourselves or problems we need to know solutions for if we are to understand how regulation of the transcriptional portion of the gene's expression is regulated. Attribution: Marc T. Facciotti (own work)
Activation and Repression of Transcription
Some basics
Let us consider a protein-coding gene and work through some logic. We start by imagining a simple case, where a protein-coding gene is encoded by a single contiguous stretch of DNA. We know that to transcribe this gene an RNA polymerase will need to be recruited to the start of the coding region. The RNA polymerase is not "smart" per se. There needs to be some mechanism, based on chemical logic, to help recruit the RNA polymerase to the start of the protein-coding gene. Likewise, if this process is to be regulated, there needs to be some mechanism, or mechanisms to dictate when an RNA polymerase should be recruited to the start of a gene, when it should not, and/or if it is recruited to the DNA, whether it should begin transcription and how many times this process should happen. Note, that the previous sentence, has several distinct subproblems/questions (e.g. when is the polymerase recruited?, if recruited should it start transcription?, if it starts transcription, how many times should this process repeat?). We can also reasonably infer, that there will need to be some mechanisms to "instruct" (more anthropomorphisms) the polymerase to stop transcription.
Recruiting RNA polymerase to specific sites
To initiate transcription, the RNA polymerase must be recruited to a segment of DNA near the start of a region of DNA encoding a functional transcript. The function of the RNA polymerase as described so far, however, is not to bind specific sequences but to move along any segment of DNA. Recruiting the polymerase to a specific site therefore seems contradictory to its usual behavior. Explaining this contradiction requires us to invoke something new. Either transcription can start anywhere and just those events that lead to a full productive transcript do anything useful or something other than the RNA polymerase itself helps to recruit the enzyme to the beginning of a gene. The latter, we now take for granted, is the case.
The recruitment of the RNA polymerase is mediated by proteins called general transcription factors. In bacteria, they have a special name: sigma factors. In archaea they are called TATA binding protein and transcription factor IIB. In eukaryotes, relatives of the archaeal proteins function with many others to recruit the RNA polymerase. The general transcription factors have at least two basic functions: (1) They can chemically recognize a specific sequence of DNA and (2) they can bind the RNA polymerase. Together these two functions of general transcription factors solve the problem of recruiting an enzyme that is otherwise not capable of binding a specific DNA site. In some texts, the general transcription factors (and particularly the sigma factor varieties) are said to be part of the RNA polymerase. While they are part of the complex when they help to target the RNA polymerase they do not continue with the RNA polymerase after it starts transcription.
The DNA site that recruits an RNA polymerase has a special name. We call it a promoter. While the DNA sequences of different promoters need not be exactly the same, different promoter sequences typically have some special chemical properties in common. Obviously, one property is that they can associate with an RNA polymerase. In addition, the promoter usually has a DNA sequence that facilitates the dissociation of the double stranded DNA such that the polymerase can begin reading and transcription the coding region. (Note: technically we could have broken down the properties of the promoter into design challenge subproblems. Here we skipped it, but you should still be able to step backwards and create the problem statements and or relevant questions once you find out about promoters).
In nearly all cases, but particularly in eukaryotic systems the complex of proteins that assembles with the RNA polymerase at promoters (typically called the pre-initiation complex) can number in the tens of proteins. Each of these other proteins has a specific function, but this is far too much detail to dive into for Bis2A.
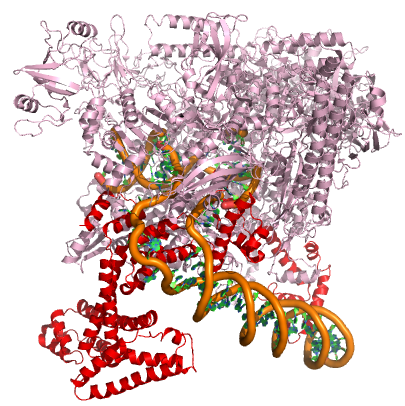
A model of the E. coli pre-initiation complex. The sigma factor is colored red. The DNA is depicted as orange tubes and opposing ring structures. The rest of the pre-initiation complex is colored pink. Note that the DNA has regions of the double helix and an open structure inside the PIC. Attribution: Structure derived from PDB coordinates (4YLN) Marc T. Facciotti (own work)
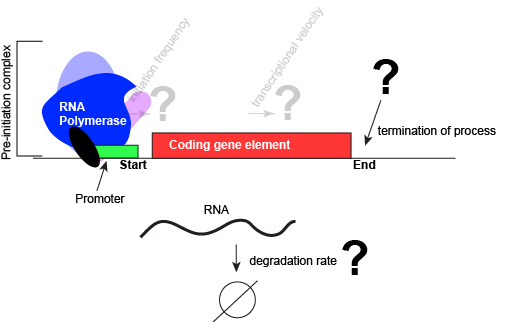
An abstract model of a generic transcriptional unit shown above with the addition of a promoter and PIC. Questions noted earlier that will probably not be covered in Bis2a have been grayed out. Attribution: Marc T. Facciotti (own work)
States of a regulated promoter
Since promoters recruit an RNA polymerase these sites and the assembly of the pre-initiation complex are obvious sites for regulating the first steps of gene expression. At the level of transcription initiation, we often classify promoter into one of three classes. We call the first constitutive. Constitutive promoters are generally not regulated very strongly. Their base state is "on". When the constitutive transcription from a promoter is very high (relative to most other promoters), we will colloquially call that promoter a "strong constitutive" promoter. If the amount of transcription from a constitutive promoter is low (relative to most other promoters) we call that promoter a "weak constitutive" promoter.
A second way to classify promoters by the use of the term activated or equivalently induced. These interchangeable terms are used to describe promoters sensitive to some external stimulus and respond to said stimulus by increasing transcription. Activated promoters have a base state exhibits little to no transcription. Transcription is then "activated" in response to a stimulus - the stimulus turns the promoter "on".
Finally, the third term used to classify promoters is by the use of the term repressed. These promoters also respond to stimuli but do so by decreasing transcription. The base state for these promoters is some basal level of transcription, and the stimulus acts to turn down or repress transcription. Transcription is "repressed" in response to a stimulus - the stimulus turns the promoter "off".
The examples given above assumed that a single stimulus acts to regulate promoters. While this is the simplest case, many promoters may integrate different information and may be alternately activated by some stimuli and repressed by other stimuli.
Possible NB Discussion  Point
Point
In Bis2A, you have learned about how important it is for the cell to be able to regulate its biology, with several examples of regulatory mechanisms. Given this, can you explain why it is advantageous to still have constitutive promoters that are generally not regulated very strongly? What kinds of genes would you expect to have a strong constitutive promoter? What about a weak constitutive promoter?
Transcription factors help to regulate the behavior of a promoter
How are promoters sensitive to external stimuli? Mechanistically, in both activation and repression require regulatory proteins to change the transcriptional output of the gene being observed. The proteins responsible for helping to regulate expression are generally called transcription factors. The specific DNA sequences bound by transcription factors are often called operators and most times the operators are very close to the promoter sequences.
Here's where the nomenclature gets potentially confusing - particularly when comparing across bacterial and eukaryotic research. In bacterial research, if the transcription factor acts by binding DNA and the RNA polymerase in a way that increases transcription, then it is typically called an activator. If, by contrast, the transcription factor acts by binding DNA to repress or decrease transcription of the gene then it is called a repressor.
Why are the classifications of activator and repressor potentially problematic? These terms describe the proteins with idealized single functions. While this may be true with some transcription factors, in reality other transcription factors may act to activate gene expression in some conditions while repressing in other conditions. Some transcription factors will act to modulate expression either up or down depending on context rather than shutting transcription "off" or turning it completely "on". To circumvent some of this confusion, some of your instructors prefer to avoid using the terms activator and repressor and instead prefer to discuss the activity of transcription various transcription factors as either a positive or a negative influence on gene expression in specific cases. If we use these terms, you might hear your instructor saying that the transcription factor in question ACTS LIKE/AS a repressor or that it ACTS LIKE/AS an activator, taking care not to call it simply an activator or repressor. It is more likely, however, that you will hear them say that a transcription factor is acting to positively or negatively influence transcription.
CAUTION: Depending on your instructor, you may cover a few real biological examples of positive and negative regulatory mechanisms. These specific examples will use the common names of the transcription factors - since the examples will typically be drawn from the bacterial literature, the names of the transcription factors may include the terms "repressor" or "activator". These names are relics of when they were first discovered. Try to spend more time examining the logic of how the system works than trying to commit to memory any special properties of that specific protein to all other cases with the same label. Just because we label a protein as a repressor does not mean that it only acts as a negative regulator in all cases or that it interacts with external signals in the same was as the example.
Possible NB Discussion  Point
Point
What types of interactions do you think happen between the amino acids of the transcription factor and the double helix of the DNA? How do transcription factors recognize their binding site on the DNA?
Allosteric Modulators of Regulatory Proteins
The activity of many proteins, including regulatory proteins and various transcription factors, can
Let us imagine a negative transcriptional regulator. In the most simple case we've considered so far, transcription of a gene with a binding site for this transcription factor would be low when the TF is present and high when the TF is absent. We can now add a small molecule to this model. Here the small molecule can bind the negative transcriptional regulator through sets of complementary hydrogen and ionic bonds. In this first example we will consider the case where the binding of the small molecule to the TF induces a conformational change to the TF that severely reduces its ability to bind DNA. If this is the case, the negative regulator - once bound by its small molecule - would release from the DNA. This would
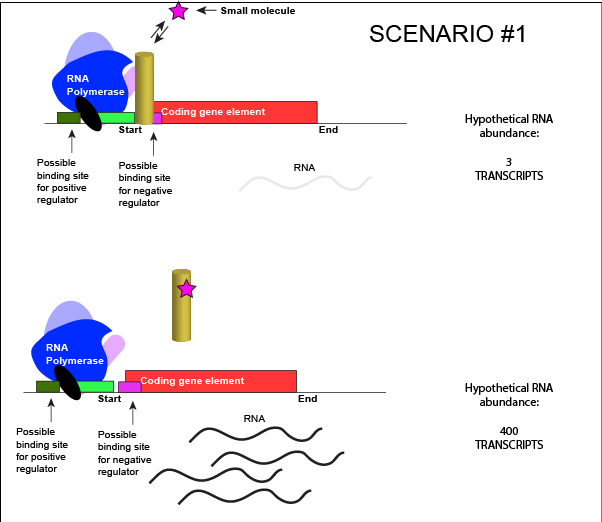
An abstract model of a generic transcriptional unit regulated by a negative regulator whose activity
We can consider a second model for how a negatively acting TF might interact with a small molecule. Here, the TF alone cannot bind its regulatory site to the DNA. However, when a small molecule binds to the
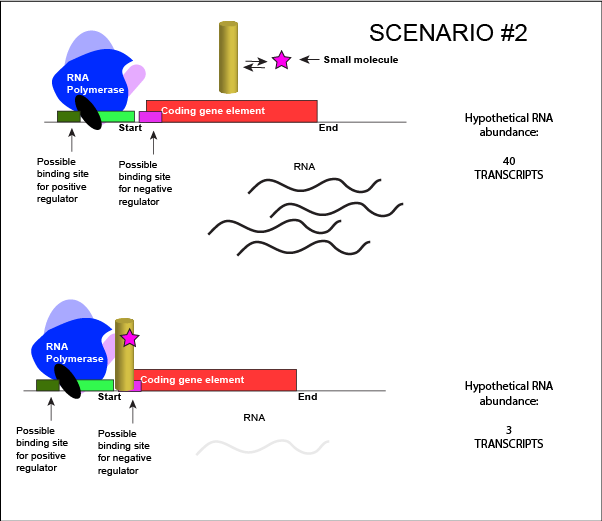
An abstract model of a generic transcriptional unit regulated by a negative regulator whose activity
Note how the activity of the TF can
In both cases proposed above, the binding of a small molecule to a TF
Is it positive or negative regulation?
Resolving a common point of confusion
It is not uncommon for many Bis2a students to
However, in both examples above, the TF
Note that sometimes a TF may act as a positive regulator at one promoter and negative regulator at a different promoter so describing the behavior of the TF on a per case basis is often important (reading too much from the name it has
A genetic test for positive or negative regulatory function of a TF
How does one determine if a regulatory protein functions in a positive or negative way? A simple genetic test is to ask "what happens to expression if the regulatory protein is absent?" This can
Termination of Transcription and RNA degradation
Termination of transcription in bacteria
Once a gene
Rho-independent termination
Termination of transcription in eukaryotes
The termination of transcription is different for the different polymerases. Unlike in prokaryotes, elongation by RNA polymerase II in eukaryotes takes place
Termination of transcription in archaea
Termination of transcription in the archaea is far less studied than in the other two domains of life and
Degradation of RNA
The lifetime of different RNA species in the cell can vary dramatically, from seconds to hours. The mean lifetime of
Tuning termination
Like all other biological processes, the termination of transcription is not perfect. Sometimes, the RNA polymerase can read through terminator sequences and transcribe adjacent genes. This is
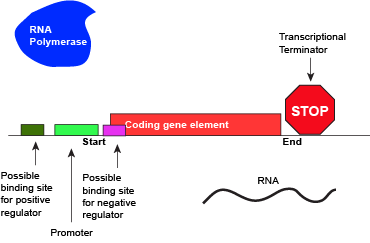
An abstract model of a full basic transcriptional unit and the various "parts" encoded on the DNA that may influence the expression of the gene. We expect students in Bis2A to
Summary of gene regulation
In the preceding text we have examined several ways to solve some design challenges associated with regulating the amount of

