5.2: Changes in Chromosome Structure
- Page ID
- 25740
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Learning Objectives
- Distinguish between the 4 types of chromosomal rearrangements and their effects on gene dosage and meiosis.
If the chromosome is altered, but still retains the three critical features of a chromosome (centromeres, telomeres, and origin of replication), it will continue to be inherited during subsequent cell divisions, however the daughter cell may not retain all the genes. For example, if a segment of the chromosome has been lost, the cell may be missing some genes. The causes and consequences of chromosome structural abnormalities are described here. They all involve breaks in the chromosomal DNA backbone.
Cause #1: Incorrect Repair of Double Strand DNA Breaks During Interphase
A chromosome is a very long but very thin molecule. In the phosphodiester backbone there are only two covalent bonds holding each base pair to the next. If one of these covalent bonds is broken the chromosome will still remain intact, although a DNA ligase will be needed to repair the nick. Problems arise when both strands are broken at or near the same location. This double strand break will cleave the chromosome into two independent pieces. Because these events do occur in cells, there is a repair system called the non-homologous end joining (NHEJ) system to fix them. Proteins bind to each broken end of the DNA and reattach them with new covalent bonds. This system is not perfect and sometimes leads to chromosome rearrangements (see next section).
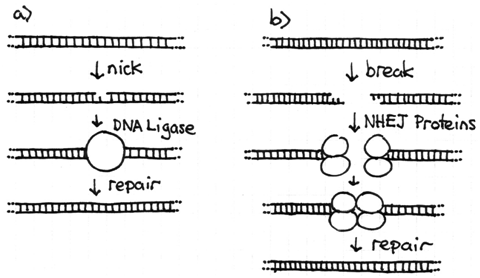
The NHEJ system proteins only function if required. If the chromosomes within an interphase nucleus are all intact, the system is not active. The telomeres at the natural ends of chromosomes prevent the NHEJ system from attempting to join the normal ends of chromosomes together. If there is one double-strand break, the two broken ends can be recognized and joined. However, if there are two (or more) double-strand breaks at the same time there will be four broken ends in total. The NHEJ system proteins recognize the breaks but do not have the ability to identify exactly which ends should be joined to restore the original chromosome. Thus, NHEJ may join the ends together correctly, but if they do not the result is a chromosome rearrangement.
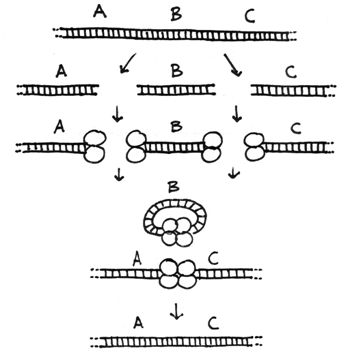
The Four Types of Chromosome Rearrangements
Errors during the repair of multiple double-strand breaks can cause four types of chromosome rearrangements. The type of chromosome rearrangement is dependent upon where the two breaks were originally and how they are rejoined. First double-strand DNA break occurs in one site; a second DNA break occurs and the NHEJ proteins mend the damage incorrectly by joining the ends.
There are four major types of rearrangements:
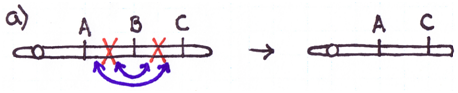
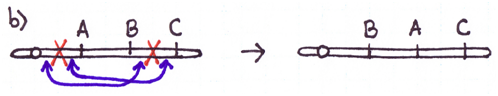
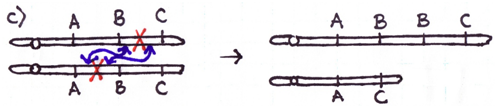
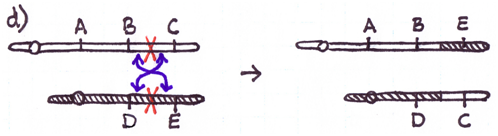
Cause #2: Incorrect Crossovers During Meiosis
Meiotic crossovers occur at the beginning of meiosis for two reasons. They help hold the homologous chromosomes together until separation occurs during anaphase I. They also allow recombination to occur between linked genes. The event itself takes place during prophase I when a double strand break on one piece of DNA is joined with a double strand break on another piece of DNA and the ends are put together. Most of the time the breaks are on non-sister chromatids and most of the time the breaks are at the same relative locations.
Problems occur when the wrong pieces of DNA are matched up along the chromosomes during crossover events. This can happen if the same or similar DNA sequence is found at multiple sites on the chromosomes. For example, if there are two Alu transposable elements on a chromosome. When the homologous chromosomes pair during prophase I the wrong Alu sequences might line up. A crossover may occur in this region. If so, when the chromosomes separate during anaphase I one of the chromatids will have a duplication and one will have a deletion. Ultimately, of the four cells produced by this meiosis, two will be normal, one will have a chromosome with extra genes, and one will have a chromosome missing some genes. Errors of this type can also cause inversions and translocations.
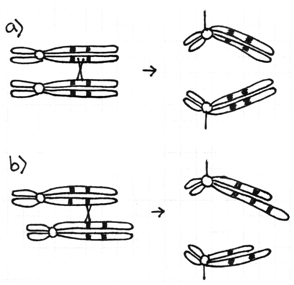
Consequence #1: Abnormal pairing at Meiosis
DNA forms loops to achieve pairing when chromosomes are rearranged
Homologous regions of chromosomes pair at meiosis I (prophase I). With rearranged chromosomes this can lead to visible abnormalities and segregation abnormalities.
Deletion chromosomes will pair up with a normal homolog along the shared regions and at the missing segment, the normal homolog will loop out (nothing to pair with) to form a deletion loop. This feature can be used to locate the deletion cytologically. The deleted region is also pseudo-dominant, in that the cell no longer has two copies of the genes in this region, therefore mutant recessive alleles may produce a phenotypic change.
When an inversion chromosome is paired up in meiosis there is an inversion loop formed. If there is a crossover within the loop then abnormal products will result and abnormal, unbalanced gametes will be produced. For example, a crossover event within the loop of a paracentric inversion will lead to a dicentric product that will break into deletion products and produce unbalanced gametes. Similarly, with a pericentric inversion, a crossover event leads to duplicate/deletion products that are unbalanced.
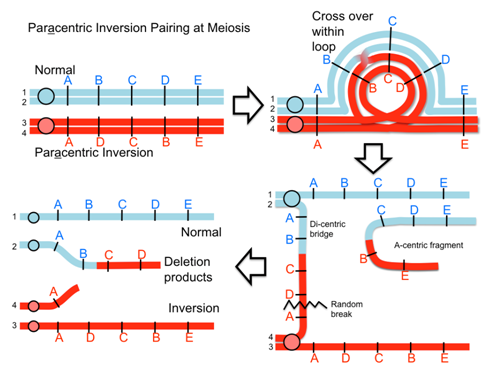
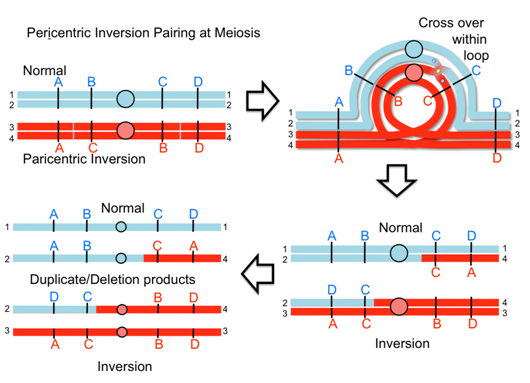
Video explanation: Crossing over in inversions
Translocated chromosomes pair with more than one homologous chromosome
For translocations, a consequence for the two chromosomes involved is that when they pair at meiosis both replicated chromosome pairs will be together. During meiosis I, homologous chromosomes separate. The four chromosomes can segregate in different ways, depending on which centromeres are pulled to each pole during anaphase. This set of paired, replicated chromosomes can segregate as Alternate (balanced) where both non-translocated (N1 and N2) chromosomes go to one pole and both translocated chromosomes go to the other pole. A gamete that contains the two translocated (T1 and T2) chromosome still contains one full copy of both chromosomes. However, the chromosomes may segregate as Adjacent-1 (unbalanced) where a cells gets one normal and one translocation chromosome; each gamete will contain extra copies of some genes and lack copies of other genes. Each of these possibilities occur with approximately equal frequency and thus about half the gametes will be unbalanced.
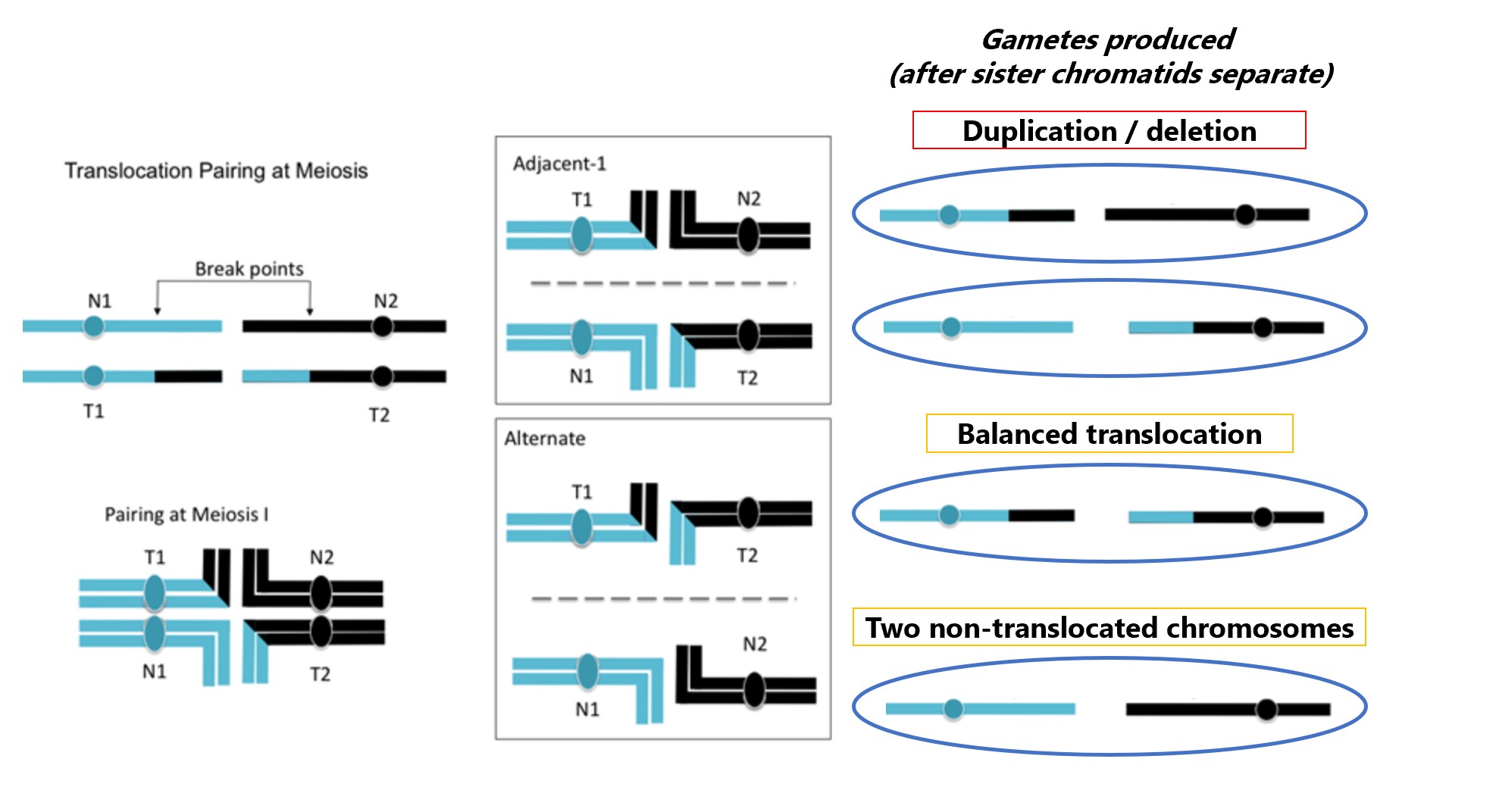
Figure \(\PageIndex{6}\): A reciprocal translocation pairing at meiosis. There are two main avenues for segregation: Adjacent-1 and Alternate. Adjacent-1 results in duplication and deletion for part of the chromosome segments. Alternate segregation . (Modified by SWL from Original-Locke-CC:AN)
Consequence #2: Decreased Viability
All of the chromosome rearrangements shown above produce functional chromosomes. Each has one centromere, two telomeres, and thousands of origins of replication. Because inversions and translocations do not change the number of genes in a cell or organism they are said to be balanced rearrangements. Unless one of the breakpoints occurred in the middle of a gene the cells will not be affected. On the other hand, deletions and duplications are unbalanced rearrangements. The larger they are (more genes involved) the more disruption they cause to the proper functioning of the cell or organism.
Note about gene dosage
Having too much or too little gene activity for a large number of genes can disrupt the cellular metabolism to generate a phenotype or reduce viability.
Consequence #3: Decreased Fertility
Recall that during meiosis I homologous chromosomes pair up. If a cell has a chromosome with a rearrangement this chromosome will have to pair with its normal homolog.
Cells heterozygous for balanced rearrangements actually have more difficulties in prophase I. There are different ways they might pair during prophase I, but if a crossover occurs in the inverted region the result will be unbalanced gametes. Embryos made with unbalanced gametes rarely survive. The consequence is that the heterozygous organism will have reduced fertility.
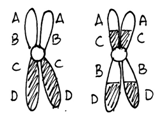
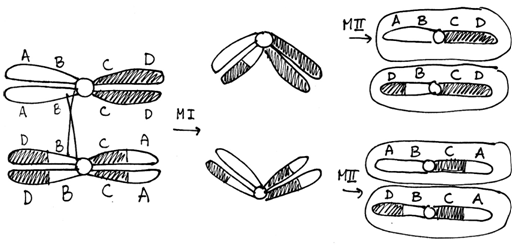
Figure \(\PageIndex{8}\): Meiosis in a cell heterozygous for the chromosomes shown in Figure 9.11. Note that of the four gametes one has a deletion of the A gene and a duplication of the D gene while another gamete has a duplication of A and a deletion of D. (Original-Harrington-CC:AN)
Note that an organism homozygous for this inversion chromosome will not be affected in this way because no loops are formed. The chromosomes can pair along their entire length and crossovers will not produce any unbalanced gametes. This is a general property of inversions and translocations. In heterozygotes there are problems during meiosis, that result in a lot many unbalanced gametes and an overall reduction in fertility. In homozygotes the rearranged chromosomes pair with one another just fine and there is no effect on fertility.
Consequence #4: Cancer
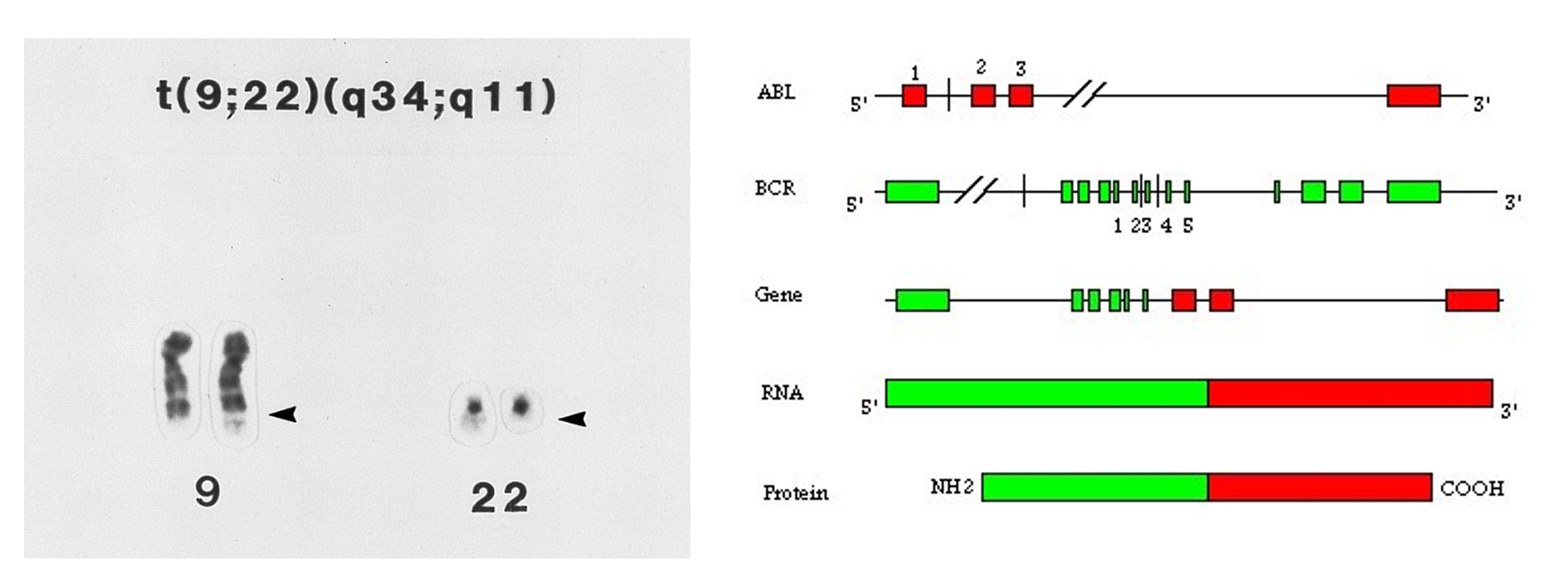
Some chromosome rearrangements have breakpoints within genes leading to the creation of hybrid genes – the first part of one gene with the last part of another. If the hybrid gene inappropriately promotes cell replication, the cell can become cancerous. One example of such a translocation t(9:22) is the cause of chronic myelogenous leukemia (CML). Association of this translocation in cancer was first suggested in a brief letter 1960 (Nowell and Hungerford, Science 132:1497). The translocation creates a fusion gene with the BCR gene promoter and 5' region and the ABL 3' end. When transcribed and translated, this fusion protein is a constitutively active kinase.
Consequence #5: Evolution
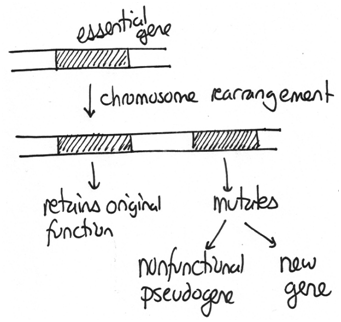
Those chromosome changes that duplicate genes are important for evolution. If an organism has an extra copy of important genes, one gene can be retained for their original function while others can mutate and potentially acquire new functions. For example, multiple copies of the globin genes are found in mammals and have unique functions.
Chromosome rearrangements that decrease fertility are also important for the origin of new species. If a rearrangement becomes common in a small isolated population, that population has 100% fertility if they mate within their group, but a reduced fertility if they mate with members of the larger population. As rearrangements accumulate the small population will become more and more reproductively isolated. When members are incapable of forming viable, fertile offspring with the original population the group will have become a new species.
Contributors and Attributions
Dr. Todd Nickle and Isabelle Barrette-Ng (Mount Royal University) The content on this page is licensed under CC SA 3.0 licensing guidelines.


