Research Paper: The following questions are based on data, graphs, and figures from the following article: Naphthoquinone-mediated Inhibition of Lysine Acetyltransferase KAT3B/p300, Basis for Non-toxic Inhibitor Synthesis. Mohankrishna Dalvoy Vasudevarao et al. The Journal of Biological Chemistry, 289, 7702-7717. March 14, 2014 . doi: 10.1074/jbc.M113.486522
BACKGROUND
Double-standed DNA is a polyanion as every phosphate in the backbone is negatively charged. To pack it into chromosome in the nucleus, DNA is complexed to a variety of proteins. In the first stage of packing, DNA is wound around a core of positively charged histones protein, forming a nucleosome. The size of the nucleosome is about the size of RNA polymerase.
Nucleosome: Jmol14 (Java) | JSMol (HTML5)
A. For RNA polymerase to transcribe DNA that is packed in a nucleosome, what must likely happen to the nucleosome complex?
The DNA must unwind from the histone core of then nucleosome to allow binding of transcription factors and ultimately RNA polymerase to the regulatory regions of the DNA (promoter, etc). Otherwise DNA-dependent RNA polymerization will not occur.
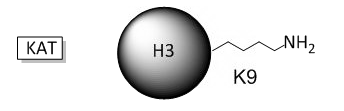
B. The phenotype of a cell depends on the DNA sequence of the cell and the subset of genes that are expressed by that cell. The emerging field of epigenetics describes how factors other than the DNA sequence control cell phenotype and heritability of traits. Key features in epigenetic regulation of gene transcription are chromatin remodeling, packing and covalent modification of DNA. Chromatin packing at the level of DNA:histone interactions are critical. Key to these interactions are the state of covalent modification of lysine (K) side chains in protein by histone acetylases (HATs) which are also called lysine acetylases (KATs) and histone deacetylases (HDACs). Two chemical modifications of lysine side chains in histones are shown below.

a. Offer a chemical explanation of why structure b will not be acetylated by KAT.
The epsilon amino group of Lys acts as a nucleophile when it attacked the electrophilic carbonyl C of acetyl CoA. A protonated amine is not nucleophilic.
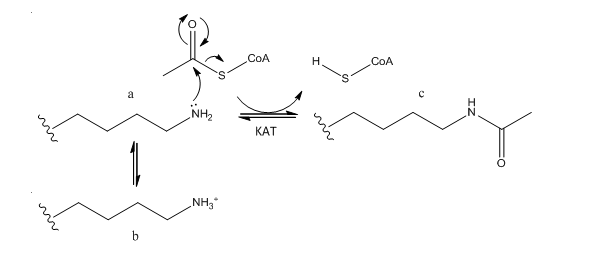
b. What affect might KAT modification of histone lysine side chains have on transcriptional competency of DNA in nucleosomes?
The pKa of Lys in aqueous solution is around 9.0. In a protein it may vary from that, but at physiological pH, it would be mainly protonated with a positive charge. On acetylation, histone Lys side chains would lose any positive charge, weakening Coulombic interactions with DNA, which would promote unwinding of DNA from the histone core of the nucleosome. This would increase the transcriptional competency of the DNA.
1,4-Naphthoquinone derivatives are naturally occurring compounds that are present in the Plumbago and Diospyros plant genera and have a variety of biological activities. One such derivative, plumbagin, is a potent inhibitor of a particular lysine acetyltransferase, KAT p300 lysine acetyltransferases 3B/3A (p300/CBP). Some of the inhibitors are toxic to cells as they react with free RSH groups in cells and generate reactive oxygen species (ROS). An ideal drug would be nontoxic and an effective inhibitor/activator of a specific enzyme. Investigators synthesized a derivative (PTK1) of plumbagin which inhibit p300 and with much lower toxicity that plumbagin. How does it work?
Fig 1A: To study the toxicity of plumbagin, investigators dissolved it in the solvent DMSO + N-acetylcysteine (NAC) which scavenges ROS. They added these reagents to three different human cells lines, SHSY-5Y, HEK293, and HeLa S3 as shown below.
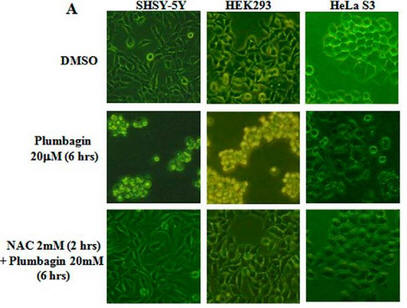
a. What effect did plumbagin have on cell toxicity?
Addition of plumbagin to the cells (row 2) clearly changes cell morphology, properties and numbers. One can infer that it leads to cell death.
b. What effect did NAC have plumbagin cell effects?
It is clear from Figure 1A row C that NAC prevents/interferes with the cytotoxic effects of plumbagin.
c. What did the investigators conduct an experiment just DMSO alone?
Plumbagin was dissolve in DMSO before it was added to the cells. To determine any observed cellular effects did not derive from DMSO, they conducted this experiment. DMSO, the “vehicle” that carried the drug, served as the control in this experiment.
Fig 1B. It has been shown that on DNA oxidative damage or breaks, one of the histones, H2AX is phosphorylated by a kinase, pI3K. Cells were treated as describe below (Plu = plumbagin_ and subjected to SDS-polyacrylamide gel electrophoresis. After the electrophoresis, a nitrocellluose membrane was placed on top and the proteins in the gel were electrophoresed into the membrane (a technique called Western blotting. Individual protein band were visualized by adding antibodies that recognized phosphorylated H2AXg, or histone H3 (without modification), followed by reagents that produce a dark band. Interpret the results. Why did the investigators do blots for unmodified H3?
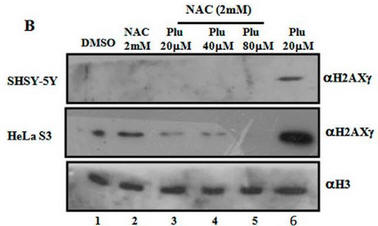
When Plumbagin was added to the cells without NAC (red boxed lanes), there was robust phosphorylation of H2AX gamma. Addition of NAC prevented the phosphorylation that arose with Plu in the absence of NAC. The third row across is another control which assure that the same amount of samples were loaded on the gel. H3 would not be expected to change in the short time course of the experiment.
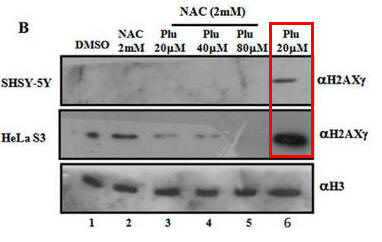
Fig 1C. What affect did plumbagin have on histone acetylation? Immuoblots were performed after plumbagin treatment in the presence or absence of NAC in the indicated cell lines as above using antibodies that recognized acetylated Lys 9 on H3 (H3K9), or histone H3 (without modfication). Does Plu inhibit H3K9 acetylation? How does NAC affect Plumbagin effect on acetylation?

Plu appears to decrease the intensity of the bands when added in the absence of NAC. This suggests the Plu inhibits H3K9 acetylation by some mechanism. As in the previous experiment, NAC seemed to completely antagonize the effect of Plu in that it prevented Plu from decreasing acetylation (compare the red and green boxed lanes). NAC inhibited the inhibitory effects of Plu on H3K9 acetylation.
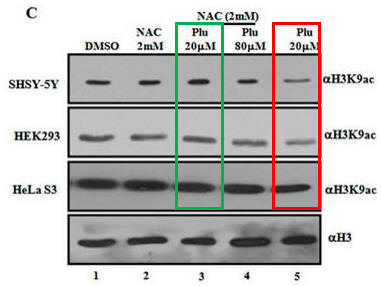
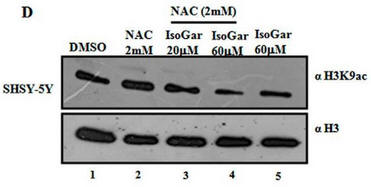
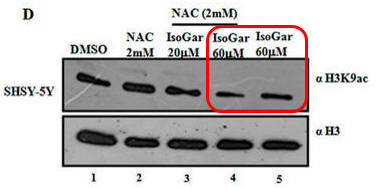
Fig 1D: Another inhibitor of H3K9 acetylation, isogarcinol (IsoGar) was studied by western blotting. Did NAC have same effect on H3K9 acetylation of cells treated with IsoGAr as those treated with Plu, shown in Fig 1C?
HINT | ANS
In contrast to the Blumgartin, another inhibitor, IsoGar, clearly inhibits acetylation of H3K9 but this inhibition is not relieved by NAC (see the red boxed lanes below).
NAC seems to behave differently in the presence of two acetylation inhibitors. These results offer two possibilities for its effects. • ROS could lead to decrease acetylation of histones, and NAC, by decreasing ROS, restore acetylation. • Alternatively, plumbagin reacts with the RSH of NAC, which decreases it effect on KAT and restores acetylation.
Draw cartoon models that would reflect these two different scenarios using a line with an arrow to indicate activation or promotion of an activity, and a line with a blunt end to show inhibition. Also use and these symbols:
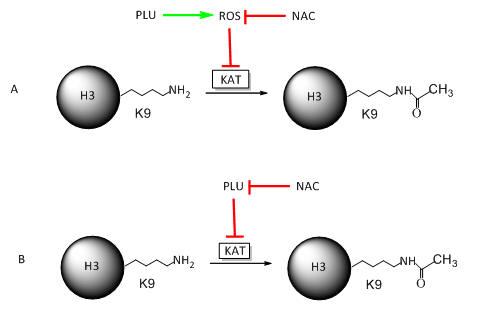
Investigators found no increase in ROS with purified KAT in a cell free in vitro produced no free radicals. Which of your models is more likely?
Model B above since Model A would lead to increased ROS such as peroxide or superoxide
Fig 2AB. To simplify study of these effects, the investigators performed in vitro (cell free) assays in which no ROS (superoxide, hydrogen peroxide) would be produced. They studied two different KATs, p300 (A) and CBP (B) using HeLa cell core histones, with 100 μm plumbagin (lanes 5–7 in A and B) either in the presence or absence of NAC, DTT, and another reducing agent Trolox (lane 8). Interpret the results. What part of NAC and DTT appear important in their effects on plumbagin inhibition.
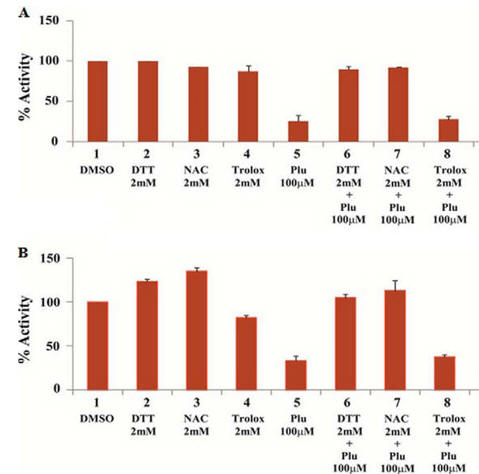
Lanes 5 and 7 and Lane 5 and 8 show that both NAC and DTT (both of which have thiols) relieve the inhibitory effect of blumbagin on KAT activity. Lanes 5 and 8 show that Trolox, which does not have a thiol group, does not relieve blumbagin inhibition.
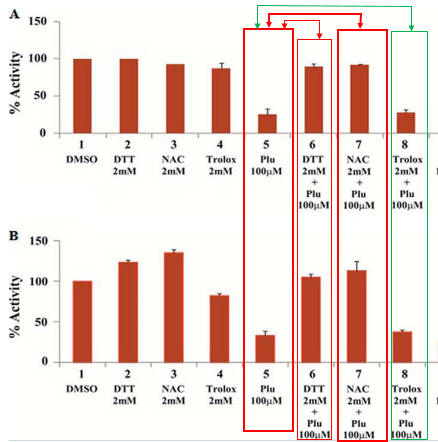
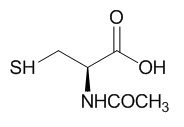
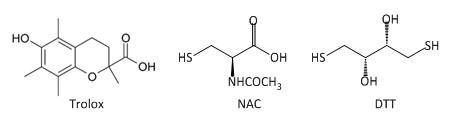
Fig 3A: These data suggest a chemical reaction of NAC and DTT with plumgartin. The generic structure of 1,4-Naphthoquinone derivatives, including plumgartin, are shown below, along with NAC and DTT.
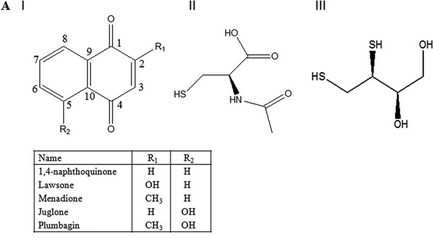
Fig 3B; The thiol group on NAC is likely to react with a sp2 C, ultimately leading to a product which retains conjugation if possible. Draw a mechanism showing how NAC might react with the structure below, and show the final products.
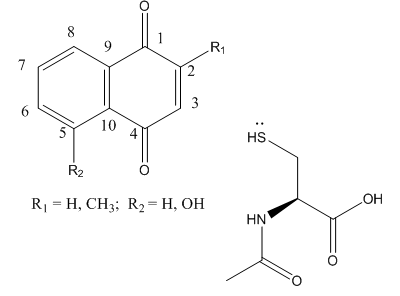
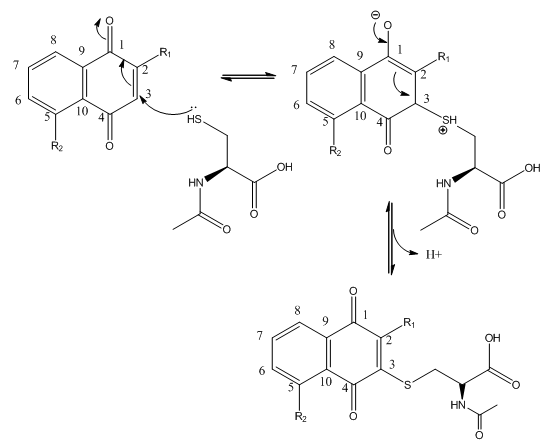
Fig 4 A: A variety of 1,4-Naphthoquinone derivatives were tested to see their effect on p300 activity in the presence and absence of NAC. One, lawsone, whose structure is given below, showed no inhibitory effects, as shown below. Give a likely chemical explanation for this observation.
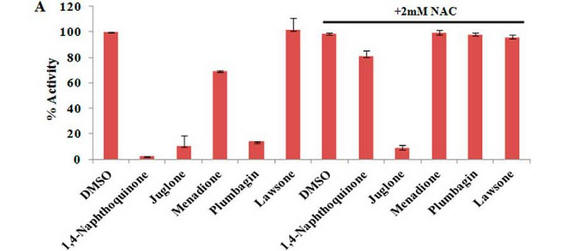
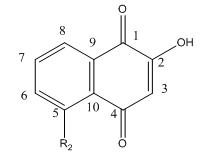
Lawsone can exists in the following tautomeric forms and hence is not likely to react with a thiol at C3.
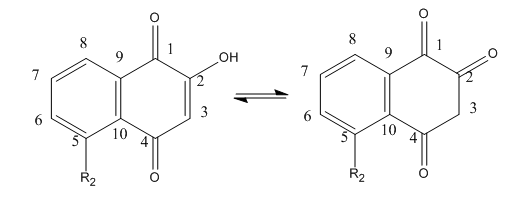
Fig 4C. If they reacted with free thiols, then the naphthoquinone derivatives should have an effect on the concentration of free thiols in an in vitro assays. Results of one such assay are shown below.
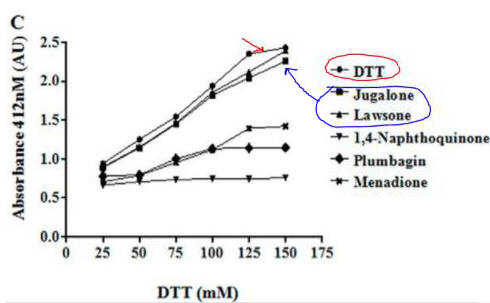
Compare this graph with the proceeding one. What conclusion can you draw?
Lawson, as predicted from its structure, does not react with thiols so can’t react with plumbagin.
Fig 5 (not shown): The authors used UV-Vis and fluorescence spectroscopy to study the effect of NAC on the spectra of 1,4-naphthoquinone analogs. What likely effects would you expect to observe on the spectrum of lawsone?
None since lawsone does not react with the thiol-containing NAC
Fig 6A: The ultimate goal of the investigators was to make a 1,4-naphthoquinone derivative that was not cytotoxic which presumably results from its interactions with free thiols. They synthesized PTK1 as shown below. Why would this reaction produce a potential inhibitor that did not react with free thols in cells?

It has a methyl at C3 which would inhibit its reaction with free thiols
FIGURE 6B. Investigators studied the effects of PTK1 on P300 acetylation of HeLa core histones with different concentrations of PTK1 in the absence and presence of NAC (Figure B) as well the presence of free thiol in the presence or absence of PTK1 (Figure C).

What can you conclude from these graphs?
As expected, PTIK1 inhibits P300 acetylation with no effects from the addition of NAC.
Figure 6D: The figure below shows the UV-visible absorption spectrum of PTK1 in the in presence or absence of NAC?
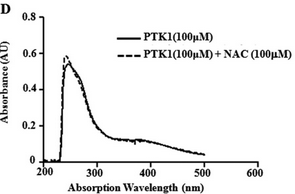
What conclusions can you draw?
PTK1 does not react with NAC, as expected.
Fig 6E: The figure below shows immunoblotting analysis (IB) of cell lysates upon 24 h of treatment by PTK1 using H3K9 acetylation antibodies. The concentrations of PTK1 used are indicated in the figure E. Error bars, S.D. What conclusions can you draw from the blots?
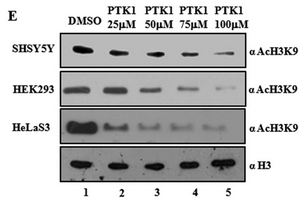
PTK1 acts as a potent inhibitor of acetylation of H3K9 in vivo.
7. How might PTK1 inhibit p300 KAT activity. Enzyme kinetic analyses would be useful. The enzyme has two substates, acetyl CoA and histones. Previous studies have shown that substrates bind to p300 , in an ordered mechanism. The non-competitive inhibitor PTK1 could bind to the free enzyme as well as to the substrate-bound form, resulting in an abortive ternary complex. T
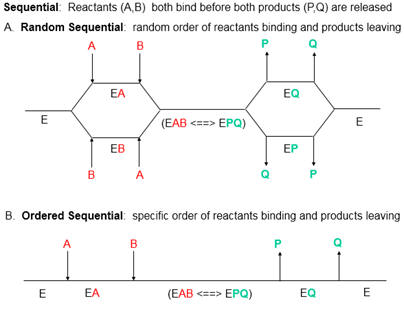
Presume that added reversible inhibitors behavior in a similar fashion with single and multi-substrate reactions. Here are some simple rules that apply to double reciprocal plots (1/v vs 1/[S} when 1 substrate is kept constant:
The slope changes when:
- I and S bind to the same form of the enzyme (for example E binds both S and I) OR
- I binds to a form of E (on the horizontal line) which is connected to the form that S binds, and I binds first (for example, I binds to E and then S binds).
The Y intercept changes when:
- I and S bind to different forms of the enzyme unless I binds first and the binding of I and S are in rapid equilibrium.
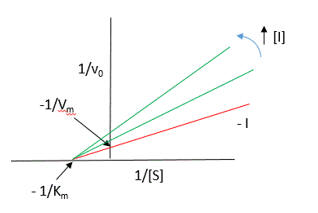
Figure 7 AB: Figure 7 A and B below show double reciprocal or Lineweaver-Burk plots (1/v vs 1/[S]) of p300 KAT with one substrate fixed and the other varying at different fixed concentrations of inhibitor PTK1.
- In Figure 7A, the concentration of core histones was kept constant at 1.7 μm while increasing the concentration of isotope labeled [3H]acetyl-CoA from 1 to 8 μm either in the absence (no inhibitor) or increasing concentrations of PTK1 (15, 20, 25, 35, and 45 μm). The direction of increasing inhibitor concentration is shown by the red arrow.
- In Figure 7B: inhibition of p300 KAT activity by PTK1 was studied at a fixed concentration of [3H]acetyl-CoA (1.6 μm) and increasing concentrations of core histones (5–70 nm) either in the absence of (no inhibitor) or increasing concentrations of PTK1 (15, 20, 25, 35, and 45 μm).
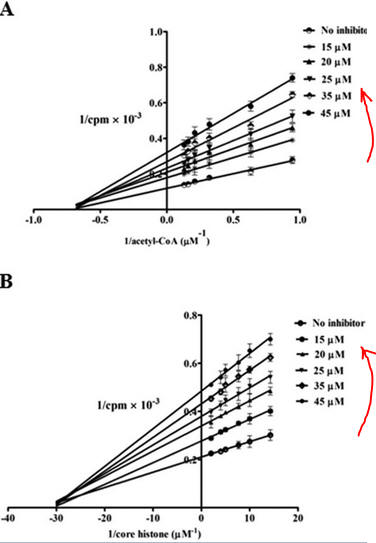
By analogy with single substrate reactions shown below in the linked chemical equations, which type of inhibition is displayed by PTK1?
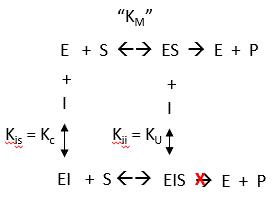
It appears to be a noncompetitive inhibit for each substrate. For a single substrate enzyme, noncompetitive inhibition is characterized by double-reciprocal plots which vary in slope and intersect on the x axis. This applies that the Km is unaltered by the inhibitor but the apparent Vm is decreased. This analysis is a bit simplified.