10.6: Lineage-through-Time Plots
- Page ID
- 21636
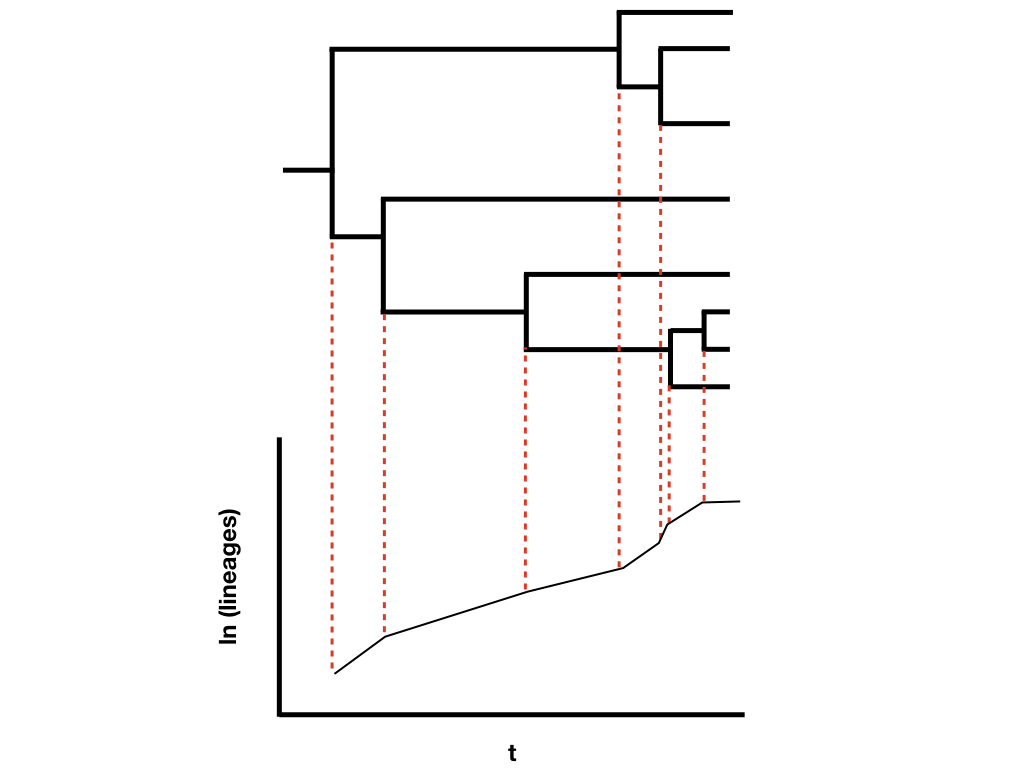
The other main way to quantify phylogenetic tree shape is by making lineage-through-time plots. These plots have time along the x axis (from the root of the tree to the present day), and the reconstructed number of lineages on the y-axis (Figure 10.8). Since we are usually considering birth-death models, where the number of lineages is expected to grow (or shrink) exponentially through time, then it is typical practice to log-transform the y-axis.
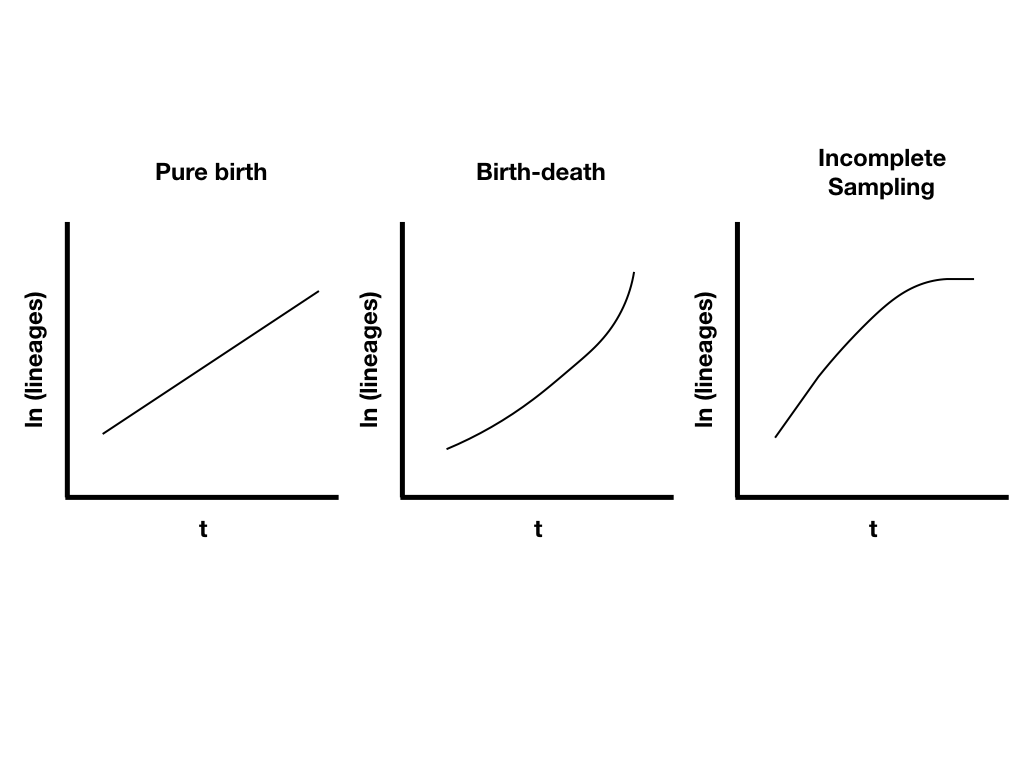
Lineage-through-time plots are effective ways to visualize patterns of lineage diversification through time. Under a pure-birth model on a semi-log scale, LTT plots follow a straight line on average (Figure 10.9A). By contrast, extinction should leave a clear signal in LTT plots because the probability of a lineage going extinct depends on how long it has been around; old lineages are much more likely to have been hit by extinction than relatively young lineages. We see this reflected in LTT plots as the “pull of the present” – an upturn in the slope of the LTT plot near the present day (Figure 10.9B). Incomplete sampling – that is, not sampling all of the living species in a clade – can also have a huge impact on the shape of LTT plots (Figure 10.9C). We will discuss LTT plots further in chapter 11, where we will use them to make inferences about patterns of lineage diversification through time.